Abstract
The completion of the Human Genome Project and the development of genome-based technologies over the past decade have set the stage for a new era of personalized medicine. By all rights, molecularly trained investigative pathologists should be leading this revolution. Singularly well suited for this work, molecular pathologists have the rare ability to wed genomic tools with unique diagnostic skills and tissue-based pathology techniques for integrated diagnosis of human disease. However, the number of pathologists with expertise in genome-based research has remained relatively low due to outdated training methods and a reluctance among some traditional pathologists to embrace new technologies. Moreover, because budding pathologists may not appreciate the vast selection of jobs available to them, they often end up choosing jobs that focus almost entirely on routine diagnosis rather than new frontiers in molecular pathology. This review calls for changes aimed at rectifying these troubling trends to ensure that pathology continues to guide patient care in a post-genomic era.
Similar content being viewed by others
Main
When researchers sequenced the first full genetic code of a human being a decade ago, many predicted this remarkable feat would forever transform medicine.1 At last, we had within our grasp the biological recipe for human health—the standard that would allow us to uncover the genetic roots of disease. This, in turn, offered a new vision of what medicine could be: rather than grouping patients into broad disease categories and treating them accordingly, physicians of the future would ascertain the exact molecular basis of each patient's disease and apply a specifically targeted intervention. At the heart of this radically different model for patient care would be the application of high-throughput genomic technologies in individual patients. Until now, the steep costs of such innovative tools has prohibited their routine use in the clinic.
Not so any longer. Recent years have seen a precipitous decline in those costs, ushering in a new era of genome-based personalized medicine. Whereas the mammoth Human Genome Project came with a whopping $3 billion price tag, the cost of sequencing a human genome today has plummeted to less than $5000 (ref. 2) and is expected to drop even further in the next few years. Cheaper sequencing technologies have yielded impressive results in the laboratory, enabling researchers to identify the underlying genetic causes of rare autosomal disorders using whole-exome or whole-genome sequencing methods.3, 4, 5
Meanwhile, biotechnology companies have developed gene-based diagnostic and prognostic tools that are already having a major impact in the clinic. For example, in 2004, the California-based life science company Genomic Health began offering patients a $4000 assay called Oncotype Dx, which measures the expression of 21 genes to predict breast cancer recurrence6 and responsiveness to certain chemotherapies.7 The test influences treatment decisions for a significant number of women with early estrogen-receptor-positive breast cancer.8
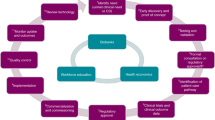
One would expect investigative pathologists to pioneer the development and application of such tools. Standing astride two historically divided realms, the best and brightest of these next-generation physician-scientists offer an unusually broad perspective and a unique set of skills perfectly suited for bridging the gap between basic science and medicine (Table 1 ). Transcending the traditional training of their predecessors, today's top investigative pathologists have acquired the rare ability to wed genomic tools with unique diagnostic skills and tissue-based pathology techniques for integrated diagnosis of human disease—a prerequisite for developing and exploiting individualized therapies for specific patients. They can also evaluate genetic and molecular information with a much greater appreciation for cellular context than most other scientists, taking into account the rich interplay between genes, cells and the microenvironment. Several recent studies led by modern molecular pathologists are a testament to their participation and leadership in genome-based research of human diseases.9, 10, 11, 12, 13, 14
Nevertheless, for the most part, pathologists have found themselves playing catch-up instead of leading the charge in this nascent field. Myriad factors contribute to this troubling trend. Traditional pathologists have avoided embracing innovation, holding to an outdated model of what pathology should be. Because these investigators are the ones who design pathology-training programs, their discomfort with new genomic and molecular technologies has trickled down to budding pathologists, leaving the next generation with critical gaps in their knowledge. Furthermore, up-and-coming pathologists tend to overlook the vast selection of research opportunities available to them, instead of choosing pathology careers that focus primarily on routine diagnosis rather than new frontiers in molecular pathology. Ultimately, this means a relatively small cadre of pathologists has the will and the necessary skills to fully integrate new molecular and genomic technologies into pathology research.
Compounding these internal struggles is an erroneous perception in the broader scientific community that genome-based tools are making traditional pathology skills less relevant. Building on the increasingly common assumption that the molecular and genetic makeup of a tumor will be its most important diagnostic and prognostic feature, large scientific consortia have forged ahead with ambitious genome sequencing projects that practically ignore the histopathological and immunohistochemical distinctions within so-called ‘common cancer types.’ They do so at their peril. More often than not, a single tumor type is not a monolithic entity but a grouping of disparate diseases with varying clinical characteristics and therapeutic responses.15, 16 Lumping them together as one disease will undoubtedly undermine the results of such endeavors.
