Abstract
Following postglacial expansion, secondary contact can occur between genetically distinct lineages. These genetic lineages may be associated with specific habitat or environmental variables and therefore, their distributions in secondary contact could reflect such conditions within these areas. Here we used mtDNA, microsatellite, and morphological data to study three genetically distinct groups of warbling vireo (Vireo gilvus) and investigate the role that elevation and habitat play in their distributions. We studied two main contact zones and within each contact zone, we examined two separate transects. Across the Great Plains contact zone, we found that hybridization between eastern and western groups occurs along a habitat and elevational gradient, whereas hybridization across the Rocky Mountain contact zone was not as closely associated with habitat or elevation. Hybrids in the Great Plains contact zone were more common in transitional areas between deciduous and mixed-wood forests, and at lower elevations (<1000 m). Hybridization patterns were similar along both Great Plains transects indicating that habitat and elevation play a role in hybridization between distinct eastern and western genetic groups. The observed patterns suggest adaptation to different habitats, perhaps originating during isolation in multiple Pleistocene refugia, is facilitating hybridization in areas where habitat types overlap.
Similar content being viewed by others
Introduction
Our understanding of the processes involved in divergence and speciation is constantly developing (Toews et al. 2016). Hybrid zones are of particular interest as genetically differentiated populations come into contact and interbreed allowing us to study a range of traits between taxa at various stages of speciation (Barton and Hewitt 1989; Barton and Gale 1993; Brelsford and Irwin 2009; Baldassarre et al. 2014). In North America, a number of physiographic features correspond to hybrid zone hotspots across taxonomic groups (Swenson and Howard 2005). Hewitt (2001) proposed that some of these were located between or at the edges of ice sheets during the Pleistocene (2.5 Mya), with isolated populations later coming into secondary contact after postglacial recolonization. The effects of past and present day climatic factors (e.g., precipitation, temperature) and the location of these hybrid zones have been well documented (Mettler and Spellman 2009; Carling and Zuckerberg 2011; Schukman et al. 2011; Walsh et al. 2020).
Hybrid zones are maintained through a number of mechanisms including extrinsic (e.g., environment) and intrinsic (pre- and post-zygotic reproductive barriers) factors, hybrid related fitness, and selection (Mettler and Spellman 2009; Billerman et al. 2016; Irwin 2020). Studying how extrinsic factors, like environment and habitat, influence hybridization is important given that habitat modifications and climate change influence hybridization (Taylor et al. 2015; Grabenstein and Taylor 2018). The frequency of hybridization events increases as landscape heterogeneity decreases, especially in modified landscapes, resulting in hybridization of ecologically distinct species that would not hybridize under normal conditions (Seehausen et al. 2008; Grabenstein and Taylor 2018; Sartor et al. 2021). Furthermore, climate change induced range expansion can increase the level of sympatry between species and populations, and lead to greater rates of hybridization (Garroway et al. 2010; Larson et al. 2019). Therefore, examining hybridization patterns along different environmental gradients, such as the transition from montane habitat in the Rocky Mountains to grassland habitat found on the Great Plains, can provide greater insight into the relationship between environment and hybridization. These transitional areas, called ecotones, act as hybrid zones for divergent taxa across North America (Walsh et al. 2016; Reding et al. 2021) and elsewhere (Culumber et al. 2012; Pavolova et al. 2013). Such hybrid zones are “natural laboratories” (Hewitt 1988) to explore how environmental variables may have influenced the initial divergence of these taxa, and how their adaptation to different environmental conditions then influences hybridization. Cryptic species, those species that are genetically distinct but exhibit low phenotypic or morphological divergence (Toews and Irwin 2008; Rush et al. 2009; Bradbury et al. 2014; Hinojosa et al. 2019), offer an exceptional opportunity to examine the impact of habitat and environment on reproductive isolation and hybridization (Coyne and Orr 2004; Chenuil et al. 2019). The absence of distinct morphological differences leads to questions about whether reproductive isolation is complete between genetically distinct groups, or whether interbreeding occurs once barriers to dispersal are removed (Chenuil et al. 2019). Further, questions remain about whether genetic differentiation is associated with environmental characteristics or niche divergence (MacDonald et al. 2020).
In this study, we assess the relationship between habitat, morphology, genetic variation, and patterns of hybridization in three genetically distinct groups of warbling vireo (Vireo gilvus) and their secondary contact zones in western North America (Fig. 1). Three of the five subspecies of warbling vireo are found in this area, including the eastern (Vireo gilvus gilvus) and two western (V. g. swainsonii and V. g. brewsteri) subspecies that diverged during the early Pleistocene (Hebert et al. 2004; Lovell et al. 2021; Carpenter et al. 2021) with hybridization occurring along the Rocky Mountain—Great Plains ecotone (Lovell et al. 2021; Carpenter et al. 2021). Hybridization between eastern and western populations in central Alberta may be facilitated by habitat, as the location of the narrow hybrid zone corresponds to a transition between deciduous and mixed forests (Lovell et al. 2021). Further to the west, Carpenter et al. (2021) revealed genetic differentiation among western populations with northwestern and southwestern groups, but whether these genetic differences are associated with habitat or elevation remains unknown. Here we test whether hybrids are more prominent in specific forest types, and whether elevation and habitat differences influence hybridization. Based on our previous study of warbling vireo phylogeographic patterns (Carpenter et al. 2021), we believe that genetic variation may also be associated with elevation differences, in addition to the habitat differences shown by Lovell et al. (2021). We examine the effects of both elevation and habitat to determine whether hybridization patterns are more closely associated with one variable or both. We also examined whether hybridization patterns differed along each of the contact zones by examining two transects within two different areas of secondary contact. Examining multiple transects allowed us to better determine how prominent hybridization is among genetically distinct groups, and ascertain how hybridization may vary across these large areas of secondary contact.
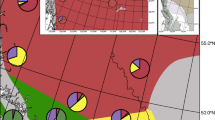
Map showing the distribution of the three warbling vireo genetic groups based on Carpenter et al. (2021) in areas of secondary contact in western North America.
