Abstract
Interactions between mutations (epistasis) can add substantial complexity to genotype-phenotype maps, hampering our ability to predict evolution. Yet, recent studies have shown that the fitness effect of a mutation can often be predicted from the fitness of its genetic background using simple, linear relationships. This phenomenon, termed global epistasis, has been leveraged to reconstruct fitness landscapes and infer adaptive trajectories in a wide variety of contexts. However, little attention has been paid to how patterns of global epistasis may be affected by environmental variation, despite this variation frequently being a major driver of evolution. This is particularly relevant for the evolution of drug resistance, where antimicrobial drugs may change the environment faced by pathogens and shape their adaptive trajectories in ways that can be difficult to predict. By analyzing a fitness landscape of four mutations in a gene encoding an essential enzyme of P. falciparum (a parasite cause of malaria), here we show that patterns of global epistasis can be strongly modulated by the concentration of a drug in the environment. Expanding on previous theoretical results, we demonstrate that this modulation can be quantitatively explained by how specific gene-by-gene interactions are modified by drug dose. Importantly, our results highlight the need to incorporate potential environmental variation into the global epistasis framework in order to predict adaptation in dynamic environments.
Similar content being viewed by others
Introduction
The topography of genotype-phenotype maps has critical consequences for the predictability of evolutionary trajectories. This topography emerges from complex interactions between genetic elements, from single nucleotides to protein residues and metabolic pathways1,2,3,4,5. Despite this complexity, recent work has shown that epistasis—the nonlinear interaction between parcels of genetic information—often has a “global” component6,7,8,9,10,11,12,13,14,15, emerging in the form of simple relationships between the fitness effect of a mutation and the fitness of the genetic background where it arises (Fig. 1a–c). Global epistasis has become a central concept in modern conversations surrounding the fundamentals of evolutionary processes. Negatively sloped correlations have been more commonly reported, both in the form of diminishing returns and increasing costs epistasis (where the fitness effect of a mutation becomes less beneficial or more deleterious, respectively, in fitter genetic backgrounds)6,7,8, 11,12,13. Positive slopes (increasing returns or decreasing costs epistasis) have also been found in low-dimensional landscapes7,16,17, and theory suggests they may be more common near fitness peaks5,18,19.
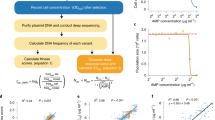
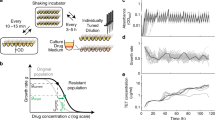
a We reanalyzed a dataset consisting of 15 genotypes of the P. falciparum parasite, each carrying a different combination of four mutations: C59R, I164L, N51I and S108N. b We examined the fitness effect of a focal mutation (in this illustration, mutation C59R) in different genetic backgrounds carrying combinations of the other three mutations. Here, colored/gray loci represent the presence/absence of the mutation. c In the absence of pyrimethamine, a negative correlation is observed between the fitness effect of mutation C59R and the fitness of its genetic background. We quantify the strength of epistasis as the variance in the mutation’s fitness effects relative to the variance in fitness across its genetic backgrounds (var Δf / var f(B)). The degree to which epistasis is global is quantified as the R2 of the linear regression between the fitness effect of the mutation and the fitness of its genetic background. d The variance ratio and the R2 can be seen as a mutation’s “coordinates” in a conceptual “epistasis map''.
The emergence of global epistasis evidences the existence of regularities in genotype-phenotype maps6,7,8,9,10, which have been leveraged in recent methodologies to infer complete adaptive landscapes under defined, steady environmental conditions20,21,22. Yet, the magnitude of mutational fitness effects23,24,25,26 and epistatic interactions27,28,29,30,31,32, and thus the topography of fitness landscapes33,34,35, can generally depend on environmental variables. Populations are often subject to natural or anthropogenic environmental fluctuations which can dictate their evolutionary fate26, 29,36,37. Importantly, this effect has been widely described in the evolution of antimicrobial resistance (AMR)29,32,38,39,40. Learning how global epistasis patterns may be shaped by environmental factors is therefore critical to our ability to predict adaptation in changing environments, and especially in the context of AMR evolution. This is yet an open question largely because understanding how global epistasis emerges from fine-grained genetic interactions (which might be subject to environmental regulation) is still in its early days.
