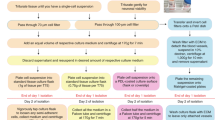
Abstract
The low number of neural progenitor cells (NPCs) present in the adult and aged primate brains represents a challenge for generating high-yield and viable in vitro cultures of primary brain cells. Here we report a step-by-step approach for the fast and reproducible isolation of high-yield and viable primary brain cells, including mature neurons, immature cells and NPCs, from adult and aged macaques. We describe the anesthesia, transcardial perfusion and brain tissue preparation; the subsequent microdissection of the regions of interest and their enzymatic dissociation, leading to the separation of single cells. The cell isolation steps of our protocol can also be used for routine cell culturing, in particular for NPC expansion and differentiation, suitable for studies of hippocampal neurogenesis in the adult macaque brain. The purified primary brain cells are largely free from myelin debris and erythrocytes, paving the way for multiple downstream applications in vitro and in vivo. When combined with single-cell profiling techniques, this approach allows an unbiased and comprehensive mapping of cell states in the adult and aged macaque brain, which is needed to advance our understanding of human cognitive and neurological diseases. The neural cell isolation protocol requires 4 h and a team of four to six users with expertize in primary brain cell isolation to avoid tissue hypoxia during the time-sensitive steps of the procedure.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
Data availability
The data supporting the findings of this study are available within the article, the primary supporting research paper of macaque hippocampus20. The scRNA-seq datasets shown in Figs. 4–6 are publicly available at ArrayExpress under the accession code E-MTAB-12399. Source data are provided with this paper.
Code availability
The code for processing the datasets is available as jupyter notebooks at GitHub (https://github.com/leitang607/macaque_Neural_cell).
References
Brewer, G. J. & Torricelli, J. R. Isolation and culture of adult neurons and neurospheres. Nat. Protoc. 2, 1490–1498 (2007).
Crouch, E. E. & Doetsch, F. FACS isolation of endothelial cells and pericytes from mouse brain microregions. Nat. Protoc. 13, 738–751 (2018).
Guo, W., Patzlaff, N. E., Jobe, E. M. & Zhao, X. Isolation of multipotent neural stem or progenitor cells from both the dentate gyrus and subventricular zone of a single adult mouse. Nat. Protoc. 7, 2005–2012 (2012).
Valtcheva, M. V. et al. Surgical extraction of human dorsal root ganglia from organ donors and preparation of primary sensory neuron cultures. Nat. Protoc. 11, 1877–1888 (2016).
Gray, D. T. & Barnes, C. A. Experiments in macaque monkeys provide critical insights into age-associated changes in cognitive and sensory function. Proc. Natl Acad. Sci. USA 116, 26247–26254 (2019).
Zhou, Y. et al. Atypical behaviour and connectivity in SHANK3-mutant macaques. Nature 570, 326–331 (2019).
Qiu, P. et al. BMAL1 knockout macaque monkeys display reduced sleep and psychiatric disorders. Natl Sci. Rev. 6, 87–100 (2019).
Park, T. I.-H. et al. Routine culture and study of adult human brain cells from neurosurgical specimens. Nat. Protoc. 17, 190–221 (2022).
Park, T. I.-H. et al. Isolation and culture of functional adult human neurons from neurosurgical brain specimens. Brain Commun. 2, fcaa171 (2020).
Nott, A., Schlachetzki, J. C. M., Fixsen, B. R. & Glass, C. K. Nuclei isolation of multiple brain cell types for omics interrogation. Nat. Protoc. 16, 1629–1646 (2021).
Lake, B. B. et al. Neuronal subtypes and diversity revealed by single-nucleus RNA sequencing of the human brain. Science 352, 1586–1590 (2016).
Habib, N. et al. Massively parallel single-nucleus RNA-seq with DroNc-seq. Nat. Methods 14, 955–958 (2017).
Zhu, Y. et al. Spatiotemporal transcriptomic divergence across human and macaque brain development. Science 362, eaat8077 (2018).