Methods
Contact zones
Genetic patterns were analyzed across multiple contact zone transects (Fig. 2). The Great Plains contact zone in Alberta, Montana, and Saskatchewan is where divergent eastern and western warbling vireo genetic lineages come into contact (Lovell et al. 2021; Carpenter et al. 2021). At the northern edge of the Great Plains in central Alberta, deciduous forest transitions into coniferous forest (Semenchuk 1992). Forest coverage is higher, and elevation is lower than at the southern edge of Alberta and northern Montana where deciduous forest is sparsely distributed among grasslands. A drastic increase in elevation occurs at the southwestern edge of the Great Plains contact zone with a transition to the Rocky Mountains, and where deciduous habitat is replaced by mixed and coniferous forests. Given how variable habitat and elevation features are across the Great Plains contact zone, we examined two separate transects (Fig. 2A, B) to determine the effect of habitat and elevation on genetic and morphological variation. Both of the transects in the Great Plains ran east–west to identify the areas where eastern genotypes are replaced by western genotypes.
In the western contact zone of the Rocky Mountains, secondary contact occurs between northwestern and southwestern genetic groups. Genetic differentiation is reduced relative to genetic differences between eastern and western birds (Carpenter et al. 2021). Coniferous and mixed forests are the dominant habitats, with deciduous forest found at lower elevations. Elevation changes continuously across the west, with the highest peaks found in the southwestern United States. Because of the considerable variation in habitat and elevation, we also analyzed two separate transects in the Rocky Mountain contact zone. One transect included birds from the two western genetic groups sampled west of the Rocky Mountains in British Columbia, Washington, Oregon, California, Idaho, and Nevada (Fig. 2C) and the other transect included all remaining samples from the two western genetic groups found east of the Rocky Mountains in Utah, Colorado, Wyoming, Montana, and Alberta (Fig. 2D). The Rocky Mountain contact zone included 203 individuals that were used in the Great Plains contact zone identified as having western genotypes. For the Rocky Mountain contact zone, we addressed similar questions of how genetic ancestry between the two western genetic groups changed across the two transects and whether hybridization was correlated with habitat or elevation differences. Our two Rocky Mountain transects were linear, and ran north to south on either side of the Rocky Mountains.
Sample collection
From 2009 to 2019, we collected samples from 544 individuals within the study area. We captured 355 adult individuals during the breeding season, May–July, using mistnets and song playbacks. Each individual was banded to avoid resampling, a small blood sample (<50 µL) was taken from the brachial wing vein, and mass (g) and uncompressed wing chord length measured to the nearest 0.1 mm were recorded. An additional 189 tissue samples, some with corresponding morphological data, were obtained from museum collections to supplement our field sampling (Table 1). Samples were stored in 95% ethanol and DNA was extracted using a modified Chelex procedure (Walsh et al. 1991; Burg and Croxall 2004).
DNA amplification
MtDNA
We developed primers targeting fixed nucleotide differences found in the cytochrome b (cyt b) gene in six eastern (gilvus) and six western (swainsonii) warbling vireo sequences (Carpenter et al. 2021) (GenBank accession numbers: MZ20223-MZ20225, MZ20227-MZ20229, MZ20254-MZ20256, MZ20258-MZ20260). Paired with the L14990 primer (Sorenson et al. 1999), we created the H15469 Gilvus primer (5′ ACGAAGGGTAGTAGCAAA 3′) and H15634 West primer (5′ GAGAATAGGGCTAGGTG 3′) to assign 192 individuals to eastern or western mtDNA clades. The 10 µL polymerase chain reaction (PCR) used 5× Green GoTaq® Flexi buffer (Promega), 0.2 mM dNTP, 2 mM MgCl2 (Promega), 0.5 µM of each H (H15469 Gilvus or H15634 West) and L14990 primer, and 0.5 U GoTaq® Flexi polymerase (Promega). The following thermocycling profile was used to amplify DNA for both H15469 Gilvus and H15634 West primers: 2 min at 94 °C, 45 s at 48 °C, 1 min at 72 °C, 37 cycles of 30 s at 94 °C, 45 s at 48 °C, 1 min at 72 °C, ending with a 5 min extension at 72 °C. PCR products were run on a 0.8% agarose gel and identified based on size; H15469 Gilvus is ~550 bp and H15634 West is ~600 bp. A positive control for both eastern and western groups was run with each set of unknown samples for accuracy of identification. Furthermore, a subset of samples (N = 20) of both eastern and western birds was sent for sequencing at NanuQ (McGill University, Montreal, QC, Canada) to confirm the reliability of these primers.
Microsatellites
To examine the nuclear genome, we genotyped 544 individuals at 14 variable microsatellite loci: BCV2-6, BCV4-5, BCV4-6New (Barr et al. 2007); Ck.1B5D, Ck.4B6D, Ck.2A5A, Ck.1B6G (Tarr and Fleischer 1998); Hofi5 (Hawley 2005); Pocc1 (Bensch et al. 1997); ApCo46 (Stenzler and Fitzpatrick 2002); Lox1 (Piertney et al. 1998); Ase18 (Richardson et al. 2003); Ppi2New (Martinez et al. 1999); and CmAAAG30 (Williams et al. 2004). PCR protocols are outlined in Carpenter et al. (2021).
Genetic analyses
We examined nuclear ancestry across both the Great Plains and Rocky Mountain contact zones using the Bayesian clustering program STRUCTURE 2.3.4 (Pritchard et al. 2000; Hubisz et al. 2009). Previous analyses (Carpenter et al. 2021) indicate that populations near both contact zones each form two distinct genetic clusters; therefore, we examined K = 2. For the east–west contact zone in the Great Plains, we used the admixture model with correlated allele frequencies and no locpriors. For our analyses of the two western genetic groups in the Rocky Mountain contact zone, we used the no admixture model with correlated allele frequencies and sampling locations set as locpriors because levels of genetic differentiation were lower (Carpenter et al. 2021). The combination of these settings allows for the detection of distinct genetic clusters within closely-related populations and when hierarchal structure is weak (Hubisz et al. 2009; Porras-Hurtado et al. 2013). For both analyses, we used a burn-in of 50,000 followed by 100,000 Markov chain Monte Carlo runs.
Spatial distribution of hybrid zones
We mapped the spatial distribution of hybridization across the Rocky Mountain and Great Plains contact zones and quantified the area of each contact zone. We mapped points where hybrids (individuals with Q values >0.2 and <0.8) were captured in QGIS (version 10.13) and used the heat map function to identify hybridization density across these contact zones. To create heat maps, we used the default settings with a three-degree buffer radius and the scaled values output. We then used the raster calculator function to convert all areas with threshold values above three, so that all pixels above this threshold received a new value of one, whereas areas below this value received a value of zero. We chose this threshold based on the distribution of hybridization events across our contact zone; areas with a value above this threshold represent concentrated areas where there was extensive hybridization, whereas areas with values below this threshold represent areas with relatively few or no hybridization events. We converted the raster file to a vector file with the raster-to-vector conversion tool and used the area calculator to quantify the area covered by hotspots on the map for each contact zone.
