Abstract
Explicitly accounting for phenotypic differentiation together with environmental heterogeneity is crucial to understand the evolutionary dynamics in hybrid zones. Species showing intra-specific variation in phenotypic traits that meet across environmentally heterogeneous regions constitute excellent natural settings to study the role of phenotypic differentiation and environmental factors in shaping the spatial extent and patterns of admixture in hybrid zones. We studied three environmentally distinct contact zones where morphologically and reproductively divergent subspecies of Salamandra salamandra co-occur: the pueriparous S. s. bernardezi that is mostly parapatric to its three larviparous subspecies neighbours. We used a landscape genetics framework to: (i) characterise the spatial location and extent of each contact zone; (ii) assess patterns of introgression and hybridization between subspecies pairs; and (iii) examine the role of environmental heterogeneity in the evolutionary dynamics of hybrid zones. We found high levels of introgression between parity modes, and between distinct phenotypes, thus demonstrating the evolution to pueriparity alone or morphological differentiation do not lead to reproductive isolation between these highly divergent S. salamandra morphotypes. However, we detected substantial variation in patterns of hybridization across contact zones, being lower in the contact zone located on a topographically complex area. We highlight the importance of accounting for spatial environmental heterogeneity when studying evolutionary dynamics of hybrid zones.
Similar content being viewed by others
Introduction
The spatial extent of hybrid zones (i.e. areas where gene flow between lineages occurs), and the degree of admixture between taxa in these areas are influenced by endogenous (genetic factors) and exogenous (environmental factors) selection, but also through extrinsic neutral processes (e.g. landscape barriers to gene flow). It has been shown that genomic incompatibilities between parent taxa at prezygotic (e.g. reduced fertilization success) and/or postzygotic level (e.g. reduce hybrid fitness) influence evolutionary dynamics in hybrid zones1,2,3,4. Environmental variation (i.e. isolation by environment, IBE) may also promote selection for or against hybrids5,6, or ecological segregation of parent taxa7, while landscape features (e.g. rivers, mountains, roads) can act as barriers to dispersal and gene flow (i.e. isolation by resistance, IBR) across heterogeneous landscapes8,9,10. Additionally, the historical context underlying the formation and maintenance of hybrid zones can also affect the extent to which contemporary gene flow occurs between species, with past demography and age of contact zones comprising relevant factors11,12,13.
Contact zones involving intraspecific evolutionary units (e.g. subspecies) often resulted from allopatric differentiation of neighbouring lineages in distinct refugia during unfavourable climatic periods of the Pleistocene, followed by population expansions during climatically favourable periods14. These lineages seldom achieve reproductive isolation and continue to hybridize in secondary contact zones15. The evolutionary fate of hybrid zones depends on the strength of barriers to gene flow between diverged lineages. On the one hand, secondary contact zones can result in extensive admixture of parental genotypes or lineage fusion when there are not effective barriers to gene flow16. On the other hand, selection against hybrids can favour the evolution of barriers to gene flow and ultimately lead to reproductive isolation and speciation17. However, an intermediate scenario commonly found in secondary contact zones involves the formation of tension zones often characterized by an equilibrium between dispersal and selection4,18.
Phenotypes are targets of selection, playing a major role in species’ performance (e.g. dispersal potential), recognition, and diversification19, ultimately dictating the extent and population dynamics of hybrid zones through processes such as assortative mating and overdominance4,20,21. Thus, gene flow between subspecies or lineages occurring in secondary contact zones is also determined by their genomic and phenotypic differentiation11,22,23,24. Accounting for this phenotypic differentiation together with the bioclimatic gradients and landscape heterogeneity across contact zones is crucial to understanding the evolution of phenotypic traits and patterns of introgression. This is especially relevant under a scenario of pronounced climate and landscape changes driven by anthropogenic activities because they can alter the evolutionary trajectories of hybridizing taxa25 and the fate of evolutionary significant units26.
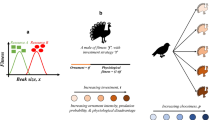
The fire salamander, Salamandra salamandra (Linnaeus, 1758), is widely distributed in Europe and offers an exceptional model system for studying neutral and adaptive processes, particularly in the Iberian Peninsula, where shows its highest intraspecific genetic and phenotypic variation27,28,29,30. This likely resulted from the complex Iberian physiography and the climatic oscillations during the Pleistocene, which promoted allopatric divergence in populations isolated within Iberian glacial refugia30,31. In this region, S. salamandra shows a remarkable intraspecific variation in external morphology (e.g. body size, head shape, colour pattern)28,29,30, and more strikingly, in reproductive modes27. Two viviparous strategies co-occur27: (1) an aquatic-breeding and widespread reproductive mode, larviparity (parturition of aquatic larvae); and (2) a terrestrial-breeding and spatially restricted reproductive mode, pueriparity (parturition of terrestrial juveniles). Pueriparity in S. salamandra evolved at least twice: in the subspecies S. s. bernardezi along the Cantabrian Mountains during the Pleistocene31, and in two Atlantic islands in the subspecies S. s. gallaica during the Holocene32,33. Salamandra s. bernardezi is currently surrounded by larviparous populations of S. s. gallaica and S. s. bejarae along its western and southern range, respectively. Whether S. salamandra populations across the southern slopes of the Cantabrian Mountains belong to S. s. bejarae (type locality in the Iberian Central System), or to an undescribed subspecies within the S. s. gallaica complex, needs taxonomic revision. However, they both represent independent and closely related lineages within one of the two main S. salamandra nuclear clades30 and, hence, we opted to consider those populations as traditionally S. s. bejarae. The second nuclear clade includes the Iberian S. s. bernardezi and S. s. fastuosa, together with the Appenninic subspecies S. s. gigliolii. On the easternmost range, S. s. bernardezi (Asturias-Cantabria border) meets S. s. fastuosa, which is considered larviparous despite exhibiting both reproductive strategies in some populations27, 34 (Fig. 1). Thus, the northern Iberian populations of fire salamanders constitute an exceptional natural system, in which multiple lineages and phenotypes meet across heterogeneous contact zones in a relatively continuous range, and as such, provide an unusual opportunity to evaluate how exogenous and endogenous factors shape patterns of introgression at the subspecific level.
Upper left: distribution of the fire salamander, Salamandra salamandra, with the study area denoted with an inset (A). Upper right: mtDNA and nuDNA trees to show subspecies relationships and approximate dates of major splits. The mtDNA tree was generated with cytochrome b sequences from this study while the nuDNA tree is adapted from the topology obtained with phylogenomic data in Burgon et al.30. Bottom: sampling sites across each of the studied contact zone: North Galicia (NG), Cantabrian Mountains (CM) and Asturias-Cantabria border (AC). Adult salamander images represent the two morphotypes, while the reproductive mode of each studied subspecies (larviparous, pueriparous or both) is represented on the polygons depicting each subspecies’ distribution. The figure was created with QGIS software (version 3.2; https://qgis.org/en/site/). The satellite image was retrieved from Google Earth using the QuickMapServices plugin.
Here we investigate patterns of gene flow and introgression, and the underlying environmental drivers, between the pueriparous S. s. bernardezi and its three larviparous subspecies neighbours across three contact zones in Northern Spain: (1) northern Galicia, where S. s. bernardezi is in contact at west of its range with S. s. gallaica35; (2) the Cantabrian Mountains, where southern populations of S. s. bernardezi meet S. s. bejarae at relatively high elevations; and (3) the Asturias-Cantabria border where admixture between eastern populations of S. s. bernardezi and S. s. fastuosa was previously reported31. We used a fine-scale individual sampling, and combined one mitochondrial gene (cytochrome b) and 15 microsatellites markers in a landscape genetics approach to: (1) delimit and characterise the range of S. s. bernardezi; (2) identify patterns of introgression and quantify levels of admixture between pueriparous and larviparous subspecies in each contact zone; and (3) evaluate the role of environmental and geographic factors in explaining genetic admixture between the different subspecies. Considering previous studies examining phenotypic and/or genetic patterns across northern Iberian populations of S. salamandra29,31,35,36,37,38, we hypothesize that some degree of genetic admixture occurs between subspecies in all contact zones. However, we expect the level of admixture to depend upon the environmental heterogeneity in each area and, therefore, we expect reduced rates of hybridization under IBE or IBR scenarios. Moreover, because of the reduced population connectivity among pueriparous S. s. bernardezi populations38, we hypothesise that introgression between larviparous and pueriparous populations will be asymmetric.