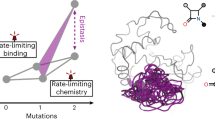
To address these questions, here we analyze a previously published fitness landscape31,34,41 consisting of 15 genotypes of the P. falciparum malaria parasite. Each genotype carries a different combination of four mutations (amino acid substitutions C59R, I164L, N51I and S108N) at four sites of the P. falciparum gene for the dihydrofolate reductase (DHFR) enzyme, which confers resistance to several antifolate drugs (Fig. 1a). These particular four mutations have been associated to antimalarial drug resistance in numerous studies across multiple geographical locations42,43,44,45. The 15 genotypes were cultured in a concentration gradient of pyrimethamine or cycloguanil (antifolate drugs commonly used to treat malaria), and fitness was quantified as the growth rate relative to that of the slowest growing genotype in the absence of drug. Notably, epistasis makes it so the genotype carrying all four mutations has lower fitness than the sum of all mutations' individual fitness effects at the highest drug concentrations (which can be explained by resistance mutations often being redundant46,47); but also reduces the deleterious effect of the four mutations when there is no drug in the environment (Supp. Figure 1). This observation highlights the need to characterize how drug dose modulates epistasis in this particular landscape in order to understand the emergence and prevalence of resistant P. falciparum genotypes.
We thus asked whether the concentration of drug in the environment may modulate the patterns of global epistasis observed for a particular mutation and, if so, whether we can trace back the origins of this modulation to specific gene-by-environment interactions. For each of the four mutations, we focused on (a) to what extent the fitness effect of the mutation depends on its genetic background (i.e., how much epistasis there is), (b) how correlated the fitness effect of the mutation is with the fitness of its genetic background (i.e., to what degree epistasis is global), and (c) what the shape of global epistasis is (e.g., diminishing returns, accelerating returns, etc.).
Our results show that drug concentration strongly modulates global epistasis in this particular landscape. Extending previous theoretical results, we demonstrate that this modulation can be explained by specific gene-by-gene interactions and how they are affected by the environment. In particular, we mathematically define a set of “effective” genetic interactions, showing that their distribution determines the strength and shape of global epistasis across drug doses. Our results highlight an avenue for quantitatively connecting fine-grained gene-by-environment and gene-by-gene-by-environment interactions with the emergence of global epistasis and the topography of fitness landscapes.
Results
Mutations exhibit variation in the strength and shape of global epistasis
We first asked to what extent mutations in our landscape exhibit variable fitness effects depending on their genetic background, that is, what the magnitude of epistasis is in this system. For a given mutation (which we denote as the focal mutation i), we refer to the set of genotypes not carrying that mutation as the genetic backgrounds (Fig. 1b). We denote f(B) the fitness of one of such backgrounds B. Calling B+i the genotype resulting from adding mutation i to the genetic background B, the fitness effect of mutation i is quantified simply as: Δfi = f(B + i)−f(B), that is, the difference in fitness between genotypes B+i and B (see Fig. 1b for an illustration with C59R as the focal mutation). If a mutation had only weak epistatic interactions with other loci, we would see that its fitness effect Δfi remains roughly constant across all genetic backgrounds (i.e., the effect of the mutation is essentially additive). On the other hand, high levels of epistasis would make this fitness effect largely dependent on the presence or absence of other mutations, exhibiting large variation across backgrounds. We thus quantified the strength of epistasis for a focal mutation i as the variance of Δfi relative to the variance in fitness across the genetic backgrounds of that mutation: var Δfi/var f(B) (see Fig. 1c for the case with C59R as the focal mutation).
We also asked the extent to which the fitness effect of a mutation may be well predicted by a simple linear model linking it to the fitness of its genetic background, that is, to what degree is epistasis global. Note that, in general, the relationship between Δf and f(B) need not always be truly linear, and other studies have found different types of relationships between the two magnitudes14,15. The linear fits to our data should be interpreted simply as the most parsimonious statistical models one can use to link the fitness effect of a mutation to the background fitness.
If epistasis was strong for a given mutation (large var Δfi/var f(B)) and we observed a high correlation between f(B) and Δf, we would interpret this as epistasis being largely global: while the fitness effect of the mutation strongly depends on the genetic structure of its background, it can still be well estimated from the background fitness through a simple linear model (Fig. 1c). If, on the other hand, the correlation between f(B) and Δf was weak, epistasis would be largely idiosyncratic (as opposed to global). We quantified the degree to which epistasis is global for a given mutation as the coefficient of determination (R2) of the regression between f(B) and Δf for that mutation (Fig. 1c).
We propose that these two quantities (the variance ratio var Δfi/var f(B) and the R2) define a “map of epistasis” in which different mutations may occupy different positions. The edges of this map correspond to limit cases where epistasis is either very strong or very weak, and either entirely global or entirely idiosyncratic (Fig. 1d). In our system, and when there is no drug in the environment, we observed that the strength of epistasis is similar for all mutations (var Δfi/var f(B)∼1 for all four), but the degree to which this epistasis is global exhibits substantial variation (R2∼0.7 for mutation C59R to R2∼0.2 for mutation S108N).
