Abstract
Primary biliary cholangitis (PBC) is a chronic cholestatic liver disease characterized pathologically by destruction of intrahepatic bile ducts. PBC is largely classified into three subtypes based on clinical course: (i) gradually progressive, (ii) portal hypertension, and (iii) hepatic failure. Previous studies have indicated that serum levels of the pro-inflammatory cytokine TNF-α, is elevated in PBC patients with fibrosis. Although the severity of cholangitis might also be related to the PBC subtype, its etiology has been unclear. Several studies have shown that microRNAs (miRNAs) demonstrate specific expression patterns in various diseases. In the present study, we evaluated miRNA expression patterns among the PBC subtypes using comprehensive deep sequencing. We also carried out histologic examination by laser capture microdissection and investigated how the identified miRNAs were involved in PBC clinical progression using the miRNA transfection method. On average, ~11 million 32-mer short RNA reads per sample were obtained, and we found that the expression levels of 97 miRNAs differed significantly among the four groups. Heat mapping demonstrated that the miRNA profiles from hepatic failure and portal hypertension type were clustered differently from those of the gradually progressive type and controls. Furthermore, we focused on miR-139-5p, which has an adequate number of total short reads. Quantitative reverse transcription PCR showed that miR-139-5p was significantly downregulated in clinically advanced PBC. Also, examination of liver tissues demonstrated that the expression of lymphocyte-derived miR-139-5p was significantly higher in hepatocytes. In vitro, the level of TNF-α was significantly elevated in supernatant of cells with upregulation of miR-139-5p. Furthermore, c-FOS gene transcription was repressed. Thus, we have demonstrated a novel inflammation-regulatory mechanism involving TNF-α and c-FOS transcription through miR-139-5p in the NF-κB signaling pathway. We conclude that the specific miRNA miR-139-5p might be involved in the pathogenesis of PBC, especially during clinical progression.
Similar content being viewed by others
Main
Primary biliary cholangitis (previously called primary biliary cirrhosis) (PBC) is a chronic cholestatic liver disease pathologically characterized by destruction of the intrahepatic bile ducts.1 The etiology of PBC appears to involve a variety of factors, and the contributions of autoimmune mechanisms, genetic predisposition, and environmental factors have been reported previously.2, 3 Moreover, PBC is characterized serologically by the presence of anti-mitochondrial antibody directed against the E2 subunit of the pyruvate dehydrogenase enzyme complex, and an increased level of serum immunoglobulin M.4, 5 PBC is also often complicated by other autoimmune diseases, such as chronic thyroiditis, Sjögren’s syndrome, and rheumatoid arthritis.6, 7 Thus, abnormal immune responses might induce inflammations of cholangiocytes and may be involved in the pathogenesis of PBC. Indeed, previous studies have demonstrated increased serum levels of pro-inflammatory cytokines (IL-6, TNF-α, IFN-γ) in PBC patients with fibrosis.8, 9
Some genome-wide association study in a PBC population from some countries have demonstrated the specific susceptibility loci, suggesting a genetic predisposition.10, 11, 12, 13 These findings suggest that genetic factors contribute to the development of PBC. Currently, some murine animal models of PBC are available.14 With these models, substantial progress have been achieved to clarify the pathogenesis of PBC. However, the exact mechanism of pathogenesis in PBC have remained unclear.
Because of wider recognition of the disease and improvements in diagnostic procedures, ~70–80% of patients with PBC are clinically diagnosed in the early or asymptomatic stage. The majority of these patients respond well to standard medical therapy, for example, using ursodeoxycholic acid (UDCA), resulting in a stable course over several decades.15, 16 However, other patients develop symptoms such as pruritus, jaundice, severe ascites, or hepatic encephalopathy owing to liver damage, and show progression to liver failure within a short period.15, 17 The remaining patients develop portal hypertension and esophageal varices without overt jaundice, and at some stage will require liver transplantation or endoscopic treatment for prevention of esophageal varices.18, 19 Therefore, the prognosis of PBC varies according to the clinical course. Patients who develop hepatic failure have a poor prognosis, with a 5-year survival rate of <50%.20, 21, 22
The clinical course of PBC can be classified into three subtypes:
-
i
Gradually progressive type: in which a major proportion of patients respond to drug therapy, showing gradual progression and remaining asymptomatic for a long period.
-
ii
Portal hypertension type: in which patients have symptoms such as pruritus and develop esophageal varices as a result of portal hypertension, without overt jaundice.
-
iii
Hepatic failure type: in which patients rapidly develop overt jaundice and ultimately hepatic failure, accounting for only a small proportion of cases.
Recently, diagnosis of PBC has become relatively easy on the basis of laboratory hallmarks and histological characteristics.23 However, prognosis at the time of initial diagnosis is still difficult, because of the lack of reliable predictive markers. Therefore, it is clinically very important to establish a reliable surrogate marker for predicting the clinical course of PBC. Furthermore, it would be expected that clarification of the mechanism responsible for clinical progression to portal hypertension or liver failure would have an impact on treatment and, possibly, prognosis.
microRNAs (miRNAs) are small, endogenous non-coding RNA molecules comprising 19 to 24 nucleotides. They bind imperfectly to complementary sequences in the 3' untranslated region (UTR) of target messenger RNAs and repress their translation,24, 25, 26, 27 and in this capacity miRNAs are important regulators of protein expression. miRNAs have also been attracting attention recently because previous studies have revealed that they show specific patterns of expression in a variety of diseases.28, 29 miRNA expression patterns in either tissues or serum of PBC patients have also been reported. For example, miR-506 is upregulated in cholangiocytes from PBC patients and related to decreased AE2 activity.30 Specific expression profiling patterns of 35 miRNAs in liver biopsy samples from PBC patients have also been reported,31 and a microarray study has shown that 17 miRNAs were differentially expressed relative to controls in peripheral blood mononuclear cells from patients with PBC.32 Deep sequencing has also revealed that PBC patients have specific serum miRNA expression patterns in comparison with healthy controls or patients with viral hepatitis.33 These reports suggest that miRNAs have potential as biomarkers and might play a role in the pathogenesis of PBC.