Han, L. et al. Cell transcriptomic atlas of the non-human primate Macaca fascicularis. Nature 604, 723–731 (2022).
Mathys, H. et al. Single-cell transcriptomic analysis of Alzheimer’s disease. Nature 570, 332–337 (2019).
Franjic, D. et al. Transcriptomic taxonomy and neurogenic trajectories of adult human, macaque, and pig hippocampal and entorhinal cells. Neuron 110, 452–469.e14 (2022).
Ayhan, F. et al. Resolving cellular and molecular diversity along the hippocampal anterior-to-posterior axis in humans. Neuron 109, 2091–2105.e6 (2021).
Tran, M. N. et al. Single-nucleus transcriptome analysis reveals cell-type-specific molecular signatures across reward circuitry in the human brain. Neuron 109, 3088–3103.e5 (2021).
Wei, J.-R. et al. Identification of visual cortex cell types and species differences using single-cell RNA sequencing. Nat. Commun. 13, 6902 (2022).
Hao, Z.-Z. et al. Single-cell transcriptomics of adult macaque hippocampus reveals neural precursor cell populations. Nat. Neurosci. 25, 805–817 (2022).
Zhong, S. et al. Decoding the development of the human hippocampus. Nature 577, 531–536 (2020).
Nowakowski, T. J. et al. Spatiotemporal gene expression trajectories reveal developmental hierarchies of the human cortex. Science 358, 1318–1323 (2017).
Schmitz, M. T. et al. The development and evolution of inhibitory neurons in primate cerebrum. Nature 603, 871–877 (2022).
Jiang, X. et al. Principles of connectivity among morphologically defined cell types in adult neocortex. Science 350, aac9462 (2015).
Ting, J. T. et al. Preparation of acute brain slices using an optimized N-methyl-d-glucamine protective recovery method. J. Vis. Exp. 26, 53825 (2018).
Huang, W. et al. Linking transcriptomes with morphological and functional phenotypes in mammalian retinal ganglion cells. Cell Rep. 40, 111322 (2022).
Mich, J. K. et al. Functional enhancer elements drive subclass-selective expression from mouse to primate neocortex. Cell Rep. 34, 108754 (2021).
Zeisel, A. et al. Molecular architecture of the mouse nervous system. Cell 174, 999–1014.e22 (2018).
Saunders, A. et al. Molecular diversity and specializations among the cells of the adult mouse brain. Cell 174, 1015–1030.e16 (2018).
Braidy, N. et al. Effects of kynurenine pathway metabolites on intracellular NAD synthesis and cell death in human primary astrocytes and neurons. Int. J. Tryptophan Res. 2, 61–69 (2009).
Bohár, Z., Toldi, J., Fülöp, F. & Vécsei, L. Changing the face of kynurenines and neurotoxicity: therapeutic considerations. Int. J. Mol. Sci. 16, 9772–9793 (2015).
Kanwar, J. R., Kanwar, R. K. & Krissansen, G. W. Simultaneous neuroprotection and blockade of inflammation reverses autoimmune encephalomyelitis. Brain 127, 1313–1331 (2004).
Minnella, A. M. et al. Excitotoxic superoxide production and neuronal death require both ionotropic and non-ionotropic NMDA receptor signaling. Sci. Rep. 8, 17522 (2018).
Lysko, P. G., Webb, C. L., Yue, T. L., Gu, J. L. & Feuerstein, G. Neuroprotective effects of tetrodotoxin as a Na+ channel modulator and glutamate release inhibitor in cultured rat cerebellar neurons and in gerbil global brain ischemia. Stroke 25, 2476–2482 (1994).
Tasic, B. et al. Adult mouse cortical cell taxonomy revealed by single cell transcriptomics. Nat. Neurosci. 19, 335–346 (2016).
Hrvatin, S. et al. Single-cell analysis of experience-dependent transcriptomic states in the mouse visual cortex. Nat. Neurosci. 21, 120–129 (2018).
Armand, E. J., Li, J., Xie, F., Luo, C. & Mukamel, E. A. Single-cell sequencing of brain cell transcriptomes and epigenomes. Neuron 109, 11–26 (2021).
