Abstract
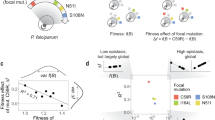
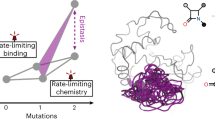
Fitness landscapes, mappings of genotype/phenotype to their effects on fitness, are invaluable concepts in evolutionary biochemistry. Although widely discussed, measurements of phenotype–fitness landscapes in proteins remain scarce. Here, we quantify all single mutational effects on fitness and phenotype (EC50) of VIM-2 β-lactamase across a 64-fold range of ampicillin concentrations. We then construct a phenotype–fitness landscape that takes variations in environmental selection pressure into account. We found that a simple, empirical landscape accurately models the ~39,000 mutational data points, suggesting that the evolution of VIM-2 can be predicted on the basis of the selection environment. Our landscape provides new quantitative knowledge on the evolution of the β-lactamases and proteins in general, particularly their evolutionary dynamics under subinhibitory antibiotic concentrations, as well as the mechanisms and environmental dependence of non-specific epistasis.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
Sequencing data are available through the NCBI SRA, under BioProject accession PRJNA634597. All data generated or analysed during this study are included in this published article and its Supplementary Information.
Code availability
All code for analysis are publicly available. ‘DMS-FastQ-processing’ (v.1) can be found at https://github.com/johnchen93/DMS-FastQ-processing. ‘DMS-EC50’ (v.1) can be found at https://github.com/johnchen93/DMS-EC50.
References
Dalziel, A. C., Rogers, S. M. & Schulte, P. M. Linking genotypes to phenotypes and fitness: how mechanistic biology can inform molecular ecology. Mol. Ecol. 18, 4997–5017 (2009).
Kaltenbach, M. & Tokuriki, N. Dynamics and constraints of enzyme evolution. J. Exp. Zool. B 322, 468–487 (2014).
Yi, X. & Dean, A. M. Adaptive landscapes in the age of synthetic biology. Mol. Biol. Evol. 36, 890–907 (2019).
Kemble, H., Nghe, P. & Tenaillon, O. Recent insights into the genotype–phenotype relationship from massively parallel genetic assays. Evol. Appl. 12, 1721–1742 (2019).
Bershtein, S., Serohijos, A. W. & Shakhnovich, E. I. Bridging the physical scales in evolutionary biology: from protein sequence space to fitness of organisms and populations. Curr. Opin. Struct. Biol. 42, 31–40 (2017).
DePristo, M. A., Weinreich, D. M. & Hartl, D. L. Missense meanderings in sequence space: a biophysical view of protein evolution. Nat. Rev. Genet. 6, 678 (2005).
Pál, C., Papp, B. & Lercher, M. J. An integrated view of protein evolution. Nat. Rev. Genet. 7, 337–348 (2006).
Rodrigues, J. V. et al. Biophysical principles predict fitness landscapes of drug resistance. Proc. Natl Acad. Sci. USA 113, E1470–1478 (2016).
Hartl, D. L., Dykhuizen, D. E. & Dean, A. M. Limits of adaptation: the evolution of selective neutrality. Genetics 111, 655–674 (1985).
Lunzer, M., Miller, S. P., Felsheim, R. & Dean, A. M. The biochemical architecture of an ancient adaptive landscape. Science 310, 499–501 (2005).
Lundin, E., Tang, P.-C., Guy, L., Näsvall, J. & Andersson, D. I. Experimental determination and prediction of the fitness effects of random point mutations in the biosynthetic enzyme HisA. Mol. Biol. Evol. 35, 704–718 (2018).
Tokuriki, N. & Tawfik, D. S. Stability effects of mutations and protein evolvability. Curr. Opin. Struct. Biol. 19, 596–604 (2009).
Sarkisyan, K. S. et al. Local fitness landscape of the green fluorescent protein. Nature 533, 397–401 (2016).