Material and methods
Study system
Salamandra s. bernardezi and S. s. fastuosa are relatively small salamanders (body size up to 180 mm27), with stripe colouration pattern (dorsal yellow and black bands), and round snout29. While the former is strictly pueriparous27,37, the latter is mostly larviparous despite also presenting pueriparity across its range34 (Fig. 1). Conversely, S. s. gallaica and S. s. bejarae are large salamanders (body size up to 250 mm), show a spotted or blotched colouration pattern and a pointed snout29. These two subspecies are larviparous, although insular pueriparous populations of S. s. gallaica exist32,33.
Despite being phenotypically different in northern Spain, these subspecies inhabit similar habitats, particularly, deciduous woodlands (Quercus spp. and Fagus spp.) with nearby aquatic systems (streams or ponds) where larviparous females can give birth to aquatic larvae27,39. They can also be found in other terrestrial habitats, such as scrublands and coniferous forests, alpine scrublands, and more rarely in eucalyptus (Eucalyptus spp.) plantations39,40. Previous studies showed agricultural and urban areas comprise strong barriers to dispersal and gene flow in both larviparous and pueriparous fire salamanders38,41,42. However, pueriparous salamanders can also survive in harsh environments where water bodies are absent34,42, despite water courses (streams and rivers) comprising semi-permeable obstacles to dispersal due to their fully terrestrial life style38.
Study areas and sampling design
We carried out individual-based sampling (1–3 samples per locality) within each contact zone because this sampling approach was shown to provide greater statistical power in detecting landscape genetic patterns compared to population-based sampling43. We selected three heterogeneous study areas (i.e. different levels of environmental and habitat variability) where different subspecies, morphotypes and the two distinct reproductive modes meet, thus covering a large portion of the secondary contact zones between S. s. bernardezi and its neighbouring lineages (Fig. 1). We performed sampling during rainy nights from November 2013 until May 2015 (Supplementary Table S1). We sampled mostly adult individuals (toe clip), but on a few occasions we also sampled larvae (tail clip). Tissue samples were stored in absolute ethanol and all individuals were released in their sampling points. At the western range of S. s. bernardezi, we collected samples of 129 salamanders from 86 localities following a west–east axis along northern Galicia (NG hereafter), where intermixed phenotypes between S. s. bernardezi and S. s. gallaica were previously observed32,35 (Fig. 2). The landscape in NG is relatively flat (elevation ranges from 0 m up to 1016 m a.s.l. using the minimum extent of the samples, Supplementary Fig. S1) and is extensively fragmented by agricultural practices and Eucalyptus spp. plantations. Remnant populations of fire salamanders in NG occur in suitable forested and scrubland patches in lowland and hilly areas. At the southern range of S. s. bernardezi, we also followed a longitudinal sampling along the Cantabrian Mountain range (CM hereafter), where we collected samples of 73 individuals from 38 localities on both sides of the mountains, representing S. s. bernardezi and S. s. bejarae from the northern and southern slopes, respectively (Fig. 3). The landscape in this study plot is characterized by high elevation, steep terrains, and high topographic complexity (elevation ranging from 179 m up to 2343 m above sea level (a.s.l.), Supplementary Fig. S1), and is highly suitable for the species due to the native forests and scrublands that dominate the region with sparse agricultural patches. At the eastern range of S. s. bernardezi, we obtained samples of 123 individuals from 58 localities across the Asturias-Cantabria border (AC hereafter, Fig. 4). We did not extend the sampling further south to avoid sampling S. s. bejarae (Fig. 1). Despite the high elevation in this region (elevation ranging from 0 m up to 2394 m a.s.l., Supplementary Fig. S1), the mountains follow a west–east direction and thus, no major topographic barriers exist between these two subspecies, which frequently occur along coastal valleys. The AC area is composed of natural forests and scrublands, with eucalypt plantations, agricultural areas and human infrastructures (e.g. villages, roads) being predominant on the coast.
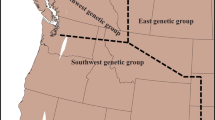
Genetic structure of S. salamandra across northern Galician populations (NG) as inferred by both mitochondrial (upper panel; two main lineages separating S. s. bernardezi and S. s. gallaica) and nuclear DNA (lower panel; STRUCTURE K = 2). Samples in the barplots are arranged from west to east. Genetic assignment to S. s. bernardezi or S. s. gallaica is colour-coded in blue and red, respectively. Black bars between structure plots indicate individuals with mitonuclear discordance (19%). The figure was created with QGIS software (version 3.2; https://qgis.org/en/site/).
Genetic structure of S. salamandra across the Cantabrian Mountains (CM) as inferred by both mitochondrial (upper panel; two main lineages separating S. s. bernardezi and S. s. bejarae) and nuclear DNA (lower panel; STRUCTURE K = 2). Samples in the barplots are arranged from west to east. Genetic assignment to S. s. bernardezi or S. s. bejarae is colour-coded in blue and yellow, respectively. Black bars between structure plots indicate individuals with mitonuclear discordance (6%). The figure was created with QGIS software (version 3.2; https://qgis.org/en/site/).
Genetic structure of S. salamandra across the Asturian-Cantabrian border (AC) as inferred by both mitochondrial (upper panel; two main lineages separating S. s. bernardezi and S. s. fastuosa) and nuclear DNA (lower panel; STRUCTURE K = 2). Samples in the barplots are arranged from west to east. Genetic assignment to S. s. bernardezi and S. s. fastuosa is colour-coded in blue and green, respectively. Black bars between structure plots indicate individuals with mitonuclear discordance (16%). The figure was created with QGIS software (version 3.2; https://qgis.org/en/site/).
Fieldwork for obtaining tissue samples was done with the corresponding permits from the regional administrations (Xunta de Galicia, Ref. 024/2014, and 410/2015; Gobierno del Principado de Asturias, Ref. 2018/2115; Castilla y León, EP/CYL/625/2013, EP/CYL/725/2015; Picos de Europa National Park CO/09/0125/2013, CO/09/012/2014, CO/09/065/2015). Sampling procedures were carried out following the Guidelines for Use of Live Amphibians and Reptiles in Field and Laboratory Research, 2nd Edition, revised by the Herpetological Animal Care and Use Committee (HACC) of the American Society of Ichthyologists and Herpetologists, 2004. All procedures complied with the country legal requirements on animal welfare (RD 53/2013) and were conducted in accordance with the guidelines of the Research Ethics Committee of the University of Oviedo (Comité Ético de Experimentación Animal de la Universidad de Oviedo) under authorisation #8-INV-2012. The members of the research team have approved licenses by the Service of Animal Welfare and Production of the Principality of Asturias to conduct experimental protocols with animals (license types C and D to A.G.N). This study was carried out in compliance with the ARRIVE guidelines (Animal Research: Reporting in Vivo Experiments) for how to report animal research in scientific publications (https://arriveguidelines.org/arrive-guidelines).
Molecular markers and laboratory procedures
We extracted genomic DNA using the Genomic DNA Tissue Kit (EasySpin), following the manufacturer’s protocol, and verified both the quantity and quality of extracted products in a 0.8% agarose gel. We used the DNA extract as template for polymerase chain reactions (PCR) to amplify and sequence one mitochondrial (mtDNA) fragment (ca. 1100 bp) of the cytochrome b and adjacent tRNAs (hereafter cytb). We used the primers Glu14100L and Pro15500H44 to amplify cytb following a previously described protocol28. We outsourced DNA sequencing to Genewiz Inc. (Leipzig, Germany), and inspected and aligned resulting chromatograms using GENEIOUS version 11.1.4 (http://www.geneious.com). Extracted products were also used to amplify and genotype 15 microsatellite loci (SST-A6-I, SST-A6-II, SST-B11, SST-C3, and SST-G945; SalE14, Sal29, SalE12, SalE7, SalE5, SalE2, SalE06, Sal3, and SalE846) distributed in five optimized multiplexes (see47 for details about the protocol optimisation and genotyping).
Our individual-based approach, together with some degree of continuous phenotypic variation in the contact zones, prevent us from grouping samples into a priori populations or genetic units, and thus from assessing accurately Hardy–Weinberg equilibrium (HWE) and linkage equilibrium (LE). However, previous phylogeographic and population genetic studies using the same set of markers33,38,41,42,48,49,50 on Iberian populations of S. salamandra support the validity of these markers for population and landscape genetic analyses.
