Abstract
Previously we described that bone morphogenetic protein-7 (BMP7) could protect prostate cancer C4-2B cells from serum starvation-induced apoptosis via survivin induction. Here, for the first time, we identify Runx2 as a key regulator of survivin transcription. In C4-2B cells grown normally, suppression of Runx2 reduced survivin expression. Using ChIP assays, two regions of the survivin promoter, −1953 to −1812 (I) and −1485 to −1119 (II) encompassing consensus Runx-binding sites were examined. Runx2 was found to be associated with both regions, with a stronger affinity to region-I. In serum-starved cells neither region was occupied, but BMP7 restored association to region-II and not region-I. In reporter assays, transcription activity by BMP7 was significantly reduced when sequences including binding sites of region-II were deleted. Additionally, Runx2 expression was enhanced by BMP7 in these cells. Along with a strong survivin expression, a trend in increased Runx2 expression in human prostate cancer cells and tissues was noted. In the conditional Pten-knockout mouse, Runx2 level increased with growth of prostate tumor. The data define a novel role of Runx2 in regulating survivin expression in malignant epithelial cells and identify it as a critical factor in BMP signaling that protects cancer cells against apoptosis.
Similar content being viewed by others
Main
The family of bone morphogenetic protein (BMP) that belongs to the transforming growth factor-β superfamily, was originally identified as osteogenic factors.1 Soon it became evident that BMP also played important roles in multiple cellular processes such as cell growth, differentiation, migration, apoptosis, and in cancer.2 We were particularly interested in the role of BMP7 in prostate cancer. BMP7 expression was reported in human metastatic prostate cancer specimens3 and was most abundant in prostate cancer skeletal metastasis.4 Furthermore, regulation of BMP7 expression by androgen was described.5 Previously, we reported that in the Pten conditional deletion mouse model of prostate cancer6, 7 BMP7 protein expression increased with growth of the prostate adenocarcinoma8 and correlated with the induction of tumor-promoting heterotypic cell interactions.9 We also demonstrated that BMP7 could induce epithelial–mesenchymal transdifferentiation in PC-3 human prostate cancer cells and exert strong protection against stress-induced apoptosis in another aggressive prostate cancer cell line, C4-2B.8 In C4-2B cells under serum starvation, the effect of BMP7 was found to be mediated by survivin, a member of the inhibitor of apoptosis (IAP) protein family. Under serum starvation BMP7 maintained the level of survivin protein but not XIAP of the IAP family or any of the Bcl-2 family members examined in these cells.8
Survivin is considered as a cancer therapeutic target as aberrant high expression of survivin was documented in many different types of human cancer.10 It is thought that survivin overexpression might allow accumulation of mutations in transformed cells and thereby promote tumor progression. Transcription of the survivin gene that is prominent in the mitotic phase of the cell cycle is also regulated by various growth factors and cytokines.11
When we analyzed the human survivin promoter with the TF Search program,12 multiple potential binding sequences for transcription factors were apparent. Among them were sites that could serve to recruit Smad and Runx proteins. The Smad pathway is indeed a canonical pathway for BMP function and we showed earlier that introduction of a dominant-negative mutant of Smad-5 could inhibit the activity of the survivin promoter.13 In mammalian cells the Runx family, also known as the acute myeloid leukaemia (AML), core-binding factor-α (CBFα), or polyoma enhancer-binding protein-2α (PEBP2α) family, include three structurally similar members (Runx1, -2, and -3). Runx proteins were described to bind to a common non-DNA binding partner, CBFβ, to form a heterodimeric complex, the target of which appeared to be a conserved nucleotide sequence.14, 15 Despite their similarities, Runx proteins, however, were determined to play divergent biological roles that were consistent with the distinct phenotypes observed in the corresponding gene knockout in mice.16
There is evidence that activated Smad proteins may interact with Runx2, the ‘master’ transcription factor for differentiation of osteoblasts, in regulating transcriptional activity.17, 18, 19 Considering these sets of existing knowledge, and our earlier demonstration that BMP-induced Smad signaling is indeed involved in the transcriptional activation of the survivin gene in prostate cancer cells,8 we hypothesized that Runx2 might also be critical in regulation of survivin expression in malignant epithelial cells and consequently required for the antiapoptotic effect of BMP7 in the C4-2B test system. Hence, among the Runx members, we wished to focus on Runx2 for this study. Here, we describe for the first time that Runx2 interacts with the survivin promoter in vivo and regulates the transcription of the survivin gene in C4-2B cells grown under either normal conditions or under serum starvation but with exposure to BMP7. We also show that BMP7 regulates the expression of Runx2 in these cells and that in the conditional Pten deletion mouse model,6, 7 similar to our previous observation of survivin overexpression,13 there is strong correlation of increased Runx2 expression with prostate tumor growth. In the human prostate cancer specimens, we confirm upregulation of survivin expression and a readily detectable Runx2 expression.