In the present study, we evaluated patterns of miRNA expression in serum samples from patients with the three distinct clinical subtypes of PBC using the comprehensive deep sequencing technique, and attempted to identify novel biomarkers for predicting these clinical subtypes. Using samples of liver tissue and a cultured cell line, we also investigated the mechanism by which the identified miRNAs might be involved in the progression and aggravation of PBC. In addition, computer analysis of differentially expressed miRNAs was conducted to predict the target genes and their biological functions. Finally, we evaluated the function of the identified miRNAs to clarify the mechanisms involved in the disease progression of PBC.
Materials and methods
PBC Patients and Healthy Volunteers
The diagnosis of all cases was based on internationally established criteria with liver biopsy.23 In this way we were able to exclude patients with chronic viral hepatitis, autoimmune hepatitis (AIH), PBC and AIH overlap syndrome, and other malignant tumors. After 2 years of treatment with UDCA, we judged the responses shown during the clinical course using international criteria.34 On the basis of the treatment response, we then classified the PBC patients into three groups according to their clinical course as follows: (i) the predominant form in which progression is gradual and the patients remain asymptomatic for a long period (gradually progressive type), (ii) a form in which the patients develop portal hypertension without jaundice (portal hypertension type), and (iii) a form in which a small proportion of patients rapidly develop hepatic failure with jaundice (hepatic failure type). Patients with each of these three PBC subtypes, and healthy subjects as controls, were enrolled in this study (n=7, 6, 6, and 6, respectively) (Table 1). In the following experiments, we initially decided to measure miRNAs in serum samples from patients with the various PBC subtypes to identify their patterns of expression. For this purpose, we prepared serum from a total of 19 PBC patients and 6 healthy controls. All the study protocol above was approved by the Ethics Committee of Yamagata University Faculty of Medicine (#25–82), and written informed consent was obtained from each individual.
Deep Sequencing Using an Illumina Sequencer
A deep sequencer is capable of analyzing huge numbers of small-molecule RNA species in a single run,35 making it an ideal tool for this type of study. The instrument employed for this purpose was an Illumina Genome Analyzer IIx (Illumina, CA, USA).
Total RNA was extracted from 800 μl of serum from each of the PBC patients and healthy controls using Trizol LS (Invitrogen, CA, USA). miRNA libraries were then prepared from total RNA using the TruSeq Small RNA Sample Prep Kit (Illumina), as described in a previous report.33 The miRNA libraries were sequenced on a deep sequencer (SCS 2.8 software; Illumina), with a 32-mer single-end sequence. Image analysis and base calling were performed using RTA 1.8 software.
Sequencing and Statistical Analysis
Raw miRNA sequence reads were checked for quality, and the 3' and 5' adapter sequences were cut by cutadapt softwere while discarding reads shorter than 20 nucleotides.36 The sequence reads were aligned with miRBase (June 2014 release) and hg19 (human genome 19, UCSC) used by bwa (0.5.9-r16), allowing one nucleotide base mismatch.37, 38 Digital expression levels were neutralized by taking account of the length of the miRNAs. The total number of miRNA reads generated in each library was expressed using trimmed mean of M values (TMM) normalization.39 Read counts for each isolated miRNA were normalized to the total number of miRNA reads, and then the ratio was multiplied by a constant set to 1 × 106. One-way analysis of variance (ANOVA) was performed on the number of sequence reads with P-values, comparing differentially expressed miRNAs in the disease group. Differentially expressed miRNAs were filtered using a P-value cutoff of 0.05. Hierarchical clustering was performed using the R platform and a heatmap described as using a function of heatmap.2 in gplots.40
Validation Study
In addition to initial samples analyzed by deep sequencing, more serum samples from patients with the three PBC subtypes and healthy controls (a total of 12, 7, 7, and 11 samples in each group) were used in a validation study employing quantitative RT-PCR. The protocol was previously reported by Mitchell.41 We used Caenorhabditis elegans microRNA-39 (cel-miR-39) (5′-UCACCGGGUGUAAAUCAGCUU-3′) as an internal control miRNA. Cel-miR-39 was synthesized by Sigma Aldrich Japan (Tokyo, Japan). Reverse transcription was conducted using a TaqMan miRNA RT kit (Applied Biosystems, CA, USA) for identification of cel-miR-39 expression in the presence of the internal control at 5 fmol/ml. qRT-PCR was conducted for detection of the identified miRNAs in 20-μl PCR reaction volumes using the TaqMan MicroRNA assay (Applied Biosystems), as reported previously.33
Levels of expression of the identified miRNAs detected by qRT-PCR were normalized to cel-miR-39 and presented as the fold change (2−ΔΔCt) above the level for the healthy control (healthy-4): ΔΔCt=(CtmiRNA-Ctcel-miR-39)patients−(CtmiRNA-Ctcel-miR-39)healthy. These normalized levels of miRNA expression were compared among the PBC subtypes using Student’s t test.