Phylogenetic lineages
We used the cytb sequences to perform coalescent-based Bayesian analyses in BEAST version 1.8.451 and identify the mitochondrial lineage (mtDNA) of each sample, which was then used to estimate the extent of cyto-nuclear discordance across each contact zone. However, we note that mtDNA phylogenetic tree does not represent the true phylogenetic relationships among S. salamandra subspecies. In particular, S. s. fastuosa (with embedded S. s. gigliolii) was placed as the sister lineage to S. s. bernardezi and formed a well differentiated clade within S. salamandra according to phylogenomic data30. Yet, both S. s. gallaica and S. s. bejarae fall within the second clade and are sister lineages in both mtDNA and phylogenomic trees. We selected the optimal nucleotide substitution model (TrN) with JMODELTEST version 2.1.452, under the Bayesian information criterion (BIC). To estimate the time to the most recent common ancestor (TMRCA) of mtDNA lineages we used both a prior substitution rate and fossil calibrations. We set the divergence rate for cytb estimated in a 2.7% Myr for Mediterranean amphibians53. We used a normal distribution with a mean of 0.0134 and SD of 0.0009. We also used two calibration points (divergence date estimates) based on the split between Chioglossa and Mertensiella (mean: 19.4; lower and upper 95% confidence intervals (CIs) of 14.2 and 27.5, respectively), and the split between Lyciasalamandra and Salamandra (mean: 17.59; 95% CIs 11.24 and 24.82). These estimates were obtained from a two-step protocol based on a combination of primary fossil calibrations and secondary calibrations from a multilocus nuclear tree on a multilocus mitochondrial tree53. However, we avoided the use of internal nodes provided within Salamandra due to some incorrect phylogenetic relationships at both inter-specific (e.g. S. algira) and intra-subspecific (e.g. S. s. morenica) levels30. We obtained cytb sequences from GenBank for Chioglossa lustitanica (AF329314), Mertensiella caucasica (EU880319), Lyciasalamandra flavimembris (EU880318) and Lyciasalamandra atifi (AF154053), which were added to our cytb alignment. We performed three independent runs using an uncorrelated relaxed clock and a constant population size model as the coalescent tree prior, with a total of 100 million generations (burn-in: 10%). We combined tree files of all runs using the software LOGCOMBINER version 1.7.5, and verified parameter convergence by examining the effective sample sizes (ESSs) in TRACER version 1.6. We obtained a maximum clade credibility summary tree with Bayesian posterior probabilities (BPP) for each node using TREEANNOTATOR version 1.8.4, and edited the resulted tree in FIGTREE version 1.4.3 (http://tree.bio.ed.ac.uk/software/figtree). We ran BEAST analyses in the CIPRES Science Gateway54. We estimated genetic distances (uncorrected p-distances) between subspecies in GENEIOUS 11.1.4 (www.geneious.com).
Genetic structure and hybridization analysis
We first used STRUCTURE 2.3.455 to assess the individual genetic membership coefficient (q-value) for each subspecies in each contact zone. Because we were not interested in determining the genetic structure of each study area but rather assigning individuals to one of the two co-occurring subspecies, we ran STRUCTURE for K = 2 (two clusters). We ran 10 replicates and used the correlated allele frequency and admixture models, without location prior, and with a burn-in period of 100,000 iterations followed by 1,000,000 iterations. We combined the results from the 10 replicate runs using CLUMPP56. To complement STRUCTURE, we used the spatially explicit method of spatial Principal Component Analyses (sPCA)57, which does not assume HWE or LE expectations. To identify if the sPCA scores were able to assess global (patches and clines where neighbours tend to be similar) and local (strong genetic differences between neighbours) structure, we used a Delaunay triangulation connection network and two separate Monte Carlo tests with 100 permutations.
Because each individual has two q-values (one for each of K=2 clusters), we considered individuals as admixed when showing low levels of coefficient membership for either of the two clusters (0.1 < q-value < 0.9). Then, we used these potential hybrids to estimate the probability of assignment (Z) of early-generation hybrid or backcrossed individuals between subspecies within each contact zone in NEWHYBRIDS v.1.158,59. We used six classes in each analysis: pure individuals of each subspecies (parental classes), first generation hybrid (F1), second-generation hybrid (F2), and first-generation backcrosses to each subspecies. To increase the power to detect hybrids, we also assigned all individuals with > 90% (q-value > 0.9) estimated ancestry in STRUCTURE from a single subspecies to the appropriate pure parental category using the ‘z’ option. We performed three independent runs per contact zone and we set 1 million generations with a burn-in of 10% per run. We used two random starting values in each NEWHYBRIDS run, and the “Jeffreys-like” prior for the estimates of θ (i.e., allele frequencies) and π (i.e., mixing proportions). We used a posterior probability threshold of P ≥ 0.760 to assign potential hybrids identified in STRUCTURE to each of the six classes.
Landscape genetics analysis
The workflow for the landscape genetic analyses is summarised in Fig. 5 and detailed as follows.
Landscape genetics analyses workflow used to derive optimized variables representing the different isolation models (IBD, IBR, and IBE). Two parameters of a logistic curve (curvature and inflection point) were applied to the isolation by distance matrix and optimized with genetic algorithms. The objective of optimization is the maximization of the logLikelihood from a GLS regression. Note that this workflow is for a single genetic distance type, thus, it was executed twice for the RQG and RTRI genetic distances. The figure was created with Inkscape software (version 1.0.2; https://inkscape.org/).
Genetic distance matrices
The use of individual-based distances is recommended to avoid potential violations of the assumption of discrete populations that can lead to erroneous estimates of genetic parameters61. We used COANCESTRY62 to calculate pairwise relatedness matrices between individuals in each studied contact zone. First, we used the relatedness coefficient from Queller & Goodnight (1989) (RQG)63, which does not assume Hardy–Weinberg equilibrium. We also used the triadic likelihood relatedness (RTRI) estimate64, which calculates pairwise relatedness between two individuals while using a third one as control, thus reducing the identical by descent error rate. The matrices were linearly scaled to the range between 0 and 1, where higher values indicate higher similarity. To obtain an indication of dissimilarity, we found the complement of each matrix element by subtracting to 1.
Spatial data
We evaluated the three main environmental features that can influence spatial connectivity and gene flow of fire salamanders across the study region: elevation, habitat and climate. Elevation data was obtained from the Spanish Instituto Geográfico Nacional (https://www.ign.es) at a 5 m resolution and aggregated to a 250 m resolution for downstream analyses. We produced two variables using a different aggregating function: (1) the average to represent mean elevation at each 250 m pixel, and (2) the standard deviation as a representation of the topographic heterogeneity. This collection of two variables is hereafter referred to as elevation data. We used the Normalized Difference Vegetation Index (NDVI) as a continuous surrogate of habitat availability. The NDVI imagery was obtained with a 16-day period covering a range of 10 years (from January 2009 to December 2018) that includes the field work period. The MOD13Q1 product was downloaded from the EarthData website (https://earthdata.nasa.gov/) with a spatial resolution of 250 m and 16-day temporal resolution, resulting in a time-series with 230 measurement layers. To provide summaries of the 16-day NDVI for a few variables, we processed the data with a harmonic regression and extracted three spatial coefficients representing the periodic behaviour in each pixel as continuous surrogates of local habitat65. The collection of three variables is hereafter referred as NDVI data. All data was processed in R with packages rgdal (v.1.5.8), raster (v.3.3.13), TSA (v.1.2.1) and zoo (v.1.8.8). Climate data was obtained from the CHELSA database (https://chelsa-climate.org/66) as 19 bioclimate variables at 30” (~ 1 km) spatial resolution. We further summarized the climate data by means of a PCA after centering and scaling. Resulting spatial components from PCA are hereafter referred to as the climate data. We projected all spatial data to the WGS84 coordinate system.
Isolation by distance, environment and resistance matrices
We generated geographic, environmental and resistance distance matrices to test for the dominant model of isolation by distance (IBD), environment (IBE), or resistance (IBR) in each contact zone. We produced IBD matrices by calculating Euclidean geographic distances between genotyped individuals in each contact zone. We transformed coordinates to the ETRS89 Lambert Azimuthal Equal-Area projection before calculating the straight line distance between sampling localities.
To account for environmental differentiation between sampling locations, we calculated one IBE matrix based on climate (IBECLIM) and another based on NDVI (IBENDVI) for each contact zone (six IBE matrices in total; see Fig. 5). The IBE matrices were calculated as the Euclidean distance in the environmental space6,67. As such, we extracted at the locations of samples the value of the PCA components from climate data and calculated the distances between them. We followed the same procedure with NDVI data.
We generated IBR matrices in relation to descriptive categories: elevation (IBRALT), climate (IBRCLIM), and NDVI (IBRNDVI). We used a common dataset combining the presence locations from the three contact zones to determine the usage of each variable in each category. Pixel duplicates were removed independently in each category due to the different spatial resolution. We extracted variable data at the presence locations and calculated a vector of average values per variable and the covariance matrix. We used this information to combine the different variables by means of a multivariate normal distribution, generating a map representing the likelihood of finding a presence for each variable category. Hence, this value is related to the usage of the multidimensional space of each combination of variables by the species. We used these maps as a resistance surfaces, after a minmax scaling, in order to generate resistance distances with the ‘commuteDistance’ function in ‘gdistance’ R package (v.1.3.6).