Drug dose modulates the strength and shape of global epistasis
To analyze how the environment may modulate epistasis in our system, we considered every concentration of pyrimethamine ranging from 10−2 μM to 103 μM, as well as the no drug control, in the dataset described above (Fig. 2a). We found that the strength and shape of global epistasis varied substantially for all mutations as the concentration of pyrimethamine increased. As an illustration, in Fig. 2b we show that mutation C59R goes from exhibiting a pattern of diminishing returns (smaller fitness effects in higher-fitness backgrounds) at low drug doses to exhibiting increasing returns (larger positive fitness effects in higher-fitness backgrounds) at high doses. For other mutations, epistasis is modulated differently (Supp. Figure 2).
a We analyzed how an environmental perturbation (in the form of a pyrimethamine concentration gradient) affects the global epistasis patterns observed in our four-mutation landscape. b As an example, we show that increasing drug dose alters the global epistasis pattern for mutation C59R, going from diminishing returns to increasing returns. c Drug concentration affects epistasis differently for each mutation, which can be thought of as mutations following different paths in our “epistasis map''. Arrows follow the direction of change as drug dose increases from 0 to 103 μM. d–f The strength of epistasis (variance ratio var Δf/var f(B)), the degree to which epistasis is global (R2) and the shape of global epistasis (slope of the regression between Δf and f(B)) are modulated differently for each mutation.
Mutations can be seen as describing a “path” through the map of epistasis we defined above (Fig. 2c) as the environment changes, i.e., as drug concentration increases. For instance, mutation S108N becomes largely idiosyncratic, moving to the bottom-right corner of the map. Epistasis for mutation N51I becomes weaker as drug dose increases, i.e., this mutation describes a path towards the left side of the map. For mutations C59R and I164L, the strength of epistasis remains roughly constant across drug concentrations (variance ratio var Δfi/var f(B)∼1), but epistasis becomes more global (higher R2) for I164L and more idiosyncratic (lower R2) for C59R at intermediate drug doses. Beyond modulating the variance ratio (Fig. 2d) and the R2 (Fig. 2e), drug dose also changes the shape of global epistasis. Interestingly, while the slopes of the linear regressions between f(B) and Δf are similar for all mutations at low doses, they become substantially different at higher concentrations (Fig. 2f). Note that these three quantities (variance ratio, R2, and slope) are not independent, as they satisfy b2 = R2 var Δf/var f(B) (where we have denoted the slope as b).
Using cycloguanil instead of pyrimethamine as the environmental perturbation modifies the paths that mutations follow in the epistasis map, but some qualitative behaviors are similar for both drugs: for instance, epistasis for mutation S108N also becomes largely idiosyncratic at high cycloguanil doses, while epistasis for mutation I164L becomes strongly global at intermediate doses (Supp. Figures 3 and 4). Together, these observations highlight the role of the environment in dictating the strength and shape of global epistasis in this particular system.
Environmental modulation of global epistasis is explained by changes in background fitness effects and interactions
Can we attribute these environmental effects on global epistasis to specific interactions between the four mutations in this landscape? Recent theory has shown that the slope of the regression between Δf and f(B) for a given mutation can be quantitatively explained in terms of fine-grained genetic interactions between that mutation and every other loci in its genetic background48, 49 (equations 1 to 3 in Box 1). Here we show that the strength of epistasis (variance ratio var Δf/var f(B)) and the degree to which epistasis is global (R2 of the Δf-vs-f(B) regression) can also be quantitatively linked to specific gene-by-gene interactions (Box 1 and Supplementary Notes).
For that, we define the “effective interaction” between mutations i and j (denoted βij) as the average magnitude of epistasis for those mutations (i.e., the average deviation between the fitness of a genotype carrying both mutations with respect to the additive expectation that they do not interact, see Box 1) with respect to the average fitness effect of mutation j across all genetic backgrounds of i (see equation 2 in Box 1). Expanding on previous theoretical results, in Box 1 we show that the variance ratio, R2, and global epistasis slope for a focal mutation i can all be quantitatively linked to the mean and standard deviation of the distribution of that mutation’s effective interactions (see also Supplementary Notes for a more detailed derivation of the expressions in Box 1).