Habitat measurements
We collected habitat and elevation data for 544 warbling vireos. Habitat data were extracted from the North American Land Monitoring System, and we classified habitats into one of four categories: non-forested (NF), deciduous forest (D), mixed deciduous and coniferous forest (M), and coniferous forest (C). Data for elevation were extracted from the World BioClim dataset (v.2.1; http://www.worldclim.org/) at 2.5 min resolution (Hijmans et al. 2005). We focused on habitat and elevational differences because the distribution of both western and eastern groups appears to be delimited by these two variables. For example, the eastern genetic group appears to be restricted to low elevation deciduous forests, whereas the western genetic groups inhabit high elevation coniferous forests (Carpenter et al. 2021).
Correlates with ancestry and habitat
First, we plotted the frequency of genotypes (mtDNA or microsatellite ancestry) against habitat and elevation to visualize genetic variation across the contact zone. Within the Great Plains contact zone, eastern mtDNA and nuclear genotypes were plotted against six elevational categories (≤600, ≤1000, ≤1400, ≤1800, ≤2200, ≥2600 m) and the four habitat categories. We repeated this for the Rocky Mountain contact zone, but plotted the frequency of northwestern nuclear genotypes.
We used two different modeling approaches to examine predictors of ancestry and genetic differentiation. First, we used Spearman’s rank correlation to examine the intercorrelations between ancestry, elevation, and habitat. We used this approach to assess the relationship between genetic variation and environmental variables because of the non-linear nature of these data. The first analysis included samples from the Great Plains contact zone (Montana, Alberta, and Saskatchewan) where eastern and western warbling vireo genetic groups come into contact. We performed this analysis on samples across the entire contact zone, as well as within the northern and southern transects (Fig. 2A, B). For the second analysis, we included all populations from the Rocky Mountain contact zone where the two western genetic groups come into contact. This was also conducted across the entire contact zone, as well as within the western and eastern Rocky Mountain transects (Fig. 2C, D). For all of the analyses, we used the ancestry coefficients (Q values) generated from STRUCTURE to quantify ancestry. We did not examine the correlation between mtDNA and habitat in the Rocky Mountain contact zone because of the low mtDNA genetic differentiation between the two western genetic groups (Carpenter et al. 2021). Lastly, we looked at the influence of latitude on the Rocky Mountain contact zone to account for the distribution of samples across a large geographic area. All analyses were conducted in Past 3.0 (Hammar et al. 2001) and correlations were considered significant at p ≤ 0.05.
For our second approach, we conducted distance-based and partial redundancy models to examine the relationship between the environment and genetic variation. This approach has been used in other studies (Riordan et al. 2016; Hindley et al. 2018) to examine predictors of genetic variation and is a more powerful option than Mantel tests (Legendre and Fortin 2010). Distance-based and partial redundancy models are multivariate statistical approaches that use canonical analyses to examine the effect of predictor variables on response variables. For these analyses, we calculated Sforza-Chord genetic distances between individuals based on our microsatellite datasets in Genodive 3.0 (Meirmans and Van Tienderen 2004). We examined the relationship between genetic distance and two variables (habitat and elevation) across the Great Plains contact zone, and three variables (habitat, elevation, and latitude) across the Rocky Mountain contact zone. In both sets of analyses, we included geographic distance as a variable to test the effect of isolation-by-distance on genetic distance. Isolation-by-distance is often viewed as the null hypothesis for genetic variance and divergence and including this in the analysis provided a reference to compare the effects of other predictor variables within the contact zones. Northern and southern transects were examined separately within the Great Plains contact zone, as well as the eastern Rocky Mountain and western Rocky Mountain transects within the Rocky Mountain contact zone. Finally, we examined mtDNA differentiation across the Great Plains contact zone using Nei’s genetic distance calculated in GenAlEx 6.5 (Peakall and Smouse 2012) for 192 individuals, and the effect of the same two variables (habitat and elevation) on the microsatellite data and isolation-by-distance on mitochondrial data.
Morphological variation
Carpenter et al. (2021) previously established that the eastern group is morphologically distinct from both western groups, but for the purpose of this study we wanted to investigate how morphology varies across the contact zones and the relationship between morphological, genetic, and environmental variation. To examine these patterns, we used Spearman’s correlation to test intercorrelations between morphological variation, habitat, elevation, and ancestry across both the Great Plains and Rocky Mountain contact zones. For the Rocky Mountain contact zone, we also tested the influence of latitude on morphological variation, as the western populations included in this study covered a much broader area.
Results
MtDNA screening
We assigned mtDNA ancestry for 192 individuals across 22 populations in the Great Plains contact zone. Two populations in central Alberta (Barrhead County, Parkland County/Edmonton) and one in southeastern Alberta (Cypress Hills) contained individuals from both eastern and western mtDNA clades, while the remaining 19 populations contained either western or eastern haplotypes only. In Barrhead County, 16 of the 24 individuals (67%) had eastern haplotypes, while the remaining eight individuals (33%) had western haplotypes. All but one of the 17 Cypress Hills birds contained western mtDNA and the opposite pattern was observed in the 14 individuals from Parkland County, with all but one individual grouping with the eastern mtDNA clade.
Microsatellites
Across the Great Plains contact zone, 9.2% (27 of 294) of individuals were identified as hybrids (Fig. 3A, B). Nine of the 27 hybrids had Q values between 0.75 and 0.80 indicative of advanced generation hybrids, and five other hybrids were possibly first generation hybrids (0.6 < Q < 0.4). Two populations contained individuals from both eastern and western genetic groups. In the first population, Barrhead County, 72% of individuals (18 of 25) had eastern genotypes, while the remaining 28% (7 of 25) had western genotypes. By comparison, in the second population in western Montana, 96.6% of individuals had western genotypes, and one (3.4%) had an eastern genotype. Hybridization rates were comparable between each of the northern and southern Great Plains transects; 9.7% of the birds (19 of 196) in the south and 8.2% (8 of 98) of individuals in the north were identified as hybrids.
Distribution of hybrids in the Great Plains contact zone across (A) elevational and (B) habitat gradients, and hybrids in the Rocky Mountain contact zone across (C) elevational and (D) habitat gradients. Triangles denote putative hybrids between eastern and western genetic lineages (A, B), while circles denote putative hybrids between the two western genetic lineages (C, D).