Optimizations and model selection
Parameter optimization procedures are common in landscape genetic studies68,69,70,71. These optimizations aim at finding find the best combination of parameters describing a species’ environmental preferences to a resistance surface that has the best fit with genetic data. Often the aims of the optimization procedure are the parameters of a linear combination of habitat and other niche descriptors by maximizing a value relating the derived resistance distances to the genetic distances68,70. Optimizations algorithms have also been used to find the best parameters of a transformation function applied to the resistance distances69,71. Usually only the matrix referring to IBR is optimized, hence, it has an advantage over other competing models under a test of the isolation mechanism that better explains genetic divergence. Here we optimized the parameters of a logistic function relating each matrix produced (IBD, IBE and IBR) to the genetic distance matrices in each contact zone. It is expected that genetic differentiation increases with geographical distance. Yet, due to limited movement and finite population sizes and genomes, this increase cannot occur indefinitely and tends to plateau72. The same reasoning can be applied to the other models, including resistance and environmental distances. We optimized a logistic function with two parameters (curvature and inflection point), allowing for a near linear arrangement with original distances to a full logistic relation with steep transition. In the latter case, larger distances tend to plateau after the inflection point thus allowing a scenario where genetic differentiation accumulates until a maximum limit. We performed optimizations with genetic algorithms using the R package GA (v.3.2) to maximize the logLikelihood of a Generalized Least Squares (GLS) regression using the genetic matrix as a dependent variable and an isolation matrix as an independent variable. Data in pairwise distance matrices show high degrees of non-independence which hampers the direct application of regression-based algorithms. The GLS regression incorporates correlation structures informing about the lack of independence between samples. We used a maximum-likelihood population effects (MLPE) model that generates such correlation structure from pairwise data71. We used the package nlme (v.3.1.140) for GLS with corMLPE (v.0.0.3, https://github.com/nspope/corMLPE). All optimizations were done with a maximum of 1000 iterations or 200 runs without change of the optimization score. All isolation model matrices were min–max scaled to the range 0–1 in order to facilitate the comparison of the optimized parameters.
We used the Akaike Information Criterion (AIC) to rank univariate models and to eliminate those isolation models that were contributing less to the overall fit of the regression (Table 2). Models with ΔAIC lower or equal to 2 were assumed to be equivalent to best model. By comparing only univariate models and the null model, we do not use competing scenarios of resistance in the same model.
Results
Phylogenetic analysis
We obtained a total of 302 sequences after discarding sequences of poor quality and trimming some sequences (from ca. 1070 bp to 350 bp). Phylogenetic analyses identified two main monophyletic (Posterior Probability > 0.95) clusters within the three studied contact zones: one includes seven well supported mtDNA lineages of putative pueriparous S. s. bernardezi, while the second clade includes three monophyletic lineages of the three focal larviparous subspecies (S. s. gallaica, S. s. bejarae and S. s. fastuosa) (Fig. 1). According to the cytb dating approach, the split between S. s. bernardezi and the monophyletic clade including S. s. gallaica, S. s. bejarae and S. s. fastuosa is estimated at 1.3 Mya (0.9–1.7) with an earlier diversification within S. s. bernardezi (ca. 1 Mya [0.8–1.4]) compared to the S. s. gallaica-bejarae-fastuosa clade (ca. 0.3 Mya [0.15–0.4]).
Nuclear genetic structure, cyto-nuclear discordances and hybridization
After discarding individuals with missing data > 10% in the microsatellite dataset, we obtained a fully comparative mtDNA-nuDNA datasets for a total of 302 samples (127, 68 and 107 from NG, CM and AC, respectively).
STRUCTURE recovered two well defined genetic groups in each contact zone that largely match the distribution of each mitochondrial lineage (Figs. 2, 3, 4). The first eigenvalue of sPCA analyses confirmed the genetic structure found in STRUCTURE, which divided the data into two groups for each contact zone (Supplementary Figs. S2–S4). We found a significant global structure in each dataset but not significant support for a local structure in sPCA.
We found relatively similar patterns of hybridization in the contact zones of NG and AC (Table 1), in which, relatively high proportions of admixed individuals and cyto-nuclear discordances were inferred. Specifically, for NG, from the 127 analysed salamanders, a total of 92 individuals (72%) were assigned either to the subspecies S. s. gallaica (49 salamanders; 38%) or S. s. bernardezi (43 salamanders; 34%), while the remaining 35 salamanders (28%) were inferred as potential hybrids (0.1 ≤ q-value ≤ 0.9 in STRUCTURE). Among these potential hybrids, 29 salamanders were labelled as F2 hybrids in NEWHYBRIDS. Similarly, in AC, from the 107 genotypes, 57 salamanders (53%) were assigned either to the subspecies S. s. fastuosa (16 individuals; 15%) or S. s. bernardezi (41 salamanders; 38%). The remaining 50 salamanders comprised potential hybrids (47%), from which, 38 individuals were categorised as F2 hybrids, two salamanders as pure S. s. bernardezi, and two others as S. s. fastuosa in NEWHYBRIDS. Unlike the latter two contact zones, CM was characterized by a very low proportion of admixed individuals. From the 68 studied salamanders, a total of 65 individuals (96%) were assigned either to the subspecies S. s. bejarae (29 individuals; 43%) or S. s. bernardezi (36 salamanders; 53%), while the remaining three salamanders (4%) that were inferred as potential hybrids were later assigned as pure S. s. bejarae in NEWHYBRIDS. Furthermore, the number of cyto-nuclear discordances was higher for NG and AC (19% and 16%, respectively) than for CM (6%).
Landscape genetics analyses
The PCA of climate data showed the first three components accounted for 91.4% of total variation. Optimizations of IBD and IBE models showed a high degree of success, with the ΔAIC of the optimized model showing values higher than 15 compared to the non-optimized version (see Supplementary Table S2). The IBR model optimization showed a differential success depending on the contact zone. In the contact zone NG, all IBR models achieved a ΔAIC lower than 4.5, revealing a lack of effect of the optimization. A single IBD univariate model was selected as the best model based on ΔAIC values for the contact zones of NG and AC, independently of the genetic matrix used (Table 2). The contact zone of CM comprises a more complex scenario because univariate models indicate a dominant action of IBR in addition to IBD, particularly IBRCLIM and IBRNDVI for RQG genetic distances, and IBRCLIM with IBD for RTRI (Table 2). The combinations of the best models suggest not only a strong effect of IBD in all contact zones, but also a strong effect of habitat and climate resistance across the CM contact zone. The relationship between pairwise genetic distance (for both RQG and RTRI) and pairwise geographic and environmental distances for each contact zone are represented in Supplementary Figs. S5 and S6.
Discussion
The degree of reproductive isolation is expected to be larger between lineages showing differentiation in parity modes73, morphology74 or behaviour75, though time since divergence also enhances reproductive barriers76. Our fine-scale sampling and molecular approach contributes to an accurate characterisation of the location and spatial extent of the three studied contact zones, where the morphologically29 and phylogenetically30 divergent pueriparous S. s. bernardezi meets three larviparous subspecies (S. s. gallaica, S. s. bejarae, and S. s. fastuosa). We detected clear signs of gene flow and introgression in two out of three regions, thus supporting the hypothesis of incomplete reproductive isolation between these highly divergent S. salamandra morphotypes. Despite the extensive patterns of introgression and admixture in two contact zones (NG and AC), the Cantabrian Mountains (CM) likely reduce gene flow between the larviparous S. s. bejarae and pueriparous S. s. bernardezi. This study suggests the combined effect of landscape and climate barriers shape patterns of hybridization, thus highlighting the need of genetic studies across multiple and heterogeneous contact zones to better delineate the spatial patterns of gene flow and hibridisation between non-reproductively isolated taxa.
Geographic range and spatial extent of Salamandra salamandra contact zones
Previous phylogeographic studies suggested the eastern Galician mountains comprise a barrier to gene flow between S. s. gallaica and S. s. bernardezi31,32. The high proportion of admixed individuals inferred from our genetic analyses demonstrates the existence of a contact zone between western S. s. bernardezi and northern S. s. gallaica populations within a relatively homogenous landscape in northern Galicia (NG), thus indicating previous works were likely biased due to an insufficient sampling in this region. We detected a north–south contact zone between S. s. bernardezi and S. s. gallaica located ca. 10–30 kms from the northern Galician coast (Fig. 2). The sampled salamanders in NG were largely characterised by a mixture of phenotypes typical of S. s. gallaica and S. s. bernardezi, as previously documented by morphological studies in this region35,77. The coloration pattern across this area varied considerable. Most admixed individuals showed a disrupted striped colouration, resembling S. s. bernardezi phenotype, but also showed intermediate colouration patterns with both stripes and blotches. These admixed individuals also showed intermediate body sizes, and variable snout shapes, thus hindering subspecies assignment and supporting the existence of a hybrid zone. Additionally, the presence of females with a mixed reproductive mode (i.e. females give birth to both metamorphosed juveniles and aquatic larvae35; unpublished data) corroborates the presence of a hybrid zone in NG.