Furthermore, in PBC patients for whom follow-up was possible, we measured the levels of miRNA at two years after initial therapy and assessed the degree of change.
Examination of miRNA Expression in Liver Tissue of PBC Patients
We identified the patterns of miRNA expression in serum samples from patients with the various PBC subtypes. For more detailed investigation of whether these miRNAs were involved in the clinical progression of PBC, we also examined the expression of miRNAs in liver tissue from the PBC patients. We searched for the possible cellular origin of the miRNAs using the laser capture microdissection (LCM) technique (PALM-MBI, Carl Zeiss, Jena, Germany) and digital PCR quantification (QuantStudio 3D Digital PCR System, Life Technologies, CA, USA) on formalin-fixed paraffin-embedded (FFPE) liver tissues. To identify the origin of miRNA expression in liver tissue from PBC patients, areas containing hepatocytes, or infiltrating lymphocytes in areas of chronic non-suppurative destructive cholangitis (CNSDC), were each dissected out selectively, and the cell-derived miRNAs were extracted using a miRNeasy FFPE Kit (QIAGEN, Hilden, Germany) in accordance with the manufacturer’s instructions. As the amounts of the extracted miRNAs were extremely small, it was very difficult to measure them quantitatively using a traditional real-time PCR system. Therefore, we employed a digital PCR apparatus capable of assaying very low concentrations with high sensitivity.42
In situ Hybridization
The sites of expression of specific miRNAs were then further confirmed using in situ hybridization (ISH) employing a miRCURY LNA microRNA ISH Optimization Kit (EXIQON, Vedbaek, Denmark) in accordance with the manufacturer’s protocol. ISH is a technique for visualizing miRNA expression in tissues. We used FFPE tissue sections from patients with gradually progressive PBC-containing areas of CNSDC and detected the miRNA signals using a double-DIG-labeled LNA probe that hybridized with the miRNA sequence. This allowed us to identify and visually confirm the localization of miRNAs in PBC tissue.
Cell Culture and Transfection of miRNA (in vitro)
The human Huh7 cell line was used for functional analysis of identified miRNAs in vitro. Specific miRNAs in Huh7 cells were artificially upregulated using a lipofection method and their cellular functions were then evaluated. First, we disseminated 1 × 105 cells at 60–70% confluency into six-well plates. After one passage overnight, we upregulated the miRNA by addition of a miRNA mimic in accordance with the manufacturer’s recommendation (miRVana miRNA Mimics, Life Technologies). A negative control was included in every experimental model. After miRNA transfection, we cultured the cells continuously for 3 days, and then harvested the cell culture supernatants. We then measured the concentrations of various pro-inflammatory cytokines using an ELISA kit (MULTIPLEX Pro-Inflammatory Cytokine Kit, COSMO BIO, Tokyo, Japan).
In silico Analysis
To determine the possible target genes of the differentially expressed miRNAs, we searched the TargetScan Human 7.0 (August 2015 release, http://www.targetscan.org) and microRNA.org (August 2010 release, http://www.microrna.org) database. TargetScan is able to predict the highest number of known miRNA targets in mammals obtained from TarBase 6.0. MicroRNA.org is a comprehensive resource of miRNA targets based on the miRanda algorithm. Additionally, they take into account the importance of conservation of target sites in the 3'-UTR of the gene across several species. As generally there are very many predicted target genes, because of the fact that a single miRNA can regulate the expression of multiple genes, we selected the predicted target genes of identified miRNAs having multiple target 3'-UTR sites and having a more marked effect. Furthermore, we predicted a pathway between the identified pro-inflammatory cytokine and the miRNA target gene.
Western Blotting
The actual target protein in Huh7 after artificial upregulation of the miRNA was evaluated by western blotting. To obtain total protein extracts, the transfected Huh7 cells were collected and cell lysates were prepared from them (CellAmp Direct Prep Kit, TaKaRa, Shiga, Japan). The miRNA-targeted protein and β-actin signals were detected by chemiluminescence using a ChemiDoc XRS Plus System (BioRad, CA, USA), and their standardized protein volumes were quantified by Image Lab Software (BioRad). All antibodies were purchased from Santa Cruz Biotechnology.
Results
Comprehensive Deep Sequencing Analysis
Using single-end deep sequencing, we analyzed circulating miRNAs detected in 25 serum samples from patients with PBC and healthy controls. On average, ~11 million 32-mer short reads per sample were obtained. After excluding the adaptor and tag sequences of the reads using cutadapt, the resulting high-quality reads were analyzed.36 The miRBase and hg19 mapping rates were 9.26 and 69.94% on average, respectively (Table 2).
We normalized the differential expression of miRNA count data using the TMM normalization process, and the number of individual miRNA reads was standardized by a total number of 1 000 000 reads per sample.39 Subsequently, for comparison of the four groups, the differential expression levels of miRNA were extracted using ANOVA. Finally, we obtained 1514 database-registered miRNAs from the total short reads. Furthermore, among a total of 1514 miRNAs obtained, the calculated P-value and the expression levels of 97 miRNAs were found to differ significantly among the four groups (P<0.05).
Heat mapping and hierarchical clustering demonstrated that the miRNA profiles from both the hepatic failure and portal hypertension groups were clustered differently from those of the gradually progressive and control groups (Figure 1). Notably, there was a characteristic difference in the patterns of miRNA expression in the portal hypertension and hepatic failure groups. The expression patterns in the gradually progressive group were similar to those in the healthy controls. Eight miRNAs were characteristically upregulated (more than fivefold) and 11 miRNAs were downregulated (<0.25-fold) in the hepatic failure and portal hypertension groups. The miRNA showing the greatest increase was miR-5096 (Table 3A), whereas that showing the greatest decrease was miR-5001-3p (Table 3B). However, as these miRNAs had small numbers of total reads sufficient confirmation of the validation was not possible. However, it was demonstrated that some specific miRNAs appeared to be associated with the clinical progression of PBC.