Dasmeh, P. & Serohijos, A. W. R. Estimating the contribution of folding stability to nonspecific epistasis in protein evolution. Proteins Struct. Funct. Bioinforma. 86, 1242–1250 (2018).
Bershtein, S., Segal, M., Bekerman, R., Tokuriki, N. & Tawfik, D. S. Robustness–epistasis link shapes the fitness landscape of a randomly drifting protein. Nature 444, 929–932 (2006).
Pokusaeva, V. O. et al. An experimental assay of the interactions of amino acids from orthologous sequences shaping a complex fitness landscape. PLoS Genet. 15, e1008079 (2019).
Diss, G. & Lehner, B. The genetic landscape of a physical interaction. eLife 7, e32472 (2018).
Wylie, C. S. & Shakhnovich, E. I. A biophysical protein folding model accounts for most mutational fitness effects in viruses. Proc. Natl Acad. Sci. USA 108, 9916–9921 (2011).
Chou, H.-H., Delaney, N. F., Draghi, J. A. & Marx, C. J. Mapping the fitness landscape of gene expression uncovers the cause of antagonism and sign epistasis between adaptive mutations. PLoS Genet. 10, e1004149 (2014).
Bershtein, S., Mu, W., Serohijos, A. W. R., Zhou, J. & Shakhnovich, E. I. Protein quality control acts on folding intermediates to shape the effects of mutations on organismal fitness. Mol. Cell 49, 133–144 (2013).
Stiffler, M. A., Hekstra, D. R. & Ranganathan, R. Evolvability as a function of purifying selection in TEM-1 β-lactamase. Cell 160, 882–892 (2015).
Mehlhoff, J. D. et al. Collateral fitness effects of mutations. Proc. Natl Acad. Sci. USA 117, 11597–11607 (2020).
Dean, A. M. A molecular investigation of genotype by environment interactions. Genetics 139, 19–33 (1995).
Otwinowski, J., McCandlish, D. M. & Plotkin, J. B. Inferring the shape of global epistasis. Proc. Natl Acad. Sci. USA 115, E7550–E7558 (2018).
Fowler, D. M. et al. High-resolution mapping of protein sequence–function relationships. Nat. Methods 7, 741–746 (2010).
Fowler, D. M., Stephany, J. J. & Fields, S. Measuring the activity of protein variants on a large scale using deep mutational scanning. Nat. Protoc. 9, 2267–2284 (2014).
Melnikov, A., Rogov, P., Wang, L., Gnirke, A. & Mikkelsen, T. S. Comprehensive mutational scanning of a kinase in vivo reveals substrate-dependent fitness landscapes. Nucleic Acids Res. 42, e112 (2014).
Wrenbeck, E. E., Azouz, L. R. & Whitehead, T. A. Single-mutation fitness landscapes for an enzyme on multiple substrates reveal specificity is globally encoded. Nat. Commun. 8, 15695 (2017).
Mavor, D. et al. Determination of ubiquitin fitness landscapes under different chemical stresses in a classroom setting. eLife 5, e15802 (2016).
Noda-García, L. et al. Chance and pleiotropy dominate genetic diversity in complex bacterial environments. Nat. Microbiol. 4, 1221–1230 (2019).
Starr, T. N., Picton, L. K. & Thornton, J. W. Alternative evolutionary histories in the sequence space of an ancient protein. Nature 549, 409–413 (2017).
Starr, T. N. et al. Deep mutational scanning of SARS-CoV-2 receptor binding domain reveals constraints on folding and ACE2 binding. Cell 182, 1295–1310 (2020).
Hietpas, R. T., Jensen, J. D. & Bolon, D. N. A. Experimental illumination of a fitness landscape. Proc. Natl Acad. Sci. USA 108, 7896–7901 (2011).
Firnberg, E., Labonte, J. W., Gray, J. J. & Ostermeier, M. A comprehensive, high-resolution map of a gene’s fitness landscape. Mol. Biol. Evol. 31, 1581–1592 (2014).