MATERIALS AND METHODS
Materials
Human recombinant BMP7 protein (generously provided by Dr A Hari Reddi of University of California Davis or purchased from R&D) was diluted in 0.1% serum medium and used at a concentration of 50 ng/ml in all the experiments, unless specified otherwise. The survivin gene promoter constructs (pLuc1430 and pLuc649)20 were generously provided by Dr Dario Altieri of the University of Massachusetts. The pAML1-ETO vector21 was provided by Dr Baruch Frenkel of the University of Southern California. The pCMV5 empty vector was used as a control in all experiments using pAML1-ETO.
Cell Culture and Transient Transfection
C4-2B cells were maintained in culture medium with 10% fetal bovine serum (FBS) as described previously.8 For serum starvation, the cells were cultured in medium containing 0.1% FBS. Transient transfection was performed with either Tfx-20/50 reagent (Promega) or Lipofectamine 2000 (Invitrogen) according to the manufacturer's instruction.
Small Interfering RNA (siRNA) Treatment
C4-2B cells were seeded in six-well plates at 1.5 × 105 cells per well and grown in the antibiotic-free medium for 2 days to reach 30–50% confluence. The cells were transfected with siRNA duplexes at final concentrations of 50 nmol/ml. The control siRNA duplex (sense 5′-UUCUCCGAACGUGUCACGUdTdT, 3′-fluorescein-labeled; antisense 5′-ACGUGACACGUUCGGAGAAdTdT) and siRNA specific for Runx222 (sense 5′-CUCUGCACCAAGUCCUUUUdTdT; antisense 5′-AAAAGGACUUGGUGCAGAGdTdT) were purchased from Qiagen. The siRNA for human survivin (sense 5′-CUGGACAGAGAAAGAGCCAdTdT; antisense 5′-UGGCUCUUUCUCUGUCCAGdTdT) was produced by Ambion.
Terminal Deoxynucleotidyltransferase-Mediated dUTP Nick-End Labeling (TUNEL) Assay
The TUNEL assay was performed to measure apoptotic cell proportion using the APO-BrdU kit (Phoenix Flow Systems). Cell death was also monitored by light microscopy every day during cell culture for a period of 24–96 h after treatment or transfection.
Chromatin Immunoprecipitation (ChIP) Assay
Cross-linking ChIP assay was performed using the ChIP-IT Express kit (Active Motif) following the manufacturer's instruction. Formaldehyde cross-linking was performed for 10 min. Nuclei were pelleted and chromatin was sheared by sonication using VirTis Virsonic 600. Approximately 9 μg of sheared chromatin was immunoprecipitated with 4 μg anti-PEBP2α antibody (M-70X; Santa Cruz Biotechnology) or normal rabbit IgG using magnetic protein-G beads. For native ChIP, the experiments were conducted in the same way except that the formaldehyde cross-linking step was omitted.23 The primers designed to amplify various regions of the human survivin gene promoters were as follows: 5′-CGGTTAGCGAGCCAATCAGCA-3′ and 5′-ACCATACTACCCCCGAAATACTTCATTCT-3′ (region-I); 5′-GCAATTATCTTTTATTTAAATTGACATCG-3′ and 5′-GCCCCCATTCTTTAATACAGTAACTT-3′ (region-II); and 5′-GCGTTCTTTGAAAGCAGTCGAG-3′ and 5′-GGAGCGCACGGCCCTCTTA-3′ (a control region). All ChIP assays were performed at least three times and the most representative results are illustrated in the figures.
Generation of Luciferase Reporter Plasmids
The truncated fragment of the survivin promoter (pLuc1270) was generated by PCR amplification using pLuc1430 as template and using a forward primer 5′-GACACCGACGGATCCAGGCTGAGGCAGGAGAATCGC-3′ and reverse primer 5′-GAATTCTAGAAGCTTAAATCTGGCGGTTAA-3′. The DNA fragment was cloned using the TA Cloning kit with One Shot TOP10F′ Chemically Competent Escherichia