In light of equations 1 to 5 (Box 1), we may expect environmental factors to modulate the strength and shape of global epistasis for a focal mutation through two different mechanisms: either by modifying the fitness effects of the background mutations, or by modulating the interactions between them and the focal (Fig. 3a, b). To test these two potential mechanisms, we quantified the terms Δfj and ϵij for every pair of mutations i and j in the dataset, in every possible genetic background and for every drug concentration between 0 and 103 μM. We started by analyzing the case with C59R as the focal mutation i, considering the other three mutations as non-focal background mutations (j).
a We quantified the fitness effect of mutations I164L (green), N51I (yellow) and S108N (blue) on each background not carrying the focal mutation (here C59R) and for every pyrimethamine dose. Each dot corresponds to a different background. Lines represent means across all backgrounds. b In every background not carrying mutation C59R, we also quantified the magnitude of epistasis between C59R and each of the three background mutations, i.e., the deviation between the fitness of the double mutant with respect to the additive expectation (see Box 1). c Distributions of effective interactions (as defined in Box 1) for mutation C59R at different drug concentrations. Each color corresponds to the effective interaction of C59R with each of the other three mutations. Dot size is proportional to the weight of that effective interaction (as defined in equation 3). Black lines and error bars represent weighted means and weighted standard deviations across the n = 3 background mutations. d–f As drug dose increases, the moments of the distribution change (dashed lines), as do the strength of epistasis (variance ratio var Δf/var f(B)), the degree to which epistasis is global (R2), and global epistasis slope for the focal mutation C59R (solid red lines).
We quantified the fitness effects Δfj of each non-focal mutation in every genetic background not carrying C59R (see Box 1). As shown in Fig. 3a, drug concentration modulates these fitness effects differently for each background mutation. For instance, mutation S108N has a very small average fitness effect (〈ΔfS108N〉∼0) at low doses, which becomes strongly positive (〈ΔfS108N〉 > 0) as the concentration of pyrimethamine increases in the environment (Fig. 3a, blue). On the other hand, mutations I164L and N51I have small and negative average fitness effects (〈ΔfI164L〉, 〈ΔfN51I〉 < 0) at low doses, but the fitness effect of N51I becomes slightly positive (〈ΔfN51I〉 > 0) at high doses (Fig. 3a, yellow).
We then analyzed the average magnitude of epistasis (〈ϵij〉, quantified as explained in Box 1) between the focal mutation C59R and each of the three background mutations. Again, we found that the environment modulates these epistatic effects differently for each background mutation (Fig. 3b). For instance, mutation S108N goes from exhibiting low positive epistasis with C59R (〈ϵC59R,S108N〉 > 0) at low doses to exhibiting very large positive epistasis (〈ϵC59R,S108N〉 > > 0) with C59R at higher concentrations (Fig. 3b, blue). In turn, mutation N51I exhibits, on average, moderate positive epistasis with the focal mutation C59R (〈ϵC59R,N51I〉 > 0) at low drug doses, and moderate negative epistasis (〈ϵC59R,N51I〉 < 0) at high doses (Fig. 3b, yellow).
Using equations 2 and 3 in Box 1, we then quantified the magnitude of the effective interaction (β) between C59R and each of the three non-focal mutations for every drug dose. We also quantified the weight (ω) corresponding to each background mutation. In Fig. 3c we show how the distribution of effective interactions of C59R changes as pyrimethamine concentration increases. Most notably, at low doses the effective interactions of C59R with I164L (green dots in Fig. 3c) and N51I (yellow dots in Fig. 3c) have the largest weights, and are negative as a result of 〈ϵij〉 and 〈Δfj〉 having opposite signs for these two mutations. As drug concentration increases, however, the weight of the positive effective interaction between C59R and S108N (blue dots in Fig. 3c) becomes larger, since the average fitness effect of S108N increases substantially.
As explained in Box 1, the (weighted) first and second moments of the distribution of a mutation’s effective interactions modulate the strength and shape of global epistasis for that mutation. In particular, the weighted mean estimates the global epistasis slope (equation 1), the weighted second moment estimates the variance ratio var Δf/var f(B) (equation 4), and the coefficient of variation estimates the R2 (equation 5). In Fig. 3d–e we show that for mutation C59R, equations 1 to 5 do a good job at capturing how the slope and variance ratio change across drug concentrations. For the R2 (Fig. 3f) the quantitative agreement is weaker, but the qualitative behavior is well captured — namely, the R2 being high at low and high doses, and dropping at intermediate concentrations.
This analysis allows us to identify the mechanisms underlying the environmental modulation of global epistasis for this mutation. As we had shown in Fig. 2b, C59R exhibits a pattern of diminishing returns at low drug doses and a pattern of increasing returns at high doses. This can be explained by the negative effective interactions of C59R with I164L and N51I dominating at low pyrimethamine concentrations, and the positive effective interaction between C59R and S108N dominating at high drug concentrations (Fig. 3c).