Jacquier, H. et al. Capturing the mutational landscape of the beta-lactamase TEM-1. Proc. Natl Acad. Sci. USA 110, 13067–13072 (2013).
Klesmith, J. R., Bacik, J.-P., Wrenbeck, E. E., Michalczyk, R. & Whitehead, T. A. Trade-offs between enzyme fitness and solubility illuminated by deep mutational scanning. Proc. Natl Acad. Sci. USA 114, 2265–2270 (2017).
Chen, J. Z., Fowler, D. M. & Tokuriki, N. Comprehensive exploration of the translocation, stability and substrate recognition requirements in VIM-2 lactamase. eLife 9, e56707 (2020).
Meer, J. et al. Using mutability landscapes of a promiscuous tautomerase to guide the engineering of enantioselective Michaelases. Nat. Commun. 7, 10911 (2016).
Thompson, S., Zhang, Y., Ingle, C., Reynolds, K. A. & Kortemme, T. Altered expression of a quality control protease in E. coli reshapes the in vivo mutational landscape of a model enzyme. eLife 9, e53476 (2020).
Kemble, H. et al. Flux, toxicity, and expression costs generate complex genetic interactions in a metabolic pathway. Sci. Adv. 6, eabb2236 (2020).
Starita, L. M. et al. Activity-enhancing mutations in an E3 ubiquitin ligase identified by high-throughput mutagenesis. Proc. Natl Acad. Sci. USA 110, E1263–E1272 (2013).
Flynn, J. M. et al. Comprehensive fitness maps of Hsp90 show widespread environmental dependence. eLife 9, e53810 (2020).
Horovitz, A., Fleisher, R. C. & Mondal, T. Double-mutant cycles: new directions and applications. Curr. Opin. Struct. Biol. 58, 10–17 (2019).
Salinas, V. H. & Ranganathan, R. Coevolution-based inference of amino acid interactions underlying protein function. eLife 7, e34300 (2018).
Mavor, D. et al. Extending chemical perturbations of the ubiquitin fitness landscape in a classroom setting reveals new constraints on sequence tolerance. Biol.Open 7, bio036103 (2018).
Rockah-Shmuel, L., Tóth-Petróczy, Á. & Tawfik, D. S. Systematic mapping of protein mutational space by prolonged drift reveals the deleterious effects of seemingly neutral mutations. PLoS Comput. Biol. 11, e1004421 (2015).
Gray, V. E., Hause, R. J., Luebeck, J., Shendure, J. & Fowler, D. M. Quantitative missense variant effect prediction using large-scale mutagenesis data. Cell Syst. 6, 116–124 (2018).
Gullberg, E. et al. Selection of resistant bacteria at very low antibiotic concentrations. PLoS Pathog. 7, e1002158 (2011).
Westhoff, S. et al. The evolution of no-cost resistance at sub-MIC concentrations of streptomycin in Streptomyces coelicolour. ISME J. 11, 1168–1178 (2017).
Liu, A. et al. Selective advantage of resistant strains at trace levels of antibiotics: a simple and ultrasensitive colour test for detection of antibiotics and genotoxic agents. Antimicrob. Agents Chemother. 55, 1204–1210 (2011).
Jasinska, W. et al. Chromosomal barcoding of E. coli populations reveals lineage diversity dynamics at high resolution. Nat. Ecol. Evol. 4, 437–452 (2020).
Fröhlich, C. et al. Cryptic β-lactamase evolution is driven by low β-lactam concentrations. mSphere https://doi.org/10.1128/mSphere.00108-21 (2021).
Andersson, D. I. & Hughes, D. Microbiological effects of sublethal levels of antibiotics. Nat. Rev. Microbiol. 12, 465–478 (2014).