Ideally, genome-based research on diseased tissues should rely on both histopathological and molecular classification systems, but the average scientist lacks expertise in histopathology and many pathologists are underskilled in genomic and molecular techniques. The situation calls for a new kind of scientist with extensive training in both disciplines—a molecular pathologist.
WHY PATHOLOGISTS?
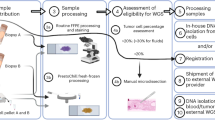
One of the greatest challenges in genomic research and the development of molecular diagnostic tools is the accrual of appropriate tissue samples. For example, many genome-based technologies work best with frozen tissue samples, and definitive validation of a particular disease marker or prognostic indicator depends on its detection in patients participating in prospective clinical trials. Because pathologists are the gateway to high-quality, well-characterized tissue specimens, they are in a unique position to help push these studies forward.
Furthermore, pathologists provide critical input in establishing the contextual foundation for all tissue-based studies because no other type of scientist has the necessary skills to identify tissues that accurately represents the patient's disease. The importance of that capability becomes clear when one considers the diagnostic limitations of molecular data. For example, 50–70% of melanomas17 and about 80% of melanocytic nevi18 harbor mutations in the proto-oncogene BRAF; however, melanomas are malignant, whereas nevi are benign skin lesions that rarely progress to more sinister forms. Thus, a molecular biologist who detects an oncogenic BRAF mutation in a patient's skin sample can only understand the biological meaning and clinical relevance of that result when a pathologist contextualizes it by identifying disease-relevant characteristics of cells in the same sample. Without that context, researchers attempting to understand and treat diseases through the molecular analysis of patient tissues would be shooting in the dark, frequently missing the target.
Moreover, investigative pathologists have the unique skills to help other scientists and clinicians make sense of complex tissue specimens comprised of many different normal and aberrant cell types. Biopsies from lung cancer patients, for instance, are notoriously heterogeneous, with actual tumor content ranging anywhere from 5 to 90%.19 Searching for lung cancer-associated genetic abnormalities, including EGFR mutations, in such samples is like looking for the proverbial needle in a haystack. Complicating matters further is the genetic heterogeneity within bona fide tumor cells—a problem highlighted by the efforts to detect drug-resistant EGFR mutations amidst a sea of more common drug-sensitive EGFR anomalies in an individual lung tumor.20 Investigative pathologists have the singular expertise to dissect such complex tissue samples using a full range of microscopy-based and molecular approaches, yielding significantly greater diagnostic power than any one strategy alone.
A natural consequence of their key role in disease diagnosis is the web of cross-disciplinary connections investigative pathologists create between experts who might never otherwise interact. A surgeon relies on them for the diagnosis of the disease; a laboratory investigator needs them to access correctly identified diseased tissue for genetic and molecular studies. Acting as a bridge between the two, investigative pathologists are in an ideal position to facilitate collaborative projects and, in many cases, form the glue that holds team-based transformational research together. In fact, we would dare to say that pathologists are uniquely placed to bring scientists and clinicians together in both clinical and research settings, serving as consultants to both groups while focusing the efforts of all parties on vexing biologic problems of clinical importance.
With this in mind, both academia and industry, eager to make the dream of personalized medicine a reality, have awakened to the growing importance of genetically and molecularly trained pathologists. Centers and departments devoted exclusively to molecular or translational pathology are springing up at research institutions and within biotechnology and pharmaceutical companies across the country. Pathologist-scientists can find particularly satisfying positions at these centers, where part of the central mission is to intergrate diagnostic skills with scientific insights. By embracing both areas of expertise rather than choosing between the two, translational pathologists face an unprecedented array of opportunities to participate in what promises to be one of the most critical scientific endeavors of our time.
BUILDING THE NEXT GENERATION OF MOLECULAR PATHOLOGISTS
Such advances certainly represent a step in the right direction, but they are still the exception rather than the rule for most pathology training programs. If the field as a whole is to remain relevant in a post-genomic era, we must focus on developing many more molecularly trained pathologists, and those efforts must come from the top down. Established pathologists must move beyond their comfort zones to try new technologies and expand their knowledge base. Training programs for up-and-coming pathologists must become more flexible, encouraging greater exploration of scientific disciplines outside the boundaries of traditional pathology. Course offerings and requirements must begin to include training in genome-based and molecular tools, not just for a few superstars but for all who would embark on a career in pathology.
Expanding training requirements to incorporate these new and developing areas will be a major challenge, particularly if morphologic and interpretive skills are to be retained in parallel. Possible solutions range from extending training time through the development of new tracks (eg anatomic pathology/translational molecular pathology). Another approach might involve the creation of new molecular pathology fellowship training programs that focus less on classical clinical genetics and more on modern genomic analysis. Changes of this type will require open-mindedness, broad input and the development of consensus among our professional groups.