Lee, K. et al. Human in vitro systems for examining synaptic function and plasticity in the brain. J. Neurophysiol. 123, 945–965 (2020).
Fattorelli, N. et al. Stem-cell-derived human microglia transplanted into mouse brain to study human disease. Nat. Protoc. 16, 1013–1033 (2021).
Hochgerner, H., Zeisel, A., Lönnerberg, P. & Linnarsson, S. Conserved properties of dentate gyrus neurogenesis across postnatal development revealed by single-cell RNA sequencing. Nat. Neurosci. 21, 290–299 (2018).
Trevino, A. E. et al. Chromatin accessibility dynamics in a model of human forebrain development. Science 367, eaay1645 (2020).
La Manno, G. et al. RNA velocity of single cells. Nature 560, 494–498 (2018).
Kang, C.-C. et al. Single cell-resolution western blotting. Nat. Protoc. 11, 1508–1530 (2016).
Stevenson, R., Samokhina, E., Rossetti, I., Morley, J. W. & Buskila, Y. Neuromodulation of glial function during neurodegeneration. Front. Cell. Neurosci. 14, 278 (2020).
Rustenhoven, J., Jansson, D., Smyth, L. C. & Dragunow, M. Brain pericytes as mediators of neuroinflammation. Trends Pharmacol. Sci. 38, 291–304 (2017).
Darmanis, S. et al. A survey of human brain transcriptome diversity at the single cell level. Proc. Natl Acad. Sci. USA 112, 7285–7290 (2015).
Zhang, Y. et al. Purification and characterization of progenitor and mature human astrocytes reveals transcriptional and functional differences with mouse. Neuron 89, 37–53 (2016).
Spaethling, J. M. et al. Primary cell culture of live neurosurgically resected aged adult human brain cells and single cell transcriptomics. Cell Rep. 18, 791–803 (2017).
Roberts, A. C. & Clarke, H. F. Why we need nonhuman primates to study the role of ventromedial prefrontal cortex in the regulation of threat- and reward-elicited responses. Proc. Natl Acad. Sci. USA 116, 26297–26304 (2019).
Cao, J. et al. Joint profiling of chromatin accessibility and gene expression in thousands of single cells. Science 361, 1380–1385 (2018).
Wu, Y. E., Pan, L., Zuo, Y., Li, X. & Hong, W. Detecting activated cell populations using single-cell RNA-seq. Neuron 96, 313–329.e6 (2017).
Iannielli, A. et al. Pharmacological inhibition of necroptosis protects from dopaminergic neuronal cell death in Parkinson’s disease models. Cell Rep. 22, 2066–2079 (2018).
Saleem, K. S. & Logothetis, N. K. A Combined MRI and Histology Atlas of the Rhesus Monkey Brain in Stereotaxic Coordinates (Academic Press, 2012).
Zhu, C. et al. An ultra high-throughput method for single-cell joint analysis of open chromatin and transcriptome. Nat. Struct. Mol. Biol. 26, 1063–1070 (2019).
Xiao, D. et al. Direct reprogramming of fibroblasts into neural stem cells by single non-neural progenitor transcription factor Ptf1a. Nat. Commun. 9, 2865 (2018).
Yates, A. D. et al. Ensembl 2020. Nucleic Acids Res. 48, D682–D688 (2020).
Wolock, S. L., Lopez, R. & Klein, A. M. Scrublet: computational identification of cell doublets in single-cell transcriptomic data. Cell Syst. 8, 281–291.e9 (2019).
Korsunsky, I. et al. Fast, sensitive and accurate integration of single-cell data with Harmony. Nat. Methods 16, 1289–1296 (2019).