Considering any of the other three mutations (instead of C59R) as the focal yields a similarly good agreement of the empirically observed variance ratio, slope, and R2 with the values estimated using equations 1 to 5. For every mutation in the dataset, we quantified its distribution of effective interactions with all other mutations at all drug doses. In Fig. 4 we show that the path described by each mutation in the “epistasis map” closely matches the path expected from estimating var Δf/var f(B) and R2 using the distribution of a mutation’s effective interactions.
a We show the paths that mutations follow in the “map of epistasis” as drug concentration increases. Solid lines: paths obtained from empirically quantifying the variance ratios and R2 (same as in Fig. 2c). Dashed lines: paths obtained by estimating variance ratios and R2 from equations 4 and 5 in Box 1. b–d We compare the empirically obtained values for the variance ratio, R2, and global epistasis slope for every mutation in the dataset and at all concentrations of pyrimethamine with the values estimated from each mutation’s distribution of effective interactions (equations 1, 4, and 5 in Box 1).
Equations 1-5 can generally provide accurate estimates for the strength and shape of global epistasis across concentrations of pyrimethamine (Fig. 4b–d) or cycloguanil (Supp. Figure 5). Importantly, however, these estimates sometimes deviate from the empirically observed variance ratio, R2 and slope. This is perhaps most evident in Figs. 3f and 4d (red dots), showing that the R2 for mutation C59R is overestimated by equation 5 at high pyrimethamine doses. As explained in Box 1, equations 1 to 5 result from partially neglecting the effect of higher-order interactions (HOIs) between mutations. Therefore, the deviations observed in this particular case point to mutation C59R engaging in strong HOIs at high drug doses. Interestingly, these deviations seem to be smaller for the cycloguanil data (Supp. Figure 5), indicating a smaller effect of HOIs in the presence of this drug. Together, these results highlight the utility of defining effective interactions in order to uncover which fine-grained genetic interactions, and through which mechanisms, play a more prominent role in shaping global epistasis across environments.
Discussion
The observation that epistasis often has a global component has set a stepping stone in our ability to understand and predict adaptation. For instance, diminishing returns epistasis can explain the strikingly conserved patterns of declining adaptability reported in long-term evolution experiments7,8,9,50,51. While there have been increasing efforts to understand how patterns of global epistasis emerge from fine-grained genetic interactions13,14,48,49,52,53, less attention has been paid to how they might be affected by the environment. This is particularly important in the context of AMR evolution, where the use of antimicrobial drugs can often exert evolutionary pressures that reshape the topography of genotype-phenotype maps and drive the emergence of resistant genotypes.
In this work, we have examined how the concentration of a drug in the environment affects the patterns of global epistasis in a particular fitness landscape defined by four mutations often found in resistant clones of the P. falciparum malaria parasite42,43,44. Our results show that the strength of epistasis, the degree to which epistasis is global, and the shape of global epistasis are strongly modulated by drug concentration in this landscape. Furthermore, we have shown that the origins of this modulation can be attributed to specific gene-by-environment and gene-by-gene-by-environment interactions, expanding on previous theoretical results connecting the emergence of global epistasis to pairwise and higher-order genetic interactions48,49. While this connection was originally formulated under the assumption of widespread epistasis48, theory had suggested that it may hold even when genetic interactions are sparse49.
Our results provide direct empirical evidence of this theoretical framework successfully explaining global epistasis patterns in a particular drug resistance fitness landscape, even when these are dominated by a single interaction. We have shown that, at high drug doses, the strength and shape of global epistasis for mutation C59R is dictated by its interaction with a single background mutation (S108N). This is consistent with recent work demonstrating that global epistasis can emerge as a result of sparse genetic interactions14. More generally, we have shown that the collective action of all mutations’ epistatic and fitness effects determines the shape of a distribution of “effective interactions,” which we have defined mathematically building on recent theory from quantitative genetics. We have demonstrated that the moments of this distribution determine the properties of global epistasis for a given mutation — namely, its strength and the shape of the relationship between the background fitness and the mutation’s fitness effect.