Patterns of genetic structure confirmed a negligible hybrid zone between southern S. s. bernardezi and northern S. s. bejarae populations across the Cantabrian Mountains (CM; Fig. 3). Contrary to NG, the contact zone in CM shows an abrupt change of phenotypes between populations inhabiting southern slopes, where fire salamanders are larviparous, large, blotched, with a relatively pointed snout, and populations located on northern slopes, in which individuals exhibit pueriparity, are small, striped and often present rounded snouts29,31,77.
Similar to NG, we also found a high proportion of admixed individuals along the Asturias-Cantabria border (AC), where eastern populations of S. s. bernardezi hybridize with S. s. fastuosa along lowlands (west–east corridors) located between mountain chains (Sierra de Cuera and Picos de Europa; Fig. 4). This area coincides with a secondary contact between amphibian lineages with Atlantic climate affinity (Lissotriton helveticus78; Rana parvipalmata-R. temporaria79). The deep divergence found in Rana points to an old split (4 Mya) and a narrow hybrid zone (25 km), which led the authors to conclude these two lineages represent distinct species. Salamandra, however, shows a more recent split (late Pleistocene). Despite this study showing the existence of a contact zone between these subspecies, previous phylogeograhic work placed this hybrid zone farther east to the Pyrenees30,31. The identification of the contact zone between S. s. bernardezi and S. s. fastuosa is not possible at the morphological level because S. s. fastuosa appears to be phenotypically indistinguishable from S. s. bernardezi. We have also detected females with mixed reproductive modes across AC, and also cases of larviparity and pueriparity within the S. s. bernardezi and S. s. fastuosa ranges, respectively, which again supports the presence of a relatively broad hybrid zone in this area. Nevertheless, future studies should be conducted to accurately delimit the range of S. s. fastuosa and the level of introgression across its range.
Genetic introgression and gene flow between S. salamandra subspecies
Patterns of cyto-nuclear discordances are common in animals80, including salamanders. Conflicting geographic patterns between mitochondrial and nuclear genetic markers have been reported across the S. salamandra range (i.e. S. s. bernardezi—S. s. fastuosa31; S. s. bernardezi—S. s. gallaica38; S. s. bejarae—S. s. almanzoris48; S. s. gigliolli—S. s. terrestris81, and between subspecies in its sister species, S. algira82. Similarly, we found cyto-nuclear discordances across the three studied contact zones, though these were higher in NG and AC than in CM.
Despite the marked genetic30 and phenotypic (i.e., colour patterns35, morphology29, and reproductive modes27,83) differences between S. s. gallaica and S. s. bernardezi, we found extensive gene flow across the entire NG, thus confirming the lack of reproductive isolation between these divergent taxa, and the widespread presence of admixed phenotypes in this contact zone. We found an asymmetric pattern of cyto-nuclear discordances between both lineages. The majority of introgressed individuals exhibited a S. s. bernardezi mtDNA, which could suggest a northward expansion by S. s. gallaica followed by a mitochondrial capture of S. s. bernardezi. This process is potentially in agreement with the male-biased dispersal hypothesis, in which, through population expansions, fire salamander males may disperse longer distances than females50,84 and move into their neighbouring subspecies range, thus capturing the local mitochondrial lineage after a few generations. Such a process was suggested as a potential driver of cyto-nuclear discordances among other S. salamandra subspecies31,48,81. However, S. s. gallaica mtDNA reached the northern Galician coast and it is known that S. s. gallaica mtDNA is present at low frequencies within the range of S. s. bernardezi38. Whether this is the result of a recent expansion of both males and females of S. s. gallaica into the range of S. s. bernardezi or a male-biased colonization of the latter into a historical and wider range of the former remains an open question, with important implications for the mid- and long-term viability of S. s. bernardezi, which warrants future research.
In CM, we found very few cases of genetic introgression at both windward and leeward sides of the Cantabrian Mountains, but no signs of hybridization. The negligible signs of admixture and hybridization between the larviparous and large S. s. bejarae and the pueriparous and small S. s. bernardezi across the CM are likely due to unfavourable environmental conditions of high-elevation areas (see next section). These conditions likely preclude the presence or reduce population densities and activity of fire salamanders, which in turn hinder dispersal across complex topographic areas. However, valleys (1400–1700 m a.s.l.) oriented in a north–south axis might act as mountain passages allowing sporadic dispersal events of fire salamander across the CM.
Like NG, we found high levels of gene flow and hybridization across AC and, consequently, absence of endogenous reproductive barriers between S. s. bernardezi and S. s. fastuosa. This is an expected result because of their phenotypic similarity and shared recent evolutionary histories. The relatively high proportion of cyto-nuclear discordances found for both subspecies comprises another pattern in common with NG, although the underlying drivers responsible for this genetic pattern are likely different. Selection of traits encoded in the nuclear genome could have boosted such discordances in AC along the expansion wave. Nuclear phylogenomic analyses place S. s. fastuosa as the sister taxon of S. s. bernardezi, yet S. s. fastuosa mtDNA falls within the S. s. gallaica-bejarae clade30. This genetic discordance fits a demographic scenario where selected traits (i.e. colouration pattern, snout shape and reproductive strategy) encoded by nuclear genes in S. s. bernardezi spread eastwards into the S. s. fastuosa area during range expansions of favourable climatic cycles in the Pleistocene31. It is possible that only the original mitochondrial legacy of S. s. fastuosa was maintained though historical mitochondrial capture when both lineages introgressed, a process that has been suggested for Rana temporaria in this region79. Alternatively, S. s. fastuosa could first diverge from S. s. bernardezi, and then followed by a process of historical mitochondrial capture of S. s. gallaica-bejarae. Further sampling across this region and the use of genomic data and demographic models will allow testing the above contrasting hypotheses.
Environmental barriers shape gene flow between S. salamandra populations
Landscape genetic analyses revealed the IBD model is a strong predictor of genetic differentiation in the NG and AC contact zones and, to a lesser extent, in CM. The marked genetic structure and absence of hybrids across the Cantabrian Mountains suggests this mountain chain is a major barrier to dispersal and gene flow between larviparous and pueriparous lineages, with mountain valleys probably allowing sporadic dispersal events. Interestingly, the observed pattern of genetic differentiation in CM could not be explained by the topographic complexity across this area, as we previously hypothesised, but it was influenced by the climate (IBRclim) and landcover (IBRndvi), albeit with some disagreement between the two genetic distance metrics used here. There is a strong environmental gradient acting between the southern and northern slopes of the Cantabrian Mountains, following the sharp transition between Atlantic and Mediterranean bioclimates, which t is reinforcing the effect that prevents rampant gene flow between S. s. bernardezi and S. s. bejarae along this CZ (see Supplementary Fig. S1). In addition, the climatic heterogeneity associated with the mountain ridges (e.g. higher wind exposure, colder climate, shorting of the growing season, or rarefaction of surface water bodies) often act as impermeable barriers to fire salamanders38,85, other salamander species86,87,88,89, and anurans90. On the other hand, the relatively smooth topography across NG and consequent softer climate transitions in the west–east corridors (i.e. lowlands) across AC likely facilitate salamander dispersal.
Other landscape barriers are likely affecting gene flow across the three studied contact zones. For instance, the reduction and conversion of woodlands to mosaics of agricultural fields, urban settlements, and plantations of exotic trees have been shown to hinder dispersal and enhance genetic differentiation among fire salamander populations38,41, though the degree of fragmentation observed in NG and AC appears to be insufficient to prevent hybridization at the spatial scale of this study. Water courses are also known to affect gene flow in S. Salamandra, particularly, in S. s. bernardezi due to its terrestrial life cycle91. Two main rivers (Deva and Nansa) cross the AC hybrid zone in a north–south axis and their locations coincide with a phylogeographic break found in other amphibians78,79. Although these rivers might have a role in reducing gene flow between S. s. bernardezi and S. s. fastuosa, the observed patterns of admixture between these lineages suggest they could act as semi-permeable barriers to gene flow. Also, larviparous individuals would have a greater advantage to overcome these geographic barriers during the larval stage91, but our data prevent us from making robust inferences about crossing rates per reproductive mode.