Acknowledgements
We gratefully acknowledge C. Xu for language editing and proofreading. This research was supported by grants from the National Key R&D Program of China (2018YFA0108300), the Natural Science Foundation of China (81961128021, 81870682, 82201231 and 32270864), the National Key R&D Program of China (2022YEF0203200), the Major Project on Brain Science and Brain-Like Computing of the Ministry of Science and Technology of China (2021ZD0200103), the Basic and Applied Basic Research Foundation of Guangdong Province (2023A1515011593), the Guangdong Provincial Key R&D Programs (2018B030335001 and 2018B030337001), the Science and Technology Program of Guangzhou (202007030010 and 202007030011), the China Postdoctoral Science Foundation (2022M713609) and Science and Technology Planning Projects of Guangzhou City (2019A1515012033).
Author information
Authors and Affiliations
Contributions
S.L. conceived and supervised the protocol. S.L., J.-R.W. and D.X. contributed to the development of the protocol and wrote the manuscript. L.T., N.X., R.L., Y.S., Z.X., X.S. J.G. and M.X. helped with protocol optimization.
Corresponding author
Ethics declarations
Competing interests
S.L. and J.-R.W. have developed the patent for this work (no. ZL202111114666.6, Sun Yat-sen University Zhongshan Ophthalmic Center).
Peer review
Peer review information
Nature Protocols thanks Orly Lazarov and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Related links
Key references using this protocol
Wei, J. R. et al. Nat. Commun. 13, 6902 (2022): https://doi.org/10.1038/s41467-022-34590-1
Hao, Z. Z. et al. Nat. Neurosci. 25, 805–817 (2022): https://doi.org/10.1038/s41593-022-01073-x
Xiao, D. et al. Nat. Commun. 9, 2865 (2018): https://doi.org/10.1038/s41467-018-05209-1
Extended data
Extended Data Fig. 1 Representative figures and electrophysiological recordings demonstrating the viability of acute brain slices in the adult macaque cortex.
a, Representative images were acquired from the cortex in acute slices from adult macaques. Scale bar, 20 μm. b, The single whole-cell patch-clamp electrophysiological recordings of pyramidal and interneuron neurons showed a multi-action potential firing cell. Pyr, pyramidal neuron; InN, inhibitory neurons.
Extended Data Fig. 2 Primary culture of cells and labeling assessments from the hippocampus of adult macaque monkeys.
a, b, The neurospheres are formed from cells cultured from the hippocampus of adult macaque monkeys. Three weeks after cell seeding, NPCs elucidated spindle morphology in a. Thirty days after cell seeding in b. D, days. Scale bars, a, 40 μm; b, 160 μm. c, d, NPCs view in P1 and P4 passages; P, passages. Scale bars, c, 80 μm; d, 60 μm. e–j, Cultured NPCs form proliferating neurospheres positive for NESTIN, MKI67, PAX6, VIMENTIN, SOX4 and HMGB2. Scale bars, 40 μm.
Extended Data Fig. 3 Preparation of fire-polishing glass Pasteur pipettes.
a, The glass pipette is heated in the top part of the flame and spun evenly. b, The heated pipette is quickly drawn out. c, The sharply drawn-out edge is quickly heated to form a smooth and round edge. d–f, Example of three fire-polishing glass Pasteur pipettes with 150-μm, 350-μm, 650-μm bores. Scale bars, 200 μm.
Supplementary information
Supplementary Video 1
The morphologies of recorded neurons.
Supplementary Video 2
The morphologies of recorded neurons.
Source data
Source Data Figs. 7 and 8
Statistical source data for Figs. 7g and 8c.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Wei, JR., Xiao, D., Tang, L. et al. Neural cell isolation from adult macaques for high-throughput analyses and neurosphere cultures. Nat Protoc 18, 1930–1957 (2023). https://doi.org/10.1038/s41596-023-00820-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41596-023-00820-z
This article is cited by
-
Patch-seq: Advances and Biological Applications
Cellular and Molecular Neurobiology (2024)
-
Multimodal Nature of the Single-cell Primate Brain Atlas: Morphology, Transcriptome, Electrophysiology, and Connectivity
Neuroscience Bulletin (2024)
-
A primate nigrostriatal atlas of neuronal vulnerability and resilience in a model of Parkinson’s disease
Nature Communications (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.