When we examined mtDNA and microsatellite patterns together for the Great Plains contact zone (N = 192), a small proportion of individuals (5 of 192 individuals, 2.6%) exhibited cytonuclear discordance; where the mtDNA genotype did not match with the nuclear genotype (e.g., individuals had eastern mtDNA, but grouped with the western genetic group based on microsatellite data). Three of these five individuals were from central Alberta, two sites that fall in the contact zone described by Lovell et al. (2021). The other two individuals with cytonuclear discordance were from Cypress Hills, AB and Helena, MT.
Across the Rocky Mountain contact zone, 36.4% (164 of 450) of individuals were identified as hybrids (Fig. 3C, D). Hybridization occurred extensively throughout the Rocky Mountains (Fig. 3); forty-one of the hybrids had Q values between 0.4 and 0.6, while the remaining 123 hybrids had Q values >0.6. Of these hybrids a large proportion of individuals (62%; 76 of 123) had higher northwestern ancestry, and 37 had higher southwestern ancestry. Hybridization rates were comparable between transects; west of the Rocky Mountains, 34.3% (72 of 210) of individuals were identified as hybrids, while east of the Rocky Mountains 38.3% (92 of 240) of individuals were identified as hybrids.
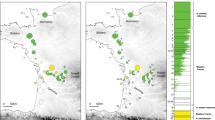
Eastern and western warbling vireos come into contact across a broad area (Fig. 4). Our spatial analyses indicate that the Great Plains contact zone encompasses an area of 294,187 km2. Within the Great Plains contact zone, the northern contact zone (73,733 km2) is about one third the size of the southern contact zone (220,454 km2). The Rocky Mountain contact zone is much larger (1,142,793 km2), approximately four times greater than the Great Plains contact zone. Within the Rocky Mountain contact zone, the eastern Rocky Mountain contact zone spans a larger area (1,102,147 km2) than the western Rocky Mountain contact zone (664,708 km2). The eastern and western Rocky Mountain contact zones overlap in a narrow area (54,181 km2) in southern British Columbia, northern Idaho, and Montana.
Correlates with ancestry and genetic distance
Plots examining the frequency of eastern microsatellite and mtDNA genotypes across habitat and elevation suggest that genetic variation is clinal across the Great Plains contact zone (Fig. 5A–D). Nuclear ancestry changed with elevation (r = 0.46) and habitat (r = 0.57) (Table 2). Eastern genotypes became less common as elevation increased and were less abundant in mixed and coniferous forests. By comparison, mtDNA haplotypes showed a strong correlation with elevation (r = 0.67) and habitat (r = 0.74). Within the northern Great Plains transect, nuclear ancestry was strongly correlated with elevation (r = 0.59) and habitat (r = 0.73), and mtDNA variation was strongly correlated with habitat (0.69), while the correlation between mtDNA haplotypes and elevation (r = 0.47) was moderate. In the southern Great Plains transect, we observed similar patterns for the correlation between nuclear ancestry and elevation (r = 0.61) and habitat (r = 0.54), whereas mtDNA haplotypes were strongly correlated with both habitat (r = 0.75) and elevation (r = 0.75).
Proportion of individuals assigned to eastern or northwestern genetic groups based on mtDNA (A, B) or nuclear (C–F) data across both the Great Plains (A–D) and Rocky Mountain (E, F) contact zones as a function of elevation and habitat. Results for individual membership probabilities for individuals analyzed in STRUCTURE at K = 2 are shown above each plot. Black lines denote groups delineated based on elevation or habitat. Orange individuals have western mtDNA (A, B) and nuclear (C, D) ancestry, and blue shows individuals with eastern mtDNA (A, B) and nuclear (C, D) ancestry. For E, F, green individuals have southwestern nuclear ancestry and purple individuals have northwestern nuclear ancestry. Eastern and western transects in the Rocky Mountain contact zone have been pooled to depict the overall pattern present across each transect (E, F).
Across the Rocky Mountain contact zone, correlations between nuclear ancestry and elevation were moderate (r = −0.46), while ancestry was weakly correlated with habitat (r = −0.21, Fig. 5E, F). Northern genotypes were more prominent at lower elevations, and in deciduous and mixed forests. The correlation between elevation and nuclear ancestry appears to be driven by patterns in the eastern Rocky Mountain contact zone, where this correlation is relatively stronger than the patterns we observed in the western Rocky Mountain contact zone (reast = −0.52 vs. rwest = −0.18). Both northwestern and southwestern nuclear genotypes had similar frequencies in coniferous forests. Within each of the Rocky Mountain transects, we observed a similar relationship between nuclear ancestry and habitat where ancestry was weakly correlated with habitat (western transect r = −0.25; eastern transect r = −0.21). Nuclear ancestry was moderately correlated with latitude, as northwestern genotypes replaced southwestern genotypes at higher latitudes; this pattern was maintained across both the western (r = 0.54) and eastern (r = 0.57) Rocky Mountain contact zone transects.
Habitat and elevation explained a small but significant proportion of genetic divergence in the Great Plains contact zone based on our analyses using dbRDA and partial-dbRDA models (Table 2). Habitat (R2 = 3.9%) accounted for greater variation (based on microsatellite genetic distance) than elevation (R2 = 2.9%), but the model that included both habitat and elevation variables accounted for a greater proportion of the variance (R2 = 5.2%). We detected a pattern of isolation-by-distance (R2 = 3.7%); habitat and elevation explained a small proportion of the genetic variation when we accounted for geographic distance (R2 = 2.9%). These patterns across the entire Great Plains contact zone were primarily maintained when we analyzed the northern and southern transects separately. Our mtDNA models using Nei’s genetic distances performed better than our models using microsatellite data, but yielded similar results. Habitat and elevation again accounted for greater variation than any other variable (R2 = 66.2%). We also detected a pattern of isolation-by-distance (R2 = 36.7%, p < 0.001); habitat and elevation explained a proportion of the genetic variation when we accounted for geographic distance (R2 = 32.7%). MtDNA patterns were comparable across both transects in the Great Plains contact zone, although elevation and isolation-by-distance accounted for greater variation in the southern transect than the northern transect (elevation: R2southern = 56.5% vs. R2northern = 22.1%; isolation-by-distance: R2southern = 47.1% vs. R2northern = 14.4%).
Habitat, elevation, and latitude (R2 = 0.2–1.0%) together accounted for a small proportion of the observed variance (using dbRDA and partial-RDA) in microsatellite genetic divergence for the Rocky Mountain contact zone, and only habitat was significant. When we analyzed the eastern and western transects separately, habitat, elevation, latitude, and geographic distance were all significant predictors, although the variables accounted for a relatively small portion of the variance (R2 = 0.5–2.6%). Patterns across both western and eastern Rocky Mountain transects were similar. Overall, habitat and elevation accounted for the greatest portion of variance and these values were comparable between transects (eastern Rocky Mountain transect: R2 = 1.9, p = 0.03; western Rocky Mountain transect: R2 = 2.6, p = 0.003).