Here we analyzed a low-dimensional landscape, which allowed us to directly test the fitness effect of every mutation, as well as every possible epistatic coefficient between all pairs of mutations, in almost every genetic background. All mutations we considered correspond to a same gene (encoding the DHFR enzyme of P. falciparum), and thus we might expect interactions between them to be stronger than with other potential mutations in more distal sites of the genome. Further work will be required to determine whether effective interactions can be reliably defined and quantified in higher-dimensional landscapes, including mutations in different genes or pathways, where empirically measuring the fitness of every genotype across environments might be more challenging. For instance, the contribution of each background locus to the global epistasis pattern of a focal mutation can be expected to become smaller (lower weight ω as defined in equation 3) as the size of the landscape increases. Whether this results in stronger or weaker global epistasis patterns will generally depend on the structure of epistasis among all mutations considered, which can be expected to vary across landscapes of different sizes. In addition, here we have focused on two specific drugs as our environmental variables. It will be important to test whether different forms of environmental perturbations (e.g., temperature or pH variations), and within what ranges, have the potential to modulate global epistasis in this landscape and others. Most importantly, our work suggests that the global epistasis framework can be readily extended to account for environmental variables, which might be critical to predict adaptive trajectories under changing environments in the context of AMR evolution.
Methods
Data analysis
Data was obtained from the original publications31,34,41. The genotype carrying mutations I164L and S108N was removed from the data due to having inconsistent fitness values across two no-drug controls. All analyses were carried out using R version 4.3.1.
Quantification of effective interactions
For a given pair of mutations i and j, we denote B a genetic background not carrying any of the two. We denote B+i and B+j the genotypes resulting from mutating i or j, respectively, in the background B. The double mutant is denoted B+i+j. To compute the effective interaction βij (see equation 1 in Box 1), we consider every possible background B and quantify
We then compute 〈Δfj〉 and 〈ϵij〉 by averaging the expressions above across every possible genetic background B. If any of the terms on the right-hand side of the equations above were missing (due to incomplete data), we computed the averages across the known terms only.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The data analyzed in this study were obtained from the original publications31, 34,41 and is also available at https://github.com/jdiazc9/env_global_epist54. Code used to generate all figures can be found in the same GitHub repository.
Code availability
All code is available at https://github.com/jdiazc9/env_global_epist54.
References
Weinreich, D. M., Delaney, N. F., DePristo, M. A. & Hartl, D. L. Darwinian evolution can follow only very few mutational paths to fitter proteins. Science 312, 111–114 (2006).
De Visser, J. & Krug, J. Empirical fitness landscapes and the predictability of evolution. Nat. Rev. Genet. 15, 480–490 (2014).
Starr, T. N. & Thornton, J. W. Epistasis in protein evolution. Protein Sci. 25, 1204–1218 (2016).
Kryazhimskiy, S. Emergence and propagation of epistasis in metabolic networks. eLife 10, e60200 (2021).
Bank, C. Epistasis and adaptation on fitness landscapes. Ann. Revi. Ecol. Evol. Syst. 53, 457–479 (2022).
MacLean, R., Perron, G. & Gardner, A. Diminishing returns from beneficial mutations and pervasive epistasis shape the fitness landscape for rifampicin resistance in pseudomonas aeruginosa. Genetics 186, 1345–1354 (2010).
Khan, A. I., Dinh, D. M., Schneider, D., Lenski, R. E. & Cooper, T. F. Negative epistasis between beneficial mutations in an evolving bacterial population. Science 332, 1193–1196 (2011).
Chou, H.-H., Chiu, H.-C., Delaney, N. F., Segrè, D. & Marx, C. J. Diminishing returns epistasis among beneficial mutations decelerates adaptation. Science 332, 1190–1192 (2011).
Perfeito, L., Sousa, A., Bataillon, T. & Gordo, I. Rates of fitness decline and rebound suggest pervasive epistasis. Evolution 68, 150–162 (2014).
Kryazhimskiy, S., Rice, D. P., Jerison, E. R. & Desai, M. M. Global epistasis makes adaptation predictable despite sequence-level stochasticity. Science 344, 1519–1522 (2014).
Schoustra, S., Hwang, S., Krug, J. & de Visser, J. A. G. Diminishing-returns epistasis among random beneficial mutations in a multicellular fungus. Proc. Royal Soc. B: Biol. Sci. 283, 20161376 (2016).
Johnson, M. S., Martsul, A., Kryazhimskiy, S. & Desai, M. M. Higher-fitness yeast genotypes are less robust to deleterious mutations. Science 366, 490–493 (2019).
Wei, X. & Zhang, J. Patterns and mechanisms of diminishing returns from beneficial mutations. Mol. Biol. Evol. 36, 1008–1021 (2019).
Bakerlee, C. W., Ba, A. N. N., Shulgina, Y., Echenique, J. I. R. & Desai, M. M. Idiosyncratic epistasis leads to global fitness–correlated trends. Science 376, 630–635 (2022).