Heat mapping and clustering analysis. Heat mapping was performed and described using a function of heatmap.2 in gplots. It uses hierarchical clustering with Euclidean distance; Pearson Linear Correlation; and Ward’s method are used to generate the hierarchical tree. Red indicates a high level of miRNA expression and blue indicates low expression. Heatmap demonstrates that 97 identified miRNAs profiles from both the hepatic failure and portal hypertension types were clustered differently from those of the gradually progressive type and controls.
miRNA Selection and Validation by Real-Time PCR
Among 97 miRNAs for which quantification was possible based on their significant expression patterns and adequate number of total short reads, we focused on those that had been commonly identified in serum of PBC patients.33 As it has been reported that miR-139-5p is persistently and specifically expressed in PBC, we speculated that it would have a specific role in the pathogenesis of various clinical subtypes of PBC.
We used qRT-PCR to verify the data obtained by deep sequencing. In addition to the initial samples, 10 additional cases of PBC and control samples were quantified. The relative expression levels were analyzed by TaqMan MicroRNA Assay and normalized relative to cel-miR-39 as an internal control. The levels of miR-139-5p expression in the hepatic failure and portal hypertension groups were significantly reduced in comparison with the healthy controls (Figure 2). In the gradually progressive group, the expression of miR-139-5p tended to be lower than in the healthy controls, but not to a significant degree. Thus, we were able to confirm that miR-139-5p was significantly downregulated in clinically advanced PBC.
Validation study using real-time PCR for miR-139-5p. In addition to the initial samples, 10 additional cases of various PBC subtypes and controls were quantified. The threshold cycle for the miR-139-5p primer/probe set was normalized against a cel-miR-39 primer/probe pair and compared with the control. The expression levels of miR-139-5p in the hepatic failure and portal hypertension groups were significantly reduced in comparison with the others. In the gradually progressive type, miR-139-5p tended to be decreased relative to the healthy controls, although the difference was not significant. *P<0.05.
Furthermore, miR-139-5p tended to increase in the gradually progressive type (supplementary Figure 1), whereas these levels tended to decrease in both portal hypertension and hepatic failure type. However, these chronological changes were not significant (paired t-test).
Evaluation of The Level of miR-139-5p Expression in Liver Tissue From Patients with PBC
Using LCM, we investigated the expression of miR-139-5p in FFPE liver sections from patients with PBC. We selectively dissected out lymphocytes in CNSDC and areas of hepatocytes in which there were no infiltrating lymphocytes (Figure 3a). When we measured the expression of miR-139-5p using digital PCR, that derived from lymphocytes was significantly higher than that derived from hepatocytes (Figure 3b). We also conducted similar examinations using disease controls (AIH, chronic hepatitis C: CHC), and found that lymphocyte-derived and hepatocyte-derived miR-139-5p had equivalent levels of expression, unlike the PBC samples (Figure 3b). Furthermore, the level of expression of lymphocyte-derived miR-139-5p from PBC was significantly higher than that from either AIH or CHC, whereas the level of expression of hepatocyte-derived miR-139-5p from PBC was significantly lower than that from either AIH or CHC (Figure 3c). These findings confirmed that the expression pattern of miR-139-5p was specific to PBC tissue.
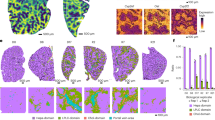
Laser capture microdissection of paraffin-embedded liver tissues from PBC patients. (a) Areas containing hepatocytes or infiltrating lymphocytes in CNSDC were selectively dissected, respectively, using LCM; the left columns show the histology before LCM and the right columns that after LCM (H&E). The bottom is the area of hepatocytes after LCM (H&E, 25×). (b) The expression levels of hepatocyte or lymphocyte-derived miR-139-5p were measured by digital PCR; lymphocyte-derived miR-139-5p expression was significantly higher than that from hepatocytes. Furthermore, lymphocyte-derived miR-139-5p and hepatocyte-derived miR-139-5p showed equivalent expression levels in control samples of other liver diseases (AIH, CHC). (c) Lymphocyte-derived miR-139-5p expression in PBC samples was significantly higher than that in AIH or CHC samples (upper). Conversely, hepatocyte-derived miR-139-5p expression in PBC samples was significantly lower than that in AIH or CHC samples (lower). Shown are means±s.d. *P<0.05; **P<0.01. Scale bare=50 μm.
In situ Hybridization For Detection of miR-139-5p in PBC Tissue
In situ hybridization demonstrated that miR-139-5p was strongly expressed in lymphocytes in areas of CNSDC in PBC liver tissue. Also, it revealed that miR-139-5p was constantly expressed in the nuclei of hepatocytes (Figure 4). This pattern of miR-139-5p expression in hepatocytes was similar to that seen in those from hepatocytes in other liver diseases, AIH and CHC (supplementary Figure 2). This result confirmed that miR-139-5p was usually present in the nuclei of hepatocytes, although its expression was specifically upregulated in lymphocytes infiltrating the liver in PBC.