The field of bioinformatics is just one of several areas in which investigative pathologists will have to master new skills. Now that whole genome sequencing has become more affordable and relatively quick, managing and interpreting the deluge of sequencing data coming out of those efforts is becoming a major challenge.21 The cost of outsourcing data analysis and validation is about $10–20 000 per genome, and even then, investigators heading up genome-based research projects must have at least some familiarity with bioinformatics in order to make sense of the results. However, most pathology training programs fail to offer even the most basic bioinformatics training to their students. Clearly, this must change.
Also, we must do a better job of inspiring the next generation to choose careers in molecular pathology by presenting them with their full range of options without oversimplifying the pros and cons of particular career paths. For instance, many young investigators think of academic and industry careers as polar opposites. According to received wisdom, researchers in the industry earn more money, work at a more rapid pace, have greater potential for developing products with direct clinical relevance and enjoy greater access to resources than their academic counterparts. Meanwhile, academic investigators may spend more time looking for research funding, but they also tend to have more autonomy, exert greater control over the scientific questions they pursue, enjoy better job security, have a stronger sense of ownership over their own work and receive much more public recognition for their findings than industry scientists. Although these generalizations are obviously valid when one paints academia and industry with broad strokes, a closer look at the diversity within each realm suggests it may be more useful to imagine a continuum of benefits and challenges linking academic and industry jobs, with significant overlap in between (Figure 1).
By sorting academic careers according to their funding structures, for example, one uncovers a more nuanced picture than that originally imagined (Figure 2). The pros and cons of an independently funded tenure-track position are significantly different from those of an investigative pathologist who relies on collaborators for funding: The former has much greater autonomy, ownership of ideas and public recognition for scientific accomplishments than the latter. somewhere in between are pathologists who are funded ‘semi-independently’ (most often, researchers running pathology cores). They usually have some authority and avoid having to write the major portion of grant proposals but lack the ability to control their research direction. Thus, in practical terms, investigative pathologists working under the ‘dependent funding’ or ‘semi-independent funding’ model may have much more in common with their industry counterparts than they would with an academic peer operating under the ‘independent funding’ model.
Blurring the line even further, pathology jobs at pharmaceutical and biotechnology companies have recently become much more comparable with those at academic institutions in terms of scientific merit. Whether an investigative pathologist is interested in pursuing basic, translational, clinical or collaborative research, he or she can find challenging and exciting projects to work on in either an academic or an industry setting. More and more companies are establishing specialized pathology groups that have increasingly important roles in drug development, animal model design, biomarker discovery and the creation of diagnostic and prognostic tools for correlative science. Furthermore, the stigma once associated with transferring from academia to industry has nearly vanished, and it is becoming increasingly common for scientists to shift back and forth between the two.
With a variety of career options available to them, next-generation pathologists have many incentives to participate in scientific research, including intellectual stimulation, interaction with other specialties, a broadening of professional perspective, facilitated access to new technologies, potential career redirection, exposure to new and different ideas and greater access to funds. Those who accept this challenge and acquire the right set of skills can rest assured that the need for their expertise is growing alongside technological advances in biomedical research. Genomic and molecular tools will undoubtedly improve the diagnosis and treatment of illness in years to come, but the ability to use sophisticated new technologies is not the same as recognizing, understanding and thereby better characterizing a given disease. This is perhaps best illustrated by the recent discovery that certain types of ovarian cancer are not ovarian at all; rather, they arise from displaced epithelial cells that originate in the fallopian tubes—an astonishing finding overlooked by the scientific community for decades before investigative pathologists finally uncovered the truth.22, 23 Such cautionary tales serve to remind us that even the most advanced technology cannot help us unless we combine it with the accurate contextual information that only a well trained investigative pathologist can provide. Clearly, the world needs this rare breed of physician-scientists, now more than ever.
References
Wade N . Reading the book of life; now, the hard part: putting the genome to work. The New York Times 2000;F1.
Drmanac R, Sparks AB, Callow MJ, et al. Human genome sequencing using unchained base reads on self-assembling DNA nano-arrays. Science 2010;327:78–81.
Ng SB, Bigham AW, Buckingham KJ, et al. Exome sequencing identifies MLL2 mutations as a cause of Kabuki Syndrome. Nat Genet 2010;42:790–793.
Lupski JR, Reid JG, Gonzaga-Jauregui C, et al. Whole-genome sequencing in a patient with Chacot-Marie-Tooth neuropathy. New Engl J Med 2010;362:1181–1191.