Johnson, M. S., Reddy, G. & Desai, M. M. Epistasis and evolution: recent advances and an outlook for prediction. BMC Biol. 21, 120 (2023).
Chou, H.-H., Berthet, J. & Marx, C. J. Fast growth increases the selective advantage of a mutation arising recurrently during evolution under metal limitation. PLoS Genet. 5, e1000652 (2009).
Otwinowski, J., McCandlish, D. M. & Plotkin, J. B. Inferring the shape of global epistasis. Proc. Natl Acad. Sci. 115, E7550–E7558 (2018).
Greene, D. & Crona, K. The changing geometry of a fitness landscape along an adaptive walk. PLoS Comput. Biol. 10, e1003520 (2014).
Blanquart, F., Achaz, G., Bataillon, T. & Tenaillon, O. Properties of selected mutations and genotypic landscapes under fisher’s geometric model. Evolution 68, 3537–3554 (2014).
Romero, P. A., Krause, A. & Arnold, F. H. Navigating the protein fitness landscape with gaussian processes. Proc. Natl Acad. Sci. 110, E193–E201 (2013).
Tonner, P. D., Pressman, A. & Ross, D. Interpretable modeling of genotype–phenotype landscapes with state-of-the-art predictive power. Proc. Natl Acad. Sci. 119, e2114021119 (2022).
Tareen, A. et al. Mave-nn: learning genotype-phenotype maps from multiplex assays of variant effect. Genome Biol. 23, 98 (2022).
Turner, P. E. & Elena, S. F. Cost of host radiation in an rna virus. Genetics 156, 1465–1470 (2000).
Bergland, A. O., Behrman, E. L., O’Brien, K. R., Schmidt, P. S. & Petrov, D. A. Genomic evidence of rapid and stable adaptive oscillations over seasonal time scales in drosophila. PLoS Genet. 10, e1004775 (2014).
Rodríguez-Verdugo, A., Carrillo-Cisneros, D., González-González, A., Gaut, B. S. & Bennett, A. F. Different tradeoffs result from alternate genetic adaptations to a common environment. Proc. Natl Acad. Sci. 111, 12121–12126 (2014).
Cvijović, I., Good, B. H., Jerison, E. R. & Desai, M. M. Fate of a mutation in a fluctuating environment. Proc. Natl Acad. Sci. 112, E5021–E5028 (2015).
You, L. & Yin, J. Dependence of epistasis on environment and mutation severity as revealed by in silico mutagenesis of phage t7. Genetics 160, 1273–1281 (2002).
Flynn, K. M., Cooper, T. F., Moore, F. B. & Cooper, V. S. The environment affects epistatic interactions to alter the topology of an empirical fitness landscape. PLoS Genet. 9, e1003426 (2013).
Lindsey, H. A., Gallie, J., Taylor, S. & Kerr, B. Evolutionary rescue from extinction is contingent on a lower rate of environmental change. Nature 494, 463–467 (2013).
Hall, A. E. et al. Environment changes epistasis to alter trade-offs along alternative evolutionary paths. Evolution 73, 2094–2105 (2019).
Ogbunugafor, C. B. The mutation effect reaction norm (mu-rn) highlights environmentally dependent mutation effects and epistatic interactions. Evolution 76, 37–48 (2022).
Ghenu, A.-H., Amado, A., Gordo, I. & Bank, C. Epistasis decreases with increasing antibiotic pressure but not temperature. Philos. Trans. Royal Soc. B 378, 20220058 (2023).
Oomen, R. A. & Hutchings, J. A. Genetic variability in reaction norms in fishes. Environ. Rev. 23, 353–366 (2015).
Ogbunugafor, C. B., Wylie, C. S., Diakite, I., Weinreich, D. M. & Hartl, D. L. Adaptive landscape by environment interactions dictate evolutionary dynamics in models of drug resistance. PLoS Comput. Biol. 12, e1004710 (2016).
Bajić, D., Vila, J. C. C., Blount, Z. D. & Sánchez, A. On the deformability of an empirical fitness landscape by microbial evolution. Proc. Natl Acad. Sci. 115, 11286–11291 (2018).
Mustonen, V. & Lässig, M. From fitness landscapes to seascapes: non-equilibrium dynamics of selection and adaptation. Trends Genet. 25, 111–119 (2009).
Iram, S. et al. Controlling the speed and trajectory of evolution with counterdiabatic driving. Nat. Phys. 17, 135–142 (2021).