In situ hybridization. miR-139-5p, 40 nM, was strongly present in lymphocytes (arrow) contributing to chronic non-suppurative destructive cholangitis in PBC samples, and was constantly expressed in nuclei of hepatocytes (arrowhead). The left lower column shows a positive control stained by U6, 0.1 nM, which is a small non-coding RNA constantly present in cells. The right lower column shows a negative control stained by scrambled, 40 nM (Nuclear Fast Red, 25×). Scale bar=50 μm.
Elevated Level of the Pro-Inflammatory Cytokine TNF-α in miRNA-Transfected Cells
For analysis of miR-139-5p function in PBC, Huh7 cells were manipulated by introduction of miR-139-5p according to the protocol described above. Among various pro-inflammatory cytokines, the level of TNF-α was significantly elevated (mean: 380.5 pg/ml) in the supernatant of cells with up-regulated miR-139-5p relative to that of control cells (mean: 38.2 pg/ml) (Figure 5a). Furthermore, RT-PCR showed that the level of TNF mRNA in transfected cells was increased more than fourfold in comparison with control cells (Figure 5b).
Elevation of the pro-inflammatory cytokine TNF-α in miRNA-transfected cells. (a) ELISA showed that the TNF-α level was significantly elevated (mean: 380.5 pg/ml) in supernatant of cells with upregulated miR-139-5p in comparison with control cells (mean: 38.2 pg/ml). (b) RT-PCR showed that expression of TNF mRNA standardized using the housekeeping gene, GAPDH, was significantly increased more than fourfold in cells with upregulated miR-139-5p in comparison with control cells. *P<0.05.
c-FOS, an Important Target Gene of miR-139-5p
We focused on miR-139-5p, which shows a specific expression pattern in clinically advanced PBC. Use of TargetScan Human 7.0 and microRNA.org database for predicting the target gene of miR-139-5p identified it as c-FOS. Among various predicted genes, c-FOS has two 3'-UTR regions (positions 96–102 and 526–532, conserved) that are repressed by miR-139-5p (Figure 6a). Especially, the conserved position 526–532 is common in both algorithms. Also, the mirSVR score was −1.2648 and the PhastCons score was 0.7442. Therefore, it was considered that c-FOS would be strongly controlled by miR-139-5p as a biological target.
c-FOS transcription is regulated by miR-139-5p. (a) In silico, the c-FOS gene was identified as the target gene of miR-139-5p using the predicting tool, TargetScan Human 7.0. The c-FOS gene has two 3'UTR regions to which miR-139-5p binds: conserved positions 526–532, and non-conserved positions 96–102. (b) RT-PCR showed that expression of c-FOS mRNA standardized by GAPDH was not changed in cells with up-regulated miR-139-5p or in control cells. N.S; not significant. (c) Western blotting showed that expression of c-FOS protein in cells with upregulated miR-139-5p was decreased in comparison with control cells. c-FOS in control cells and β-actin in both cell types showed the same level of expression. (d) Expression of c-FOS protein standardized by β-actin in miR-139-5p upregulated cells was significantly decreased relative to control cells.
Although c-FOS mRNA expression in miR-139-5p-transfected cells was not changed (Figure 6b), western blotting showed that expression of c-FOS protein standardized against β-actin in miR-139-5p upregulated cells was decreased in comparison to that of control cells (Figure 6CD). Also, c-FOS in control cells or β-actin in both cells showed the same level of expression. This result indicates that c-FOS gene transcription is obviously regulated by miR-139-5p, and thus c-FOS protein expression is repressed.
Mechanism of Inflammation Regulation Through miR-139-5p in the NF-κB Signaling Pathway
We examined the relationship of c-FOS regulation by miR-139-5p with TNF-α using pathway analysis. This revealed a novel inflammation-regulatory mechanism between TNF-α and c-FOS transcription through miR-139-5p in the NF-κB signaling pathway (Kyoto Encyclopedia of Genes and Genomes: KEGG).
Generally, TNF-α first binds to the tumor necrosis factor receptor1 (TNFR1) in the NF-κB signaling pathway, and activates the downstream part of the pathway. Subsequently, IκB proteins of NF-κB are degraded by phosphorylation, and then NF-κB p65/p50 subunits move to the nucleus and accelerate the inflammatory response. c-FOS protein acts on this p65 subunit, and this has a role in inhibiting the induction of TNF-α.43
The increased miR-139-5p inhibits the transcription of c-FOS and TNF-α transcription is accelerated. Thus, the NF-κB signaling pathway is finally activated and it could induce the further production of TNF-α. Furthermore, elevation of the TNF-α level might exacerbate bile ducts injury by positive feedback acceleration (Figures 7a and b).
miR-139-5p inhibits c-FOS transcription, and induces activation of the inflammatory response through TNF-α. (a) c-FOS inhibits the p65 subunit of NF-κB activation in the NF-κB signaling pathway. Therefore, c-FOS regulates abnormal inflammation. (b) The novel inflammation-regulatory mechanism involving TNF-α and c-FOS transcription through miR-139-5p in the NF-κB signaling pathway. The up-regulated miR-139-5p inhibits c-FOS transcription, so that the production of TNF-α is accelerated. Finally, TNF-α activates NF-κB signaling through positive feedback and the increased TNF-α induces more severe inflammation of cholangiocytes.
Discussion
The diagnosis of PBC is established on the basis of sero-biochemical and histopathological criteria. This allows many patients to be diagnosed at an early stage, thus facilitating prompt treatment, although some patients do not respond to standard therapies and show progression to liver failure or portal hypertension, eventually requiring either liver transplantation or endoscopic therapy.23 Thus, although the diagnostic criteria for PBC have been established, the clinical progression of PBC has been unpredictable. To clarify the different clinical subtypes of PBC, we focused on miRNAs that have been reported to be related to the pathogenesis of a number of human diseases.28, 41, 44, 45, 46 Although several previous studies have reported the expression patterns of miRNAs in tissue or serum from patients with PBC, those operating in the clinical progression of PBC have remained unclear.