Roach JC, Glusman G, Smit AF, et al. Analysis of genetic inheritance in a family quartet by whole-genome sequencing. Science 2010;328:636–639.
Paik S, Shak S, Tang G, et al. A multigene assay to predict recurrence of tamoxifen-treated, node-negative breast cancer. New Engl J Med 2004;351:2817–2826.
Paik S, Tang G, Shak S, et al. Gene expression and benefit of chemotherapy in women with node-negative, estrogen receptor-positive breast cancer. J Clin Oncol 2006;24:3726–3734.
Lo SS, Mumby PB, Norton J, et al. Prospective multicenter study of the impact of the 21-gene recurrence score assay on medical oncologist and patient adjuvant breast cancer treatment selection. J Clin Oncol 2010;28:1671–1676.
Shah SP, Morin RD, Khattra J, et al. Mutational evolution in a lobular breast tumour profiled at single nucleotide resolution. Nature 2009;461:809–813.
Shah SP, Köbel M, Senz J, et al. Mutation of FOXL2 in granulosa-cell tumors of the ovary. N Engl J Med 2009;360:2719–2729.
Wiegand KC, Shah SP, Al-Agha Om, et al. ARID1A mutations in endometriosis-associated ovarian carcinomas. N Engl J Med 2010;363:1532–1543.
Campbell PJ, Yachida S, Mudie LJ, et al. The patterns and dynamics of genomic instability in metastatic pancreatic cancer. Nature 2010;467:1109–1113.
Geyer FC, Weigelt B, Natrajan R, et al. Molecular analysis reveals a genetic basis for the phenotypic diversity of metaplastic breast carcinomas. J Pathol 2010;220:562–573.
Kim JH, Dhanasekaran SM, Prensner JR, et al. Deep sequencing reveals distinct patterns of DNA methylation in prostate cancer. Genome Res 2011;21:1028–1041.
Aparicio SA, Huntsman DG . Does massively parallel DNA resequencing signify the end of histopathology as we know it? J Patho 2010;220:307–315.
Ashworth A, Lord CJ, Reis-Filho JS . Genetic interactions in cancer progression and treatment. Cell 2011;145:30–38.
Davies H, Bignell GR, Cox C, et al. Mutations of the BRAF gene in human cancer. Nature 2002;417:949–954.
Pollock PM, Harper UL, Hansen KS, et al. High frequency of BRAF mutations in nevi. Nat Genet 2002;33:19–20.
Jänne PA, Borras AM, Kuang Y, et al. A rapid and sensitive enzymatic method for epidermal growth factor receptor mutation screening. Clin Cancer Res 2006;12:751–758.
Jänne PA . Challenges of Detecting EGFR T790M in gefitinib/erlotinib-resistant tumours. Lung Cancer 2008;60 (Suppl 2):S3–S9.
Mardis ER . The $1000 genome, the $100,000 analysis? Genome Med 2010;2:84.
Levanon K, Crum C, Drapkin R . New insights into the pathogenesis of serous ovarian cancer and its clinical impact. J Clin Oncol 2008;26:5284–5293.
Crum CP, Drapkin R, Miron A, et al. The distal fallopian tube: a new model for pelvic serous carcinogenesis. Curr Opin Obstet Gynecol 2007;19:3–9.
Acknowledgements
We thank the journal's reviewers for insightful and significant contributions to the manuscript. We also gratefully acknowledge the USCAP Education Committee, including Dr J Goldblum and Dr D Huntsman, for hosting a course on investigative pathology careers, which formed the basis of this review. Finally, we thank our financial supporters: the National Institutes of Health (RO1DK072000 and R01CA131945) and the Prostate Cancer Foundation.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Molecular pathologists have the rare ability to wed genomic tools with diagnostic skills and tissue-based pathology techniques for integrated diagnosis of human disease. However, the number of pathologists with expertise in genome-based research is relatively low. This situation must be rectified so that pathology can guide patient care in the post-genomic era.
Rights and permissions
About this article
Cite this article
Berman, D., Bosenberg, M., Orwant, R. et al. Investigative pathology: leading the post-genomic revolution. Lab Invest 92, 4–8 (2012). https://doi.org/10.1038/labinvest.2011.147
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/labinvest.2011.147
Keywords
This article is cited by
-
MALDI Imaging mass spectrometry: current frontiers and perspectives in pathology research and practice
Laboratory Investigation (2015)
-
αVβ3 Integrin Regulation of Respiratory Burst in Fibrinogen Adherent Human Neutrophils
Cellular and Molecular Bioengineering (2014)
-
Melanoma genotypes and phenotypes get personal
Laboratory Investigation (2013)