Baquero, F., Negri, M. C., Morosini, M. I. & Blázquez, J. Selection of very small differences in bacterial evolution. Int. Microbiol. 1, 295–300 (1998).
Drlica, K. & Zhao, X. Mutant selection window hypothesis updated. Clin. Infect. Dis. 44, 681–688 (2007).
Blondeau, J. M. New concepts in antimicrobial susceptibility testing: the mutant prevention concentration and mutant selection window approach. Vet. Dermatol. 20, 383–396 (2009).
Das, S. G., Direito, S. O., Waclaw, B., Allen, R. J. & Krug, J. Predictable properties of fitness landscapes induced by adaptational tradeoffs. eLife 9, e55155 (2020).
Starr, T. N. & Thornton, J. W. Epistasis in protein evolution. Protein Sci. 25, 1204–1218 (2016).
Domingo, J., Baeza-Centurion, P. & Lehner, B. The causes and consequences of genetic interactions (epistasis). Annu. Rev. Genomics Hum. Genet. 20, 433–460 (2019).
Yang, G. et al. Higher-order epistasis shapes the fitness landscape of a xenobiotic-degrading enzyme. Nat. Chem. Biol. 15, 1120–1128 (2019).
Nishikawa, K. K., Hoppe, N., Smith, R., Bingman, C. & Raman, S. Epistasis shapes the fitness landscape of an allosteric specificity switch. Nat. Commun. 12, 5562 (2021).
Baier, F. et al. Cryptic genetic variation shapes the adaptive evolutionary potential of enzymes. eLife 8, e40789 (2019).
Schenk, M. F., Szendro, I. G., Salverda, M. L. M., Krug, J. & de Visser, J. A. G. M. Patterns of epistasis between beneficial mutations in an antibiotic resistance gene. Mol. Biol. Evol. 30, 1779–1787 (2013).
Bank, C., Hietpas, R. T., Jensen, J. D. & Bolon, D. N. A. A systematic survey of an intragenic epistatic landscape. Mol. Biol. Evol. 32, 229–238 (2015).
Manrubia, S. et al. From genotypes to organisms: state-of-the-art and perspectives of a cornerstone in evolutionary dynamics. Phys. Life Rev. 38, 55–106 (2021).
Araya, C. L. et al. A fundamental protein property, thermodynamic stability, revealed solely from large-scale measurements of protein function. Proc. Natl Acad. Sci. USA 109, 16858–16863 (2012).
Olson, C. A., Wu, N. C. & Sun, R. A comprehensive biophysical description of pairwise epistasis throughout an entire protein domain. Curr. Biol. 24, 2643–2651 (2014).
Palmer, A. C. et al. Delayed commitment to evolutionary fate in antibiotic resistance fitness landscapes. Nat. Commun. 6, 7385 (2015).
Miton, C. M. & Tokuriki, N. How mutational epistasis impairs predictability in protein evolution and design. Protein Sci. 25, 1260–1272 (2016).
Russ, D. et al. Escape mutations circumvent a tradeoff between resistance to a beta-lactam and resistance to a beta-lactamase inhibitor. Nat. Commun. 11, 2029 (2020).
Tack, D. S. et al. The genotype–phenotype landscape of an allosteric protein. Mol. Syst. Biol. 17, e10847 (2021).
Steinberg, B. & Ostermeier, M. Environmental changes bridge evolutionary valleys. Sci. Adv. 2, e1500921 (2016).
Zwart, M. P. et al. Unraveling the causes of adaptive benefits of synonymous mutations in TEM-1 β-lactamase. Heredity 121, 406–421 (2018).
Edgar, R. C. & Flyvbjerg, H. Error filtering, pair assembly and error correction for next-generation sequencing reads. Bioinformatics 31, 3476–3482 (2015).
Sebaugh, J. L. & McCray, P. D. Defining the linear portion of a sigmoid-shaped curve: bend points. Pharm. Stat. 2, 167–174 (2003).