In the present study, we identified specific patterns of miRNA expression in serum from patients with the three clinical PBC subtypes using both deep sequencing and digital PCR. Hierarchical clustering of the 97 identified miRNA profiles revealed that the patterns characterizing the hepatic failure and portal hypertension types differed from those characterizing the gradually progressive type and controls, suggesting that some miRNAs may be involved in the clinical progression of PBC, or that some cases may progress as a result of specific changes in the expression of some miRNAs. It appeared that some miRNAs had important roles in the mechanisms underlying disease progression in the various clinical subtypes. Among the 97 miRNAs identified in this study, some were represented in very small amounts, which made the validation experiments difficult. Nevertheless, we suspected that these miRNAs had an important role in the disease process, and we focused on one potentially important miRNA, miR-139-5p, which had previously been reported in cases of PBC.33 We found that the expression of miR-139-5p changed significantly: real-time PCR confirmed that miR-139-5p in serum from patients with advanced PBC (portal hypertension type and hepatic failure type) showed lower expression in comparison with early PBC (gradually progressive type) and healthy controls.
Furthermore, in PBC liver tissue, the expression of miR-139-5p in lymphocytes constituting CNSDC was significantly higher than that in hepatocytes from the same patient. This difference of miR-139-5p expression was not observed in samples from patients with other liver diseases (AIH and CHC). Also, the expression of miR-139-5p in lymphocytes from PBC tissue was higher than that in lymphocytes from the other liver disease controls. On the other hand, the expression of miR-139-5p in hepatocytes from PBC liver was lower than that in hepatocytes from other liver disease controls. Although the main site of action of miR-139-5p in the liver (ie, lymphocytes or hepatocytes) remains undetermined, we confirmed that miR-139-5p likely has an important role in the progression of PBC, and that from a histologic viewpoint altered expression of miR-139-5p appears to be specific to PBC. Taken together with the data from serum deep sequencing, it appears that miR-139-5p has a specific role in the progression of PBC.
Several studies have reported that miRNAs have specific expression patterns in various diseases. However, there have been few detailed examinations of miRNA biological function in the liver. Therefore, as a next step, we also attempted to clarify the function of miR-139-5p which we had identified.
Among various predicted genes, we identified c-FOS as the target gene of miR-139-5p in silico. This result was consistent for the conserved position and scoring in algorithms. c-FOS is a member of the FOS family, which also includes AP-1 with the Jun proteins (jun-B, c-jun).47, 48 c-FOS protein also functions to suppress inflammatory responses, ie, c-FOS protein acts directly on the NF-κB p65 subunit, thus inhibiting the pathway downstream of NF-κB and induction of pro-inflammatory cytokines including TNF-α.49 A previous study reported that the c-FOS gene was downregulated in peripheral blood mononuclear cells isolated from PBC patients. Furthermore, expression of c-FOS recovered after treatment with UDCA.50 The pro-inflammatory cytokine TNF-α has also been reported to be increased in sera of PBC patients at the advanced stage.8 Thus, reduced c-FOS protein and increased TNF-α would exacerbate cholangitis in PBC. Therefore, in terms of treatment response, c-FOS might be an important factor responsible for more severe inflammation in PBC.
In the present study, we revealed that miR-139-5p suppresses c-FOS transcription in Huh7 cells. In silico, miR-139-5p regulates only c-FOS not another target of the pathway from the results of two predictive algorithms and confident scoring. Downregulated c-FOS protein in cells with upregulation of miR-139-5p induces further production of TNF-α through NF-κB activation. In situ hybridization and RT-digital PCR showed that miR-139-5p was overexpressed in lymphocytes around portal areas in the liver of PBC patients. Therefore, expression of c-FOS protein was reduced by miR-139-5p inhibition in lymphocytes of PBC patients, in accord with the previous report.50 In vitro, our study showed that miR-139-5p regulates c-FOS transcription and induces greater production of TNF-α in the NF-κB signaling pathway. As the limitation in the current study, we tried to measure TNF-α and c-FOS levels in hepatocytes and lymphocytes obtained from LMD to confirm if they correlate with the findings of miR-139-5p. However, we are not able to measure them in the current isolated cells by LMD method. As the expression levels of total RNAs and proteins in FFPE tissue is quite limited, it is technically impossible to measure the protein levels of these using standard laboratory methods (eg, ELISA or Immunoblotting). Finally, we examined the relationship among miR-139-5p, c-FOS and TNF-α in PBC, and found a novel inflammation-regulatory mechanism involving TNF-α and c-FOS transcription through miR-139-5p in the NF-κB-signaling pathway.
The present results revealed that infiltrating lymphocytes were the main source of miR-139-5p, although the level of expression did not reflect that in serum samples. Therefore, we speculate that miRNA from surrounding hepatocytes might mask the increase of miR-139-5p in lymphocytes.
Our present finding that miR-139-5p is a novel regulatory factor in PBC progression is very important from two aspects. One is that the identified miRNA might have potential as a predictive biomarker. The change of miR-139-5p over time after an initial treatment, UDCA administration did not cause significant difference in each group. Thus, regardless of treatment, miR-139-5p specifically reflects according to the clinical condition of PBC. This means a possibility of evaluating miR-139-5p as a novel disease biomarker of PBC. Establishment of novel biomarkers for prognostication of PBC, especially at the time of diagnosis, would seem feasible, and miRNAs are now emerging as highly tissue-specific biomarkers with potential clinical applicability. Therefore, we think that this specific miRNA could become a novel biomarker for predicting the clinical course of PBC at the time of diagnosis.