Besides the aforementioned environmental factors, we cannot discount that other phenomena may be at play here. Adaptation to ecological environments has been proposed to explain behavioural (pond vs. stream reproduction92, but see93) and morphological (colour polymorphism94) differentiation in fire salamanders, and it may be a common underlying mechanism explaining the elevated intra-specific phenotypic variation that characterizes this species27,29,95. Our study did not identify divergent natural selection (i.e. support for an IBE scenario) as a driver of the distinct morphological and reproductive types across northern Iberian S. salamandra populations. Although environmental differentiation observed between southern and northern slopes of the Cantabrian Mountains (i.e. climate and habitat, see Supplementary Fig. S1) resulted in a negligible effect from the IBE model, we cannot discount possible latent adaptive divergence to the different ecological environments found at each side of this mountain chain. We note that the other contact zones also have strong gradients, yet, they occur along the genetic transition. This implies that both subspecies are in contact with the same environmental gradient available in each contact zone. In the Cantabrian Mountains, however, the contact zone coincides with a marked change of the environmental gradient, producing favourable conditions for putative adaptive divergence. These mountains constitute the clearest contact zone where subspecies occupy different environmental niches. While NG and AC are located in a heterogeneous landscape (mosaic of eucalypts plantations, and agricultural bocage in lowland areas, and native forests and scrublands in mid-elevation areas), with a soft change in climatic conditions, in the CM, the wetter northern slopes dominated by Atlantic deciduous forests strongly contrast with the dryer southern slopes exhibiting vegetation typical of Mediterranean ecosystems96. Hence, fire salamanders at both sides of the CM likely experience different environmental regimes that may influence their evolutionary trajectories.
The barrier effect imposed by the Cantabrian Mountains, together with a potential ecological adaptation to the local habitat and climate present in the northern and southern slopes of this mountain chain, may also contribute to the reduced levels of admixture and introgression observed between S. s. bernardezi and S. s. bejarae, which ultimately can lead to speciation events. However, considering the ongoing gene flow between S. s. bernardezi and other S. salamandra subspecies in environmentally homogeneous contact zones (e.g. NG and AC), our study reinforces the intra-specific polymorphic nature of this species at various phenotypic levels. Finally, we cannot neglect potential historical differences underlying the formation and maintenance of these contact zones. These differences may involve an older and well established hybrid zone in northern Galicia (S. s. gallaica–S. s. bernardezi), and a recent secondary contact zone where population edges are still expanding across the CM (S. s. bejarae–S. s. bernardezi).
Conclusions
Our study shows that patterns of hybridization between S. salamandra subspecies are highly heterogeneous across space, thus emphasizing the role of environmental factors in shaping patterns of hybridization and reproductive isolation. This is best illustrated by the Cantabrian Mountains, which likely comprise an effective barrier to gene flow between the pueriparous S. s. bernardezi and larviparous S. s. bejarae. Despite the fact that prezygotic and postzygotic barriers were not detected in our study, it is unclear whether hybridization among these subspecies negatively affects survival and fertility of hybrids. This study also has important conservation and taxonomic implications, as the elevation of intra-specific lineages to species level is often based on the reduction of gene flow found between lineages across a small area of their entire contact zone, thus neglecting the environmental heterogeneity, and putative different outcomes of hybridization. Future multidisciplinary studies (integrating phenotypic, behavioural, ecological, and genomic-wide data) are needed to evaluate the potential genomic consequences of hybridization in the fire salamander, the spatio-temporal dynamics of these hybrid zones (i.e. location and spatial extent through time), and the potential role of ecological divergence in the Cantabrian Mountains9,10,48.
Data availability
The dataset containing microsatellite genotype data, environmental layers, input files, and the R code used to perform optimisation analyses are available upon request. Sequences generated for the mtDNA cytb fragment have been submitted to GenBank.
References
Abbott, R. J., Barton, N. H. & Good, J. M. Genomics of hybridization and its evolutionary consequences. Mol. Ecol. 25, 2325–2332 (2016).
Harrison, R. G. & Larson, E. L. Heterogeneous genome divergence, differential introgression, and the origin and structure of hybrid zones. Mol. Ecol. 25, 2454–2466 (2016).
Gompert, Z., Mandeville, E. G. & Buerkle, C. A. Analysis of population genomic data from hybrid zones. Annu. Rev. Ecol. Evol. Syst. 48, 207–229 (2017).
Jiggins, C. D. & Mallet, J. Bimodal hybrid zones and speciation. Trends Ecol. Evol. 15, 250–255 (2000).
Doebeli, M. & Dieckmann, U. Speciation along environmental gradients. Nature 421, 259–264 (2003).
Wang, I. J. & Bradburd, G. S. Isolation by environment. Mol. Ecol. 23, 5649–5662 (2014).
Tarroso, P., Pereira, R. J., Martínez-Freiría, F., Godinho, R. & Brito, J. C. Hybridization at an ecotone: Ecological and genetic barriers between three Iberian vipers. Mol. Ecol. 23, 1108–1123 (2014).
Newman, C. E. & Rissler, L. J. Phylogeographic analyses of the southern leopard frog: The impact of geography and climate on the distribution of genetic lineages vs. subspecies. Mol. Ecol. 20, 5295–5312 (2011).
Smith, K. L. et al. Spatio-temporal changes in the structure of an Australian frog hybrid zone: A 40-year perspective. Evolution 67, 3442–3454 (2013).
Visser, M., Leeuw, M. D., Zuiderwijk, A. & Arntzen, J. W. Stabilization of a salamander moving hybrid zone. Ecol. Evol. 7, 689–696 (2017).
Carneiro, M. et al. Steep clines within a highly permeable genome across a hybrid zone between two subspecies of the European rabbit. Mol. Ecol. 22, 2511–2525 (2013).
Gompert, Z., Parchman, T. L. & Buerkle, C. A. Genomics of isolation in hybrids. Philos. Trans. R. Soc. B 367, 439–450 (2012).
Zieliński, P. et al. Differential introgression across newt hybrid zones–evidence from replicated transects. Mol. Ecol. 28, 4811–4824 (2019).
Hewitt, G. M. Quaternary phylogeography: The roots of hybrid zones. Genetica 139, 617–638 (2011).
Naciri, Y. & Linder, H. P. The genetics of evolutionary radiations. Biol. Rev. 95, 1055–1072 (2020).
Kearns, A. M. et al. Genomic evidence of speciation reversal in ravens. Nat. Commun. 9, 1–13 (2018).
Butlin, R. Speciation by reinforcement. Trends Ecol. Evol. 2, 8–13 (1987).
Arntzen, J. W., de Vries, W., Canestrelli, D. & Martínez-Solano, I. Hybrid zone formation and contrasting outcomes of secondary contact over transects in common toads. Mol. Ecol. 26, 5663–5675 (2017).
Zamudio, K. R., Bell, R. C. & Mason, N. A. Phenotypes in phylogeography: Species’ traits, environmental variation, and vertebrate diversification. Proc. Natl. Acad. Sci. 113, 8041–8048 (2016).
Devitt, T. J., Baird, S. J. & Moritz, C. Asymmetric reproductive isolation between terminal forms of the salamander ring species Ensatina eschscholtzii revealed by fine-scale genetic analysis of a hybrid zone. BMC Evol. Biol. 11, 245 (2011).
Melo, M. C., Salazar, C., Jiggins, C. D. & Linares, M. Assortative mating preferences among hybrids offers a route to hybrid speciation. Evolution 63, 1660–1665 (2009).
Cornetti, L. et al. Reproductive isolation between oviparous and viviparous lineages of the Eurasian common lizard Zootoca vivipara in a contact zone. Biol. J. Linn. Soc. 114, 566–573 (2015).
Rafati, N. et al. A genomic map of clinal variation across the European rabbit hybrid zone. Mol. Ecol. 27, 1457–1478 (2018).
Shipilina, D., Serbyn, M., Ivanitskii, V., Marova, I. & Backström, N. Patterns of genetic, phenotypic, and acoustic variation across a chiffchaff (Phylloscopus collybita abietinus/tristis) hybrid zone. Ecol. Evol. 7(7), 2169–2180 (2017).
Grabenstein, K. C. & Taylor, S. A. Breaking barriers: Causes, consequences, and experimental utility of human-mediated hybridization. Trends Ecol. Evol. 33(3), 198–212 (2018).
Coates, D. J., Byrne, M. & Moritz, C. Genetic diversity and conservation units: Dealing with the species-population continuum in the age of genomics. Front. Ecol. Evol. 6, 165 (2018).