Acknowledgements
We thank A. Serohijos, P. Dasmeh and members of the Tokuriki laboratory for insightful comments on the manuscript. We thank the Canadian Institute of Health Research Foundation Grant (FDN-148437, N.T.) for the financial support. N.T. is a Michael Smith Foundation of Health Research career investigator.
Author information
Authors and Affiliations
Contributions
J.Z.C. was responsible for conceptualization, software, formal analysis, investigation, methodology, writing the original draft and reviewing and editing it. D.M.F. contributed to software, validation, methodology and reviewed and edited the manuscript. N.T. was involved in conceptualization, supervision, funding acquisition, project administration and reviewing and editing of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Ecology & Evolution thanks the anonymous reviewers for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Example dose–response curves for EC50 calculation based on deep sequencing data.
Each pair of plots represent the two replicate dose–response curves of a single variant in the library. The solid curve indicates the results of fitting to the sigmoidal equation (2). The horizontal dashed line marks the average population size of the variant in the non-selected conditions. Dose–response curves shown here were randomly sampled from the entire pool of all variants with a successful fit.
Extended Data Fig. 2 Heat map of EC50 values for all variants.
Each cell in the heat map represents the EC50 value of a single amino acid variant. Synonymous variants are indicated by dark grey circles and variants that are not present in the library are in grey. The x-axis under the heat map indicates the wild-type residue and position, while the y-axis indicates the variant residue at that position. The colouring is scaled so that the centre of the colour range (white) is at the EC50-wt of 84 µg/mL.
Extended Data Fig. 3 Replicate correlation of DMS fitness scores.
Each scatterplot shows the replicate fitness scores of all variants under selection in (a) 4.0, (b) 8.0, (c) 16, (d) 32, (e) 64, (f) 128 and (g) 256 µg/mL AMP. In each panel, variants are coloured according to mutation type with the legend in the lower right. The solid black line indicate the line of best fit for a linear regression, with the R2 and P value of the regression above each plot.
Extended Data Fig. 4 Fit parameters for the phenotype–fitness relationship across AMP concentrations.
Each plot shows one of the final fitted value of a parameter in the four-parameter sigmoidal curve in equation (3) when fitted to the fitness scores and EC50 values from the AMP concentration indicated in the x-axis. (a) Plot for maximum fitness (fmax) (b) Plot for minimum fitness (fmin), (c) Plot for Hill coefficient (n) (d) Plot for the inflection point (X50). For all plots, solid points represent AMP concentrations where the linear portion of the fit were not limited by the AMP concentration range, while hollow points indicate the opposite. For d), a linear regression was conducted with the solid data points. The black line indicates the line of best fit for a linear regression, and the R2, P value and the equation of the fit shown at the top.
Extended Data Fig. 5 Fitted parameters of sigmoidal phenotype–fitness relationships.
a All parameters shown are calculated from a fit to equation (3). b Values are displayed as optimal fit ± S.D. The S.D.s are calculated as the square root of the variance for the parameter, estimated during fitting by the ‘SciPy.optimize.cuve_fit’ function.
Supplementary information
Supplementary Data
Supplementary Data 1, Fitness scores of VIM-2 variants across AMP concentrations; Data 2, EC50 of all VIM-2 variants; Data 3, EC50 of VIM-2 variants measured individually; Data 4, OD600 of VIM-2 libraries after 6 h of selection at 37 °C.
Rights and permissions
About this article
Cite this article
Chen, J.Z., Fowler, D.M. & Tokuriki, N. Environmental selection and epistasis in an empirical phenotype–environment–fitness landscape. Nat Ecol Evol 6, 427–438 (2022). https://doi.org/10.1038/s41559-022-01675-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41559-022-01675-5
This article is cited by
-
Understanding activity-stability tradeoffs in biocatalysts by enzyme proximity sequencing
Nature Communications (2024)
-
Rapid evolutionary change in trait correlations of single proteins
Nature Communications (2024)