Another important finding is that miR-139-5p could be specifically involved in the progression of PBC, and thus could become a potential therapeutic target. miRNAs regulate post-transcription and have an important role in biological processes. Therefore, regulation of miR-139-5p could be exploited for molecular targeting in the treatment of PBC, and this miRNA could become not only a biomarker of disease progression, but also a novel therapeutic target for patients with progressive PBC subtypes.
References
Nakanuma Y, Ohta G . Histometric and serial section observations of the intrahepatic bile ducts in primary biliary cirrhosis. Gastroenterology 1979; 76: 1326–1332.
Selmi C, Mayo MJ, Gish RG et al, Primary biliary cirrhosis in monozygotic and dizygotic twins: genetics, epigenetics, and environment. Gastroenterology 2004; 127: 485–492.
Boonstra K, Beuers U, Ponsioen CY . Epidemiology of primary sclerosing cholangitis and primary biliary cirrhosis: a systematic review. J Hepatol 2012; 56: 1181–1188.
Gershwin ME, Mackay IR, Coppel RL . Identification and specificity of a cDNA encoding the 70 kd mitochondrial antigen recognized in primary biliary cirrhosis. J Immunol 1987; 138: 3525–3531.
Coppel RL, McNeilage LJ, Whittingham S et al, Primary structure of the human M2 mitochondrial autoantigen of primary biliary cirrhosis: dihydrolipoamide acetyltransferase. Proc Natl Acad Sci USA 1988; 85: 7317–7321.
Selmi C, Meroni PL, Gershwin ME . Primary biliary cirrhosis and Sjogren's syndrome: autoimmune epithelitis. J Autoimmun 2012; 39: 34–42.
Teufel A, Weinmann A, Worns M et al, Concurrent autoimmune diseases in patients with autoimmune hepatitis. J Clin Gastroenterol 2010; 44: 208–213.
Neuman M, Angulo P, Dickson ER et al, Tumor necrosis factor-alpha and transforming growth factor-beta reflect severity of liver damage in primary biliary cirrhosis. J Gastroenterol Hepatol 2002; 17: 196–202.
Golovanova EV, Il'chenko L, Gudkova RB et al, Cytokines in primary biliary cirrhosis (diagnostic and prognostic value). Ter Arkh 2004; 76: 8–11.
Nakamura M, Nishida N, Yasunami M et al, Genome-wide association study identifies TNFSF15 and POU2AF1 as susceptibility loci for primary biliary cirrhosis in the Japanese population. Am J Hum Genet 2012; 91: 721–728.
Hirschfield GM, Liu X, Lu Y et al, Primary biliary cirrhosis associated with HLA, IL12A, and IL12RB2 variants. N Engl J Med 2009; 360: 2544–2555.
Liu X, Invernizzi P, Bianchi I et al, Genome-wide meta-analyses identify three loci associated with primary biliary cirrhosis. Nat Genet 2010; 42: 658–660.
Mells GF, Floyd JA, Shin SY et al, Genome-wide association study identifies 12 new susceptibility loci for primary biliary cirrhosis. Nat Genet 2011; 43: 329–332.
Katsumi T, Tomita K, Ueno Y et al, Animal models of primary biliary cirrhosis. Clin Rev Allergy Immunol 2015; 48: 142–153.
Pares A, Caballeria L, Rodes J . Excellent long-term survival in patients with primary biliary cirrhosis and biochemical response to ursodeoxycholic Acid. Gastroenterology 2006; 130: 715–720.
Prince M, Chetwynd A, James OF et al. Survival and symptom progression in a geographically based cohort of patients with primary biliary cirrhosis: follow-up for up to 28 years. Gastroenterology 2002; 123: 1044–1051.
Springer J, Cauch-Dudek K, Heathcote EJ et al, Asymptomatic primary biliary cirrhosis: a study of its natural history and prognosis. Am J Gastroenterol 1999; 94: 47–53.
Christensen E, Neuberger J, Altman DG et al, Azathioprine and prognosis in primary biliary cirrhosis. Gastroenterology 1986; 90: 508–509.
Gores GJ, Wiesner RH, Langworthy A et al, Prospective evaluation of esophageal varices in primary biliary cirrhosis: development, natural history, and influence on survival. Gastroenterology 1989; 96: 1552–1559.
Corpechot C, Carrat F, Poupon R et al, The effect of ursodeoxycholic acid therapy on the natural course of primary biliary cirrhosis. Gastroenterology 2005; 128: 297–303.
Corpechot C, Poupon R . Geotherapeutics of primary biliary cirrhosis: bright and sunny around the Mediterranean but still cloudy and foggy in the United Kingdom. Hepatology 2007; 46: 963–965.
Metcalf JV, Mitchison HC, James OF et al, Natural history of early primary biliary cirrhosis. Lancet 1996; 348: 1399–1402.
Lindor KD, Gershwin ME, Heathcote EJ et al, Primary biliary cirrhosis. Hepatology 2009; 50: 291–308.
Lau NC, Lim LP, Bartel DP et al, An abundant class of tiny RNAs with probable regulatory roles in Caenorhabditis elegans. Science 2001; 294: 858–862.