Morphological variation
Correlations between ancestry and mass (r = −0.54) and wing length (r = −0.42) were moderate across the Great Plains contact zone (Table 3). Birds with western genotypes are smaller than birds with eastern genotypes. Within the Great Plains transects, wing length (r = −0.69) and mass (r = −0.77) were more strongly correlated with ancestry in the northern transect than the southern transect (wing length: r = −0.37; mass: r = −0.57). Mass and wing length were also correlated with habitat and elevation, but mass and wing length decreased with increases in elevation (mass: r = −0.51; wing length: r = −0.27) and the transition from deciduous to mixed and coniferous forests (mass: r = −0.45; wing length: r = −0.38).
Across the Rocky Mountain contact zone, the correlation between morphology and nuclear genotype was less apparent. Wing length was weakly correlated with ancestry (r = −0.12) and mass was not correlated with ancestry (r = 0.02); these patterns were not maintained when we analyzed the eastern and western Rocky Mountain transects separately. Wing length showed weak correlations with both elevation (r = 0.26) and habitat (r = −0.17), but mass was not correlated with either of these variables (r = −0.03 and −0.01, respectively). The relationship between wing length, habitat, and elevation appears to be driven by the eastern Rocky Mountain transect (elevation: r = 0.16; habitat: r = −0.21), as wing length was not correlated with either variable in the western Rocky Mountain transect. Wing length was weakly correlated with latitude (r = −0.20), a pattern maintained in the eastern Rocky Mountain transect (r = −0.17), but not present in the western Rocky Mountain transect (r = −0.03).
Discussion
The results of our study provide evidence for the complexity and variability of secondary contact and hybrid zones. Genetic divergence, in the Great Plains contact zone, between eastern and western warbling vireos is closely tied to habitat and elevation with hybrids found predominately in transitional areas. Comparatively, genetic structure and hybridization in the Rocky Mountain contact zone is weakly correlated with habitat and elevation. While morphological differences are correlated with habitat and elevation in the Great Plains contact zone, both of these variables are also strongly correlated with ancestry suggesting that morphological variation may be better predicted by genetic differentiation.
Habitat type explained nuclear ancestry and mtDNA clade assignment across the Great Plains contact zone, with a decrease in the frequency of eastern genotypes and haplotypes as habitat transitioned from non-forested and deciduous forests to mixed and coniferous forests. Although previous work brought attention to this relationship between habitat and genetic ancestry in the central Alberta contact zone (Lovell et al. 2021), our study provides empirical evidence that ancestry varies with both habitat and elevation. This correlation between habitat and genetic divergence is common among vireonids (Johnson 1995; Cicero and Johnson 1998; Zwartjes 2001) and other animal species (Carling and Thomassen 2012; Tarroso et al. 2014; Martin et al. 2017; Bell and Irian 2019; MacDonald et al. 2020). Western and eastern populations of warbling vireos likely diversified as the result of isolation in multiple refugia during Pleistocene glaciations (Lovell et al. 2021; Carpenter et al. 2021), a pattern common for other boreal (Weir and Schluter 2004) and temperate species (Johnson and Cicero 2004; Spellman and Klicka 2007; Manthey et al. 2011). This relationship between genetic ancestry and habitat likely reflects different environmental conditions during the Last Glacial Maximum (Avise 2000; Swenson 2006), and postglacial range expansion by both eastern and western warbling vireos likely followed, and were limited by, the recolonization of certain tree species and climate gradients (Williams 2003; Swenson 2006).
Lovell et al. (2021) found that hybridization between eastern and western warbling vireo populations occurs across a narrow hybrid zone in central Alberta, a transitional zone between deciduous-dominated parkland habitat and coniferous-dominated boreal forest. Our study of a larger geographic area and more samples indicates that hybridization in the Great Plains occurs across a broader area than previously reported. Our analyses also demonstrate that elevation influences genetic ancestry across the Great Plains contact zone, especially for mtDNA patterns where eastern haplotypes are almost entirely absent above 1000 m. Given the role of avian mitochondria in meeting energetic demands, selection for western haplotypes may potentially occur in response to the physiological requirements of higher elevation environments (Cheviron and Brumfield 2012; Abbott and Brennan 2014; Toews et al. 2014).
Ecological and habitat differences are known to act as mechanisms of diversification within the genus Vireo (Cicero and Johnson 1998) and these results from the Great Plains contact zone adds to this literature. In contrast, these same variables did not explain hybridization patterns across the Rocky Mountain contact zone between western warbling vireo genetic groups. The greater size of the Rocky Mountain contact zone and distribution of hybrids compared to the Great Plains contact zone reflects the weak relationship between ancestry and environmental variables. While individuals of northwestern ancestry were most common in mixed forests and those with southwestern ancestry in non-forested habitat, the proportion of individuals with northwestern and southwestern ancestry in both deciduous and coniferous forests was similar. Our characterization of habitat types into four broad categories may not have been able to detect discrete fine-scale habitat patterns between the more closely-related western warbling vireo genetic groups, such as whether northwestern or southwestern populations are associated with certain tree species, as has been suggested for other taxa (van Els et al. 2012; Graham et al. 2021). Moreover, we believe that a plausible explanation for the weak relationship between nuclear genotype and habitat is attributed to the recent isolation of these taxa. Mitochondrial divergence between western warbling vireos (northwestern and southwestern groups, respectively 0.2–0.3%; Carpenter et al. 2021) is low when compared to ecologically diverged eastern and western warbling vireos (3–4%; Lovell et al. 2021; Carpenter et al. 2021). Furthermore, the relatively low mtDNA genetic variation within this western clade may be indicative of a selective sweep for haplotypes adapted to high elevation environments (Dubay and Witt 2014).
Similar to habitat, the correlation between ancestry and elevation across the Rocky Mountain contact zone did not entirely explain the patterns of nuclear genetic divergence among populations of western warbling vireos. Overall, birds with southwestern genotypes are more prominent at higher elevations than those with northwestern genotypes. This trend likely reflects the biogeographic nature of the southwestern genetic group’s range, with much of its distribution in the southwestern United States, where some of the tallest mountains in the contiguous United States (e.g., Sierra Nevada’s) are found. Other vertebrate and plant species exhibit similar genetic patterns (Spellman and Klicka 2007; Haselhorst et al. 2019), and these patterns likely arose as a result of barriers to gene flow and Pleistocene glaciations.