Palmer, A. C. & Kishony, R. Understanding, predicting and manipulating the genotypic evolution of antibiotic resistance. Nat. Rev. Genet. 14, 243–248 (2013).
Berendonk, T. U. et al. Tackling antibiotic resistance: the environmental framework. Nat. Rev. Microbiol. 13, 310–317 (2015).
Bengtsson-Palme, J., Kristiansson, E. & Larsson, D. J. Environmental factors influencing the development and spread of antibiotic resistance. FEMS Microbiol. Rev. 42, fux053 (2018).
Ogbunugafor, C. B. & Eppstein, M. J. Competition along trajectories governs adaptation rates towards antimicrobial resistance. Nat. Ecol. Evol. 1, 1–8 (2016).
Tahar, R. & Basco, L. K. Molecular epidemiology of malaria in cameroon. xxii. geographic mapping and distribution of Plasmodium falciparum dihydrofolate reductase (DHFR) mutant alleles. Am. J. Trop. Med. Hygiene 75, 396–401 (2006).
Gebru-Woldearegai, T., Hailu, A., Grobusch, M. P. & Kun, J. F. Molecular surveillance of mutations in dihydrofolate reductase and dihyropteroate synthase genes of Plasmodium falciparum in ethiopia. Am. J. Trop. Med. Hygiene 73, 1131–1134 (2005).
Heidari, A. et al. Genotypes and in vivo resistance of Plasmodium falciparum isolates in an endemic region of iran. Parasitol.Res. 100, 589–592 (2007).
Ahmed, A. et al. Quadruple mutations in dihydrofolate reductase of Plasmodium falciparum isolates from Car Nicobar Island, India. Antimicrobial Agents Chemother. 50, 1546–1549 (2006).
Porse, A., Jahn, L. J., Ellabaan, M. M. & Sommer, M. O. Dominant resistance and negative epistasis can limit the co-selection of de novo resistance mutations and antibiotic resistance genes. Nat. Commun. 11, 1199 (2020).
Wong, A. Epistasis and the evolution of antimicrobial resistance. Front. Microbiol. 246 https://doi.org/10.3389/fmicb.2017.00246 (2017).
Reddy, G. & Desai, M. M. Global epistasis emerges from a generic model of a complex trait. eLife 10, e64740 (2021).
Diaz-Colunga, J. et al. Global epistasis on fitness landscapes. Philos. Trans. Royal Soc. B 378, 20220053 (2023).
Wiser, M. J., Ribeck, N. & Lenski, R. E. Long-term dynamics of adaptation in asexual populations. Science 342, 1364–1367 (2013).
Lenski, R. E. et al. Sustained fitness gains and variability in fitness trajectories in the long-term evolution experiment with escherichia coli. Proc. Royal Soc. B: Biol. Sci. 282, 20152292 (2015).
Lyons, D. M., Zou, Z., Xu, H. & Zhang, J. Idiosyncratic epistasis creates universals in mutational effects and evolutionary trajectories. Nat. Ecol. Evol. 4, 1685–1693 (2020).
Berger, D. & Postma, E. Biased estimates of diminishing-returns epistasis? empirical evidence revisited. Genetics 198, 1417–1420 (2014).
Diaz-Colunga, J., Sanchez, A. & Ogbunugafor, C. B. Environmental modulation of global epistasis in a drug resistance fitness landscape. Zenodo https://zenodo.org/doi/10.5281/zenodo.10067162 (2023).
Acknowledgements
We thank members of the Sanchez and Ogbunu labs for helpful discussions. C.B.O. acknowledges support from National Science Foundation’s Division of Environmental Biology Award Number 2142720 and from the MLK Visiting Scholars and Professors Program at the Massachusetts Institute of Technology. J.D.-C. & A.S. were partially supported by grant PID2021-125478NA-100 funded by MCIN/AEI/10.13039/501100011033 and by “ERDF: A way of making Europe.”
Author information
Authors and Affiliations
Contributions
J.D.-C., A.S. and C.B.O. conceived the study. J.D.-C. analyzed and interpreted data. J.D.-C. wrote the paper, with input from A.S. and C.B.O.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Jacob Scott and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Diaz-Colunga, J., Sanchez, A. & Ogbunugafor, C.B. Environmental modulation of global epistasis in a drug resistance fitness landscape. Nat Commun 14, 8055 (2023). https://doi.org/10.1038/s41467-023-43806-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41467-023-43806-x
This article is cited by
-
Global epistasis in plasmid-mediated antimicrobial resistance
Molecular Systems Biology (2024)
-
Environmental modulation of global epistasis in a drug resistance fitness landscape
Nature Communications (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.