Lagos-Quintana M, Rauhut R, Tuschl T et al, Identification of novel genes coding for small expressed RNAs. Science 2001; 294: 853–858.
Lee RC, Ambros V . An extensive class of small RNAs in Caenorhabditis elegans. Science 2001; 294: 862–864.
Bartel DP . MicroRNAs: genomics, biogenesis, mechanism, and function. Cell 2004; 116: 281–297.
Chen X, Ba Y, Wang K et al, Characterization of microRNAs in serum: a novel class of biomarkers for diagnosis of cancer and other diseases. Cell Res 2008; 18: 997–1006.
Lu J, Getz G, Peck D et al, MicroRNA expression profiles classify human cancers. Nature 2005; 435: 834–838.
Banales JM, Saez E, Splinter P et al, Up-regulation of microRNA 506 leads to decreased Cl-/HCO3- anion exchanger 2 expression in biliary epithelium of patients with primary biliary cirrhosis. Hepatology 2012; 56: 687–697.
Padgett KA, Lan RY, Pfeiff J et al, Primary biliary cirrhosis is associated with altered hepatic microRNA expression. J Autoimmun 2009; 32: 246–253.
Katsushima F, Takahashi A, Ohira H et al, Expression of micro-RNAs in peripheral blood mononuclear cells from primary biliary cirrhosis patients. Hepatol Res 2013; 44: E189–E197.
Ninomiya M, Kondo Y, Kakazu E et al, Distinct microRNAs expression profile in primary biliary cirrhosis and evaluation of miR 505-3p and miR197-3p as novel biomarkers. PloS One 2013; 8: e66086.
Corpechot C, Abenavoli L, Johanet C et al, Biochemical response to ursodeoxycholic acid and long-term prognosis in primary biliary cirrhosis. Hepatology 2008; 48: 871–877.
Ninomiya M, Ueno Y, Shimosegawa T . Application of deep sequence technology in hepatology. Hepatol Res 2014; 44: 141–148.
Martin M . Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet J 2011; 17: 10–12.
Maglott D, Ostell J, Tatusova T et al, Entrez Gene: gene-centered information at NCBI. Nucleic Acids Res 2011; 39: D52–D57.
Kozomara A, Griffiths-Jones S . miRBase: integrating microRNA annotation and deep-sequencing data. Nucleic Acids Res 2011; 39: D152–D157.
Robinson MD, Oshlack A . A scaling normalization method for differential expression analysis of RNA-seq data. Genome Biol 2010; 11: R25.
Team RDC R . A language and environment for statistical computing. The R Foundation for Statistical Computing: Vienna, Austria, 2011.
Mitchell PS, Parkin RK, Pogosova-Agadjanyan EL et al, Circulating microRNAs as stable blood-based markers for cancer detection. Proc Natl Acad Sci USA 2008; 105: 10513–10518.
Day E, Dear PH, McCaughan F . Digital PCR strategies in the development and analysis of molecular biomarkers for personalized medicine. Methods 2013; 59: 101–107.
Koga K, Takaesu G, Kinjyo I et al, Cyclic adenosine monophosphate suppresses the transcription of proinflammatory cytokines via the phosphorylated c-Fos protein. Immunity 2009; 30: 372–383 42.
Ai J, Zhang R, Jiao J et al, Circulating microRNA-1 as a potential novel biomarker for acute myocardial infarction. Biochem Biophys Res Commun 2010; 391: 73–77.
Chim SS, Shing TK, Chiu RW et al, Detection and characterization of placental microRNAs in maternal plasma. Clin Chem 2008; 54: 482–490.
Starkey Lewis PJ, Dear J, Antoine DJ et al, Circulating microRNAs as potential markers of human drug-induced liver injury. Hepatology 2011; 54: 1767–1776.
Chinenov Y, Kerppola TK . Close encounters of many kinds: Fos-Jun interactions that mediate transcription regulatory specificity. Oncogene 2001; 20: 2438–2452.
Shaulian E, Karin M . AP-1 as a regulator of cell life and death. Nat Cell Biol 2002; 4: 131–136.
Ray N, Kuwahara M, Tsubone H et al, c-Fos suppresses systemic inflammatory response to endotoxin. Int Immunol 2006; 18: 671–677.
Ogino H, Inagaki Y, Kobayashi K et al, Reduced expression of cellular fos gene in peripheral blood mononuclear cells of patients with primary biliary cirrhosis. Int Hepatol Commun 1994; 2: 367–374.
Acknowledgements
This study was supported in part by Health and Labour Sciences Research Grants for Research on Measures for Intractable Diseases (from the Ministry of Health, Labour and Welfare of Japan), and a Grant-in-Aid for Challenging Exploratory Research (26670376) from JSPS.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on the Laboratory Investigation website
Rights and permissions
About this article
Cite this article
Katsumi, T., Ninomiya, M., Nishina, T. et al. MiR-139-5p is associated with inflammatory regulation through c-FOS suppression, and contributes to the progression of primary biliary cholangitis. Lab Invest 96, 1165–1177 (2016). https://doi.org/10.1038/labinvest.2016.95
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/labinvest.2016.95
This article is cited by
-
miR-139-5p sponged by LncRNA NEAT1 regulates liver fibrosis via targeting β-catenin/SOX9/TGF-β1 pathway
Cell Death Discovery (2021)
-
RNA sequencing reveals changes in the microRNAome of transdifferentiating hepatic stellate cells that are conserved between human and rat
Scientific Reports (2020)
-
Laser capture microdissection: techniques and applications in liver diseases
Hepatology International (2019)