Finally, morphology varies across both the Great Plains and Rocky Mountain contact zones. Morphological differences between eastern and western warbling vireos are more pronounced than those between the two western groups (Carpenter et al. 2021). Across the Great Plains contact zone, mass and wing length are moderately correlated with habitat and elevation, and the strength of these relationships is comparable to the relationship between ancestry and morphological variation. Eastern and western warbling vireos follow separate migratory routes (Voelker and Rohwer 1998), and differences in wing length may reflect the fact that eastern birds migrate longer distances than western birds, as has been shown for other migratory species (Marchetti et al. 1995; Nowakowski et al. 2014). Morphological differences and nuclear ancestry were not correlated in the Rocky Mountain contact zone; however, the absence of morphological variation between western groups could be due to these birds following similar migratory pathways, exploit similar habitat, or are found at similar elevation.
Conclusions
Our analyses of two large contact zones, and multiple transects within each, demonstrates how variable admixture and hybridization can be. The Great Plains and Rocky Mountain contact zones vary substantially in size and shape; looking at independent transects showcases the dynamic nature of each of these zones, an aspect that has been poorly studied. Our research highlights the importance of habitat and elevation in promoting and maintaining isolation among divergent lineages, like eastern and western warbling vireo genetic groups along the Great Plains–Rocky Mountain ecotone. Although neither of these variables in our study promote isolation between the two western genetic groups, this result likely reflects their recent divergence and similar niches. Combined, our work adds novel insight into patterns of secondary contact across an expansive latitudinal scale, and the environment’s role in mediating hybridization between ecologically-adapted and divergent cryptic groups.
Data availability
Sequence data are available in GenBank with the following accession numbers for some of the samples used in this study: ON087602-ON087609 https://doi.org/10.5061/dryad.p5hqbzkrc.
Change history
24 May 2022
A Correction to this paper has been published: https://doi.org/10.1038/s41437-022-00544-y
References
Abbott RJ, Brennan AC (2014) Altitudinal gradients, plant hybrid zones and evolutionary novelty. Philos Trans R Soc B Biol Sci 369:6–9
Avise JC (2000) Phylogeography: the history and formation of species. Harvard University Press, Cambridge, MA
Baldassarre DT, White TA, Karubian J, Webster MS (2014) Genomic and morphological analysis of a semipermeable avian hybrid zone suggests asymmetrical introgression of a sexual signal. Evolution 68:2644–2657
Barr KR, Dharmarajan G, Rhodes OE, Lance R, Leberg PL (2007) Novel microsatellite loci for the study of the black-capped vireo (Vireo atricapillus). Mol Ecol Notes 7:1067–1069
Barton NH, Gale KS (1993) Hybrid zones and the evolutionary process. In: Harrison RG (ed.) Hybrid Zones and the Evolutionary Process. Oxford University Press, New York, NY
Barton NH, Hewitt GM (1989) Adaption, speciation and hybrid zones. Nature 341:497–503
Billerman SM, Murphy MA, Carling MD (2016) Changing climate mediates sapsucker (Aves: Sphyrapicus) hybrid zone movement. Ecol Evol 6:7976–7990
Bell RC, Irian CG (2019) Phenotypic and genetic divergence in reed frogs across a mosaic hybrid zone on São Tomé Island. Biol J Linn Soc 128:672–680
Bensch S, Price T, Kohn J (1997) Isolation and characterization of microsatellite loci in a Phylloscopus warbler. Mol Ecol 6:91–92
Bradbury IR, Bowman S, Borza T, Snelgrove PVR, Hutchings JA, Berg PR et al. (2014) Long distance linkage disequilibrium and limited hybridization suggest cryptic speciation in Atlantic cod. PLoS ONE 9:e106330
Brelsford A, Irwin DE (2009) Incipient speciation despite little assortative mating: the yellow-rumped warbler hybrid zone. Evolution 63:3050–3060
Burg TM, Croxall JP (2004) Global population structure and taxonomy of the wandering albatross species complex. Mol Ecol 13:2345–2355
Carling MD, Zuckerberg B (2011) Spatio-temporal changes in the genetic structure of the Passerina bunting hybrid zone. Mol Ecol 20:1166–1175
Carling MD, Thomassen HA (2012) The role of environmental heterogeneity in maintaining reproductive isolation between hybridizing Passerina (Aves: Cardinalidae) buntings. Int J Ecol 2012:295463
Carpenter AM, Graham BA, Spellman GM, Klicka J, Burg TM (2021) Genetic, bioacoustic and morphological analyses reveal cryptic speciation in the warbling vireo complex (Vireo gilvus: Vireonidae: Passeriformes). Zool J Linn Soc zlab036 https://doi.org/10.1093/zoolinnean/zlab036
Cicero C, Johnson NK (1998) Molecular phylogeny and ecological diversification in a clade of New World songbirds (genus Vireo). Mol Ecol 7:1359–1370
Chenuil A, Cahill AE, Délémontey N, Du Salliant du Luc E, Fanton H (2019) Problems and questions posed by cryptic species. A framework to guide future studies. Assessing to conserving biodiversity. History, philosophy and theory of the life sciences, Vol. 24. Springer. Daubenmire, Cham
Cheviron ZA, Brumfield RT (2012) Genomic insights into adaptation to high-altitude environments. Heredity 108:354–361
Coyne JA, Orr HA (2004) Speciation. Sinauer and Associates, Sunderland, Massachusetts
Culumber ZW, Shepard DB, Colemans SW, Rosenthal GG, Tobler M (2012) Physiological adaptation along environmental gradients and replicated hybrid zone structure in swordtails (Teleostei: Xiphophorus). J Evol Biol 25:1800–1814
Dubay SG, Witt CC (2014) Differential high-altitude adaptation and restricted gene flow across a mid-elevation hybrid zone in Andean tit-tyrant flycatchers. Mol Ecol 23:3551–3565