Velo-Antón, G., Santos, X., Sanmartín-Villar, I., Cordero-Rivera, A. & Buckley, D. Intraspecific variation in clutch size and maternal investment in pueriparous and larviparous Salamandra salamandra females. Evol. Ecol. 29(1), 185–204 (2015).
Beukema, W., Nicieza, A. G., Lourenço, A. & Velo-Antón, G. Colour polymorphism in Salamandra salamandra (Amphibia: Urodela), revealed by a lack of genetic and environmental differentiation between distinct phenotypes. J. Zool. Syst. Evol. Res. 54(2), 127–136 (2016).
Alarcón-Ríos, L., Nicieza, A. G., Kaliontzopoulou, A., Buckley, D. & Velo-Antón, G. Evolutionary history and not heterochronic modifications associated with viviparity drive head shape differentiation in a reproductive polymorphic species, Salamandra salamandra. Evol. Biol. 47(1), 43–55 (2020).
Burgon, J. D. et al. Phylogenomic inference of species and subspecies diversity in the Palearctic salamander genus Salamandra. Mol. Phylogenet. Evol. 157, 107063 (2021).
García-París, M., Alcobendas, M., Buckley, D. & Wake, D. Dispersal of viviparity across contact zones in Iberian populations of Fire salamanders (Salamandra) inferred from discordance of genetic and morphological traits. Evolution 57(1), 129–143 (2003).
Velo-Antón, G., García-París, M., Galán, P. & CorderoRivera, A. The evolution of viviparity in holocene islands: ecological adaptation versus phylogenetic descent along the transition from aquatic to terrestrial environments. J. Zool. Syst. Evol. Res. 45(4), 345–352 (2007).
Velo-Antón, G., Zamudio, K. R. & Cordero-Rivera, A. Genetic drift and rapid evolution of viviparity in insular fire salamanders (Salamandra salamandra). Heredity 108(4), 410–418 (2012).
Uotila, E., Díaz, A. C., Azkue, I. S. & Rubio Pilarte, X. Variation in the reproductive strategies of Salamandra salamandra (Linnaeus, 1758) populations in the province of Gipuzkoa (Basque Country). Munibe Cienc. Nat. Nat. Zientziak 61, 91–101 (2013).
Galán, P. Viviparismo y distribución de Salamandra salamandra bernardezi en el norte de Galicia. Bol. Asoc. Herpetol. Esp. 18, 44–49 (2007).
Alcobendas, M., Dopazo, H. & Alberch, P. Geographic variation in allozymes of populations of Salamandra salamandra (Amphibia: Urodela) exhibiting distinct reproductive modes. J. Evol. Biol. 9(1), 83–102 (1996).
Alarcón-Ríos, L., Nicieza, A. G., Lourenço, A. & Velo-Antón, G. The evolution of pueriparity maintains multiple paternity in a polymorphic viviparous salamander. Sci. Rep. 10, 14744 (2020).
Lourenço, A., Gonçalves, J., Carvalho, F., Wang, I. J. & Velo-Antón, G. Comparative landscape genetics reveals the evolution of viviparity reduces genetic connectivity in fire salamanders. Mol. Ecol. 28(20), 4573–4591 (2019).
Velo-Antón, G., & Buckley, D. Salamandra común—Salamandra salamandra. in Enciclopedia Virtual de los Vertebrados Españoles (L.M. Carrascal, A Salvador, Eds.) (Museo Nacional de Ciencias Naturales, 2015). Retrieved from http://www.vertebradosibericos.org/anfibios/salsal.html
Cordero, A., Velo-Antón, G. & Galán, P. Ecology of amphibians in small coastal Holocene islands: Local adaptations and the effect of exotic tree plantations. Munibe 25, 94–103 (2007).
Antunes, B. et al. Combining phylogeography and landscape genetics to infer the evolutionary history of a short-range Mediterranean relict, Salamandra salamandra longirostris. Conserv. Genet. 19(6), 1411–1424 (2018).
Lourenço, A., Álvarez, D., Wang, I. J. & Velo-Antón, G. Trapped within the city: Integrating demography, time since isolation and population-specific traits to assess the genetic effects of urbanization. Mol. Ecol. 26(6), 1498–1514 (2017).
Landguth, E. L., Cushman, S. A., Murphy, M. A. & Luikart, G. Relationships between migration rates and landscape resistance assessed using individual-based simulations. Mol. Ecol. Resour. 10(5), 854–862 (2010).
Zhang, P., Papenfuss, T. J., Wake, M. H., Qu, L. & Wake, D. B. Phylogeny and biogeography of the family Salamandridae (Amphibia: Caudata) inferred from complete mitochondrial genomes. Mol. Phylogenet. Evol. 49(2), 586–597 (2008).
Hendrix, R., Hauswaldt, S., Veith, M. & Steinfartz, S. Strong correlation between cross-amplification success and genetic distance across all members of ‘True Salamanders’ (Amphibia: Salamandridae) revealed by Salamandra salamandra-specific microsatellite loci. Mol. Ecol. Resour. 10(6), 1038–1047 (2010).
Steinfartz, S., Kuesters, D. & Tautz, D. Isolation and characterization of polymorphic tetranucleotide microsatellite loci in the Fire salamander Salamandra salamandra (Amphibia: Caudata). Mol. Ecol. Notes 4(4), 626–628 (2004).
Álvarez, D., Lourenço, A., Oro, D. & Velo-Antón, G. Assessment of census (N) and effective population size (N e) reveals consistency of N e single-sample estimators and a high N e/N ratio in an urban and isolated population of fire salamanders. Conserv. Genet. Resour. 7(3), 705–712 (2015).
Antunes, B., Velo-Antón, G., Buckley, D., Pereira, R. & Martínez-Solano, I. Physical and ecological isolation contribute to maintain genetic differentiation between fire salamander subspecies. Heredity. https://doi.org/10.1038/s41437-021-00405-0 (2021).
Lourenço, A., Sequeira, F., Buckley, D. & Velo-Antón, G. Role of colonization history and species-specific traits on contemporary genetic variation of two salamander species in a Holocene island-mainland system. J. Biogeogr. 45(5), 1054–1066 (2018).
Lourenço, A., Antunes, B., Wang, I. J. & Velo-Antón, G. Fine-scale genetic structure in a salamander with two reproductive modes: Does reproductive mode affect dispersal?. Evol. Ecol. 32(6), 699–732 (2018).
Drummond, A. J., Suchard, M. A., Xie, D. & Rambaut, A. Bayesian phylogenetics with BEAUti and the BEAST 17. Mol. Biol. Evol. 29(8), 1969–1973 (2012).
Darriba, D., Taboada, G. L., Doallo, R. & Posada, D. jModelTest 2: More models, new heuristics and parallel computing. Nat. Methods 9(8), 772 (2012).
Ehl, S., Vences, M. & Veith, M. Reconstructing evolution at the community level: A case study on Mediterranean amphibians. Mol. Phylogenet. Evol. 134, 211–225 (2019).
Miller, M. A., Pfeiffer, W., & Schwartz, T. Creating the CIPRES Science Gateway for inference of large phylogenetic trees. in 2010 gateway computing environments workshop (GCE pp. 1–8) (2010).
Pritchard, J. K., Stephens, M. & Donnelly, P. Inference of population structure using multilocus genotype data. Genetics 155(2), 945–959 (2000).
Jakobsson, M. & Rosenberg, N. A. CLUMPP: A cluster matching and permutation program for dealing with label switching and multimodality in analysis of population structure. Bioinformatics 23(14), 1801–1806 (2007).
Jombart, T. adegenet: A R package for the multivariate analysis of genetic markers. Bioinformatics 24, 1403–1405 (2008).
Anderson, E. C. & Thompson, E. A. A model-based method for identifying species hybrids using multilocus genetic data. Genetics 160(3), 1217–1229 (2002).
Anderson, E. C. Bayesian inference of species hybrids using multilocus dominant genetic markers. Philos. Trans. R. Soc. B 363(1505), 2841–2850 (2008).
Shurtliff, Q. R., Murphy, P. J. & Matocq, M. D. Ecological segregation in a small mammal hybrid zone: Habitat-specific mating opportunities and selection against hybrids restrict gene flow on a fine spatial scale. Evolution 68(3), 729–742 (2014).
Shirk, A. J., Landguth, E. L. & Cushman, S. A. A comparison of individual-based genetic distance metrics for landscape genetics. Mol. Ecol. Resour. 17(6), 1308–1317 (2017).
Wang, J. COANCESTRY: A program for simulating, estimating and analysing relatedness and inbreeding coefficients. Mol. Ecol. Resour. 11(1), 141–145 (2011).