Garroway CJ, Bowman J, Cascaden TJ, Holloway GL, Mahan CG, Malcolm JR et al. (2010) Climate change induced hybridization in flying squirrels. Glob Chang Biol 16:113–121
Grabenstein KC, Taylor SA (2018) Breaking barriers: Causes, consequences, and experimental utility of human-mediated hybridization. Trends Ecol Evol 33:198–212
Graham BA, Cicero C, Strickland D, Woods JG, Coneybeare H, Dohms KM et al. (2021) Cryptic genetic diversity and cytonuclear discordance characterize contact among Canada jay (Perisoreus canadensis) morphotypes in western North America. Biol J Linn Soc 132:725–740
Hammer Ø, Harper DA, Ryan PD (2001) Paleontological statistics software package for education and data analysis. Palaeontol Electron 4:9
Haselhorst MSH, Parchman TL, Buerkle CA (2019) Genetic evidence for species cohesion, substructure and hybrids in spruce. Mol Ecol 28:2029–2045
Hijmans RJ, Cameron SE, Parra JL, Jones PG, Jarvis A (2005) Very high resolution interpolated climate surfaces for global land areas. Int J Climatol 25:1965–1978
Hawley DM (2005) Isolation and characterization of eight microsatellite loci from the house finch (Carpodactus mexicanus). Mol Ecol Notes 5:443–445
Hebert PDN, Stoeckle MY, Zemlak TS, Francis CM (2004) Identification of birds through DNA barcodes. PLoS Biol 2:e312
Hewitt GM (1988) Hybrid zones-natural laboratories for evolutionary studies. Trends Ecol Evol 3:158–167
Hewitt GM (2001) Speciation, hybrid zones and phylogeography—or seeing genes in space and time. Mol Ecol 10:537–549
Hijmans RJ, Cameron SE, Parra JL, Jones PG, Jarvis A (2005) Very high resolution interpolated climate surfaces for global land areas. Int J Climatol 25:1965–1978
Hindley JA, Graham BA, Pulgarin-R PC, Burg TM (2018) The influence of latitude, geographic distance, and habitat discontinuities on genetic variation in a high latitude montane species. Sci Rep. 8:11846
Hinojosa JC, Koubínová D, Szenteczki MA, Pitteloud C, Dincă V, Alvarez N et al. (2019) A mirage of cryptic species: Genomics uncover striking mitonuclear discordance in the butterfly Thymelicus sylvestris. Mol Ecol 28:3857–3868
Hubisz MJ, Falush D, Stephens M, Pritchard JK (2009) Inferring weak population structure with the assistance of sample group information. Mol Ecol Res 9:1322–1332
Irwin DE (2020) Assortative mating in hybrid zones is remarkably ineffective in promoting speciation. Evolution 195:E150–E167
Johnson NK (1995) Speciation in vireos. I. Macrogeographic patterns of allozymic variation in the Vireo solitarius complex in the contiguous United States. Condor 97:903–919
Johnson NK, Cicero C (2004) New mitochondrial DNA data affirm the importance of Pleistocene speciation in North American birds. Evolution 58:1122–1130
Larson EL, Tinghitella RM, Taylor SA (2019) Insect hybridization and climate change. Front Ecol Evol 7:348
Legendre P, Fortin M-J (2010) Comparison of the Mantel test and alternative approaches for detecting complex multivariate relationships in the spatial analysis of genetic data. Mol Ecol Resour 10:831–844
Lovell SF, Lein MR, Rogers SM (2021) Cryptic speciation in the warbling vireo (Vireo gilvus). Ornithology 138:ukaa071
MacDonald ZG, Dupuis JR, Davis CS, Acorn JH, Nielsen SE, Sperling FAH (2020) Gene flow and climate-associated genetic variation in a vagile habitat specialist. Mol Ecol 29:3889–3906
Manthey JD, Klicka J, Spellman GM (2011) Cryptic diversity in a widespread North American songbird: phylogeography of the brown creeper (Certhia americana). Mol Phylogenet Evol 58:502–512
Marchetti K, Price T, Richman A (1995) Correlates of wing morphology with foraging behaviour and migration distance in the genus Phylloscopus. J Av Biol 26:177–181
Martin H, Touzet P, Dufay M, Gode C, Schmitt E, Lahiani E et al. (2017) Lineages of Silene nutans developed rapid, strong, asymmetric postzygotic reproductive isolation in allopatry. Evolution 71:1519–1531
Martinez JG, Soler JJ, Soler M, Moller AP, Burke T (1999) Comparative population structure and gene flow of a brood parasite, the great spotted cuckoo (Clamator glandarius) and its primary host, the magpie (Pica pica). Evolution 53:269–278
Mettler RD, Spellman GM (2009) A hybrid zone revisited: Molecular and morphological analysis of the maintenance, movement, and evolution of a Great Plains avian (Cardinalidae: Pheucticus) hybrid zone. Mol Ecol 18:3256–3267
Meirmans PG, Van Tienderen PH (2004) GENOTYPE and GENODIVE: two programs for the analysis of genetic diversity of asexual organisms. Mol Ecol Notes 4:792–794
Nowakowski JK, Szulc J, Remisiewicz M (2014) The further the flight, the longer the wing: relationship between wing length and migratory distance in Old World reed and bush warblers (Acrocephalidae and Locustellidae). Ornis Fennica 91:178–186
Pavolova A, Amos JN, Joseph L, Loynes K, Austin JJ, Keogh JS et al. (2013) Perched at the mito-nuclear crossroads: divergent mitochondrial lineages correlate with environment in the face of ongoing nuclear gene flow in an Australian bird. Evol 67:3412–3428
Piertney SB, Marquiss M, Summers R (1998) Characterization of tetranucleotide microsatellite markers in the Scottish crossbill (Loxia scotica). Mol Ecol 7:1261–1263
Porras-Hurtado L, Ruiz Y, Santos C, Phillips C, Carracedo A, Lareu MV (2013) An overview of STRUCTURE: Applications, parameter settings, and supporting software. Front Genet 4:98
Pritchard J, Stephens M, Donnelly P (2000) Inference of population structure using multilocus genotype data. Genetics 155:945–959
Reding DM, Castañeda-Rico S, Shirazi S, Hofman CA, Cancellare IA, Lance SL et al. (2021) Mitochondrial genomes of the United States distribution of gray fox (Urocyon cinereoargenteus) reveal a major phylogeographic break at the Great Plains suture zone. Front Ecol Evol. https://doi.org/10.3389/fevo.2021.666800.