Queller, D. C. & Goodnight, K. F. Estimating relatedness using genetic markers. Evolution 43, 258–275 (1989).
Wang, J. Triadic IBD coefficients and applications to estimating pairwise relatedness. Genet. Res. 89, 135–153 (2007).
Estrada-Peña, A., Estrada-Sánchez, A. & de la Fuente, J. A global set of Fourier-transformed remotely sensed covariates for the description of abiotic niche in epidemiological studies of tick vector species. Parasit. Vectors 7(1), 302 (2014).
Karger, D. N. et al. Climatologies at high resolution for the earth’s land surface areas. Sci. Data 4, 170122 (2017).
Nosil, P., Egan, S. P. & Funk, D. J. Heterogeneous genomic differentiation between walking-stick ecotypes: “Isolation by adaptation” and multiple roles for divergent selection. Evolution 62, 316–336 (2008).
Graves, T. A., Beier, P. & Royle, J. A. Current approaches using genetic distances produce poor estimates of landscape resistance to interindividual dispersal. Mol. Ecol. 22(15), 3888–3903 (2013).
Peterman, W. E., Connette, G. M., Semlitsch, R. D. & Eggert, L. S. Ecological resistance surfaces predict fine-scale genetic differentiation in a terrestrial woodland salamander. Mol. Ecol. 23(10), 2402–2413 (2014).
Tarroso, P., Carvalho, S. B. & Velo-Antón, G. Phylin 2.0: Extending the phylogeographical interpolation method to include uncertainty and user-defined distance metrics. Mol. Ecol. Resour. 19(4), 1081–1094 (2019).
Peterman, W. E. ResistanceGA: An R package for the optimization of resistance surfaces using genetic algorithms. Methods Ecol. Evol. 9(6), 1638–1647 (2018).
Hutchison, D. W. & Templeton, A. R. Correlation of pairwise genetic and geographic distance measures: Inferring the relative influences of gene flow and drift on the distribution of genetic variability. Evolution 53(6), 1898–1914 (1999).
Horreo, J. L. et al. Genetic introgression among differentiated clades is lower among clades exhibiting different parity modes. Heredity 123(2), 264–272 (2019).
Sota, T. & Tanabe, T. Multiple speciation events in an arthropod with divergent evolution in sexual morphology. Proc. R. Soc. B 277(1682), 689–696 (2010).
Merrill, R. M., Van Schooten, B., Scott, J. A. & Jiggins, C. D. Pervasive genetic associations between traits causing reproductive isolation in Heliconius butterflies. Proc. R. Soc. B 278(1705), 511–518 (2011).
Singhal, S. & Moritz, C. Reproductive isolation between phylogeographic lineages scales with divergence. Proc. R. Soc. B 280(1772), 20132246 (2013).
Donaire, D. & Rivera, X. L. salamandra común Salamandra salamandra (Linnaeus, 1758) en el subcantábrico: Origen, dispersión, subspecies y zonas de introgresión. Bull. Soc. Catal. Herpetol. 23, 7–38 (2016).
Recuero, E. & García-París, M. Evolutionary history of Lissotriton helveticus: multilocus assessment of ancestral vs. recent colonization of the Iberian Peninsula. Mol. Phylogenet. Evol. 60(1), 170–182 (2011).
Dufresnes, C. et al. Are glacial refugia hotspots of speciation and cyto-nuclear discordances? Answers from the genomic phylogeography of Spanish common frogs. Mol. Ecol. 29, 986–1000 (2020).
Toews, D. P. & Brelsford, A. The biogeography of mitochondrial and nuclear discordance in animals. Mol. Ecol. 21(16), 3907–3930 (2012).
Bisconti, R., Porretta, D., Arduino, P., Nascetti, G. & Canestrelli, D. Hybridization and extensive mitochondrial introgression among fire salamanders in peninsular Italy. Sci. Rep. 8(1), 1–10 (2018).
Dinis, M. et al. Allopatric diversification and evolutionary melting pot in a North African Palearctic relict: The biogeographic history of Salamandra algira. Mol. Phylogenet. Evol. 130, 81–91 (2019).
Buckley, D., Alcobendas, M., García-París, M. & Wake, M. H. Heterochrony, cannibalism, and the evolution of viviparity in Salamandra salamandra. Evol. Dev. 9(1), 105–115 (2007).
Helfer, V., Broquet, T. & Fumagalli, L. Sex-specific estimates of dispersal show female philopatry and male dispersal in a promiscuous amphibian, the alpine salamander (Salamandra atra). Mol. Ecol. 21(19), 4706–4720 (2012).
Vörös, J. et al. Increased genetic structuring of isolated Salamandra salamandra populations (Caudata: Salamandridae) at the margins of the Carpathian Mountains. J. Zool. Syst. Evol. Res. 55(2), 138–149 (2017).
Dudaniec, R. Y., Spear, S. F., Richardson, J. S. & Storfer, A. Current and historical drivers of landscape genetic structure differ in core and peripheral salamander populations. PLoS ONE 7(5), e36769 (2012).
Richardson, J. L. Divergent landscape effects on population connectivity in two co-occurring amphibian species. Mol. Ecol. 21(18), 4437–4451 (2012).
Mulder, K. P., Cortes-Rodriguez, N., Campbell Grant, E. H., Brand, A. & Fleischer, R. C. North-facing slopes and elevation shape asymmetric genetic structure in the range-restricted salamander Plethodon shenandoah. Ecol. Evol. 9(9), 5094–5105 (2019).
Velo-Antón, G., Parra, J. L., Parra-Olea, G. & Zamudio, K. R. Tracking climate change in a dispersal-limited species: Reduced spatial and genetic connectivity in a montane salamander. Mol. Ecol. 22(12), 3261–3278 (2013).
Sánchez-Montes, G., Wang, J., Ariño, A. H. & Martínez-Solano, Í. Mountains as barriers to gene flow in amphibians: Quantifying the differential effect of a major mountain ridge on the genetic structure of four sympatric species with different life history traits. J. Biogeogr. 45(2), 318–331 (2018).
Figueiredo-Vázquez, C., Lourenço, A. & Velo-Antón, G. Riverine barriers to gene flow in a salamander with both aquatic and terrestrial reproduction. Evol Ecol https://doi.org/10.1007/s10682-021-10114-z (2021).
Czypionka, T., Goedbloed, D. J., Steinfartz, S. & Nolte, A. W. Plasticity and evolutionary divergence in gene expression associated with alternative habitat use in larvae of the European Fire Salamander. Mol. Ecol. 27(12), 2698–2713 (2018).
Arntzen, J. W. & van Belkom, J. ‘Mainland-island’population structure of a terrestrial salamander in a forest-bocage landscape with little evidence for in situ ecological speciation. Sci. Rep. 10(1), 1–15 (2020).
Burgon, J. D. et al. Functional colour genes and signals of selection in colour-polymorphic salamanders. Mol. Ecol. 29(7), 1284–1299 (2020).
Velo-Antón, G. & Cordero-Rivera, A. Ethological and phenotypic divergence in insular fire salamanders: Diurnal activity mediated by predation?. Acta Ethol. 20(3), 243–253 (2017).
González, T. E. D., & Penas, Á. The high mountain area of Northwestern Spain: The Cantabrian Range, the Galician-Leonese Mountains and the Bierzo Trench. In The vegetation of the Iberian Peninsula (pp. 251–321). (Springer, 2017).
Acknowledgements
This work was supported by National Funds through FCT – Foundation for Science and Technology (SALOMICS: PTDC/BIA-EVL/28475/2017); the authors also acknowledge funding from other FCT projects and FEDER funds through the Operational Programme for Competitiveness Factors – COMPETE (EVOVIV: PTDC/BIA-EVF/3036/2012; FCOMP-01-0124-FEDER-028325 and UIDB/500027/2020); and by National/FEDER funds through Ministerio de Economía y Competitividad (CGL2012-40246-C02-02). G.V.-A., A.L. and P.T. were supported by FCT (IF/01425/2014 and CEECIND/00937/2018, PD/BD/106060/2015, DL57/2016/CP1440/CT0008, respectively).
Author information
Authors and Affiliations
Contributions
G.V.-A. conceived the study. G.V.-A, P.G., A.N. and A.L. conducted field work. A.L. performed laboratory work. G.V.-A, A.L., and P.T. carried out data analysis. G.V.-A wrote the manuscript with contributions from all authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Velo-Antón, G., Lourenço, A., Galán, P. et al. Landscape resistance constrains hybridization across contact zones in a reproductively and morphologically polymorphic salamander. Sci Rep 11, 9259 (2021). https://doi.org/10.1038/s41598-021-88349-7
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-021-88349-7
This article is cited by
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.