Richardson DS, Jury FL, Dawson DA, Salgueiro P, Komdeur J, Burke T (2003) Fifty Seychelles warbler (Acrocephalus sechellensis) microsatellite loci polymorphic in Sylviidae species and their cross-species amplification in other passerine birds. Mol Ecol 9:2225–2230
Riordan EC, Gugger PF, Ortego J, Smith C, Gaddis K, Thompson P et al. (2016) Association of genetic and phenotypic variability with geography and climate in three southern California oaks. Am J Bot 103:73–85
Rush AC, Cannings RJ, Irwin DE (2009) Analysis of multilocus DNA reveals hybridization in a contact zone between Empidonax flycatchers. J Avian Biol 40:614–624
Sartor CC, Cushman SA, Wan HY, Kretschmer R, Pereira JA, Bou N et al. (2021) The role of the environment in the spatial dynamics of an extensive hybrid zone between two neotropical cats. J Evol Biol 34:614–627
Schukman JM, Lira-Noriega A, Townsend Peterson A (2011) Multiscalar ecological characterization of Say’s and eastern phoebes and their zone of contact in the Great Plains. Condor 113:372–384
Seehausen O, Takimoto G, Roy D, Jokela J (2008) Speciation reversal and biodiversity dynamics with hybridization in changing environments. Mol Ecol 17:30–44
Semenchuk GP (1992) The Atlas of Breeding Birds of Alberta. Fed. of Alberta Naturalists, Edmonton, p 243
Peakall R, Smouse PE (2012) GenAlEx 6.5: Genetic analysis in Excel. Population genetic software for teaching and research–an update. Bioinformatics 28:2537–2539
Sorenson MD, Ast JC, Dimcheff DE, Yuri T, Mindell DP (1999) Primers for a PCR-based approach to mitochondrial genome sequencing in birds and other vertebrates. Mol Phylogent Evol 12:105–114
Spellman GM, Klicka J (2007) Phylogeography of the white-breasted nuthatch (Sitta carolinensis): diversification in North American pine and oak woodlands. Mol Ecol 16:1729–1740
Stenzler LM, Fitzpatrick JW (2002) Isolation of microsatellite loci in the Florida scrub jay Aphelocoma coerulescens. Mol Ecol Notes 2:547–550
Swenson NG (2006) GIS-based niche models reveal unifying climatic mechanisms that maintain location of avian hybrid zones in a North America suture zone. J Evol Biol. 19:717–725
Swenson NG, Howard DJ (2005) Clustering of contact zones, hybrid zones, and phylogeographic breaks in North America. Am Nat 166:581–591
Tarr CL, Fleischer RC (1998) Primers for polymorphic GT microsatellites isolated from the Mariana crow, Corvus kubaryi. Mol Ecol 7:253–255
Tarroso P, Pereira RJ, Martínez-Freiría F, Godinho R, Brito JC (2014) Hybridization at an ecotone: Ecological and genetic barriers between three Iberian vipers. Mol Ecol 23:1108–1123
Taylor SA, Larson EL, Harrison RG (2015) Hybrid zones: windows on climate change. Trends Ecol Evol 30:398–406
Toews DPL, Mandic M, Richards JG, Irwin DE (2014) Migration, mitochondria and the yellow-rumped warbler. Evolution 68:241–255
Toews DPL, Campagna L, Taylor SA, Balakrishnan CN, Baldassarre DT, Deane-Coe PE et al. (2016) Genomic approaches to understanding population divergence and speciation in birds. Auk 133:13–30
Toews DPL, Irwin DE (2008) Cryptic speciation in a Holarctic passerine revealed by genetic and bioacoustic analyses. Mol Ecol 17:2691–2705
van Els P, Cicero C, Klicka J (2012) High latitudes and high genetic diversity: Phylogeography of a widespread boreal bird, the gray jay (Perisoreus canadensis). Mol Phylogenet Evol 63:456–465
Voelker G, Rohwer S (1998) Contrasts in scheduling of molt and migration in eastern and western warbling vireos. Auk 155:142–155
Walsh J, Billerman SM, Rohwer VG, Butcher BG, Lovette IJ (2020) Genomic and plumage variation across the controversial Baltimore and Bullock’s oriole hybrid zone. Auk 137:1–15
Walsh J, Rowe RJ, Olsen BJ, Shriver WG, Kovach AI (2016) Genotype-environment associations support a mosaic hybrid zone between two tidal marsh birds. Ecol Evol 6:279–294
Walsh P, Metzger D, Higuchi R (1991) Chelex 100 as a medium for simple extraction of DNA for PCR-based typing from forensic material. Biotechniques 10:506–513
Weir JT, Schluter D (2004) Ice sheets promote speciation in boreal birds. Proc R Soc B 271:1881–1887
Williams JW (2003) Variations in tree cover in North America since the last glacial maximum. Glob Planet Change 35:1–23
Williams DA, Berg EC, Hale AM, Hughes CR (2004) Characterization of microsatellites for parentage studies of white-throated magpie-jays (Calocitta formosa) and brown jays (Cyanocorax morio). Mol Ecol Notes 4:509–511
Zwartjes PW (2001) Genetic structuring among migratory populations of the black-whiskered vireo, with a comparison to the red-eyed vireo. Condor 103:439–448
Acknowledgements
We are grateful for the hard work of graduate students and field assistants over the years that helped with field sample collections. Thank you to the following museums whose voucher sample contributions made our latitudinal-scale research possible: the Denver Museum of Nature and Science, the Burke Museum of Natural History and Culture, and the University of Michigan Museum. A special acknowledgement to Scott Lovell for giving us access to samples from your study, and to Jocelyn Hudon at the Royal Alberta Museum for letting us acquire said samples on short notice. Finally, a thank you to Dr. Mable, the associate editor, and the two anonymous reviewers for their comments that helped to improve the paper.
Funding
This work was funded by the Natural Sciences and Engineering Research Council of Canada (NSERC Discovery Grant RGPIN-2019-05068) (TMB) and the National Science Foundation (DEB0814841) (GMS). Samples were collected under the appropriate provincial, state, and federal permits.
Author information
Authors and Affiliations
Contributions
AMC was responsible for the development of the research project, planning and conducting field work, DNA extractions and PCRs, mtDNA and microsatellite genotyping, assisting with analyses and figures, and writing and revisions of the paper. BAG was responsible for conducting field work, taking the lead on analyses and figures, and helping with writing and revisions of the paper. GMS was responsible for loaning samples and morphological data for the study and providing input for the paper and paper revisions. TMB was responsible for the development of the research project, funding the project, and providing input for the paper and paper revisions.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Associate editor Sam Banks.
Rights and permissions
About this article
Cite this article
Carpenter, A.M., Graham, B.A., Spellman, G.M. et al. Do habitat and elevation promote hybridization during secondary contact between three genetically distinct groups of warbling vireo (Vireo gilvus)?. Heredity 128, 352–363 (2022). https://doi.org/10.1038/s41437-022-00529-x
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41437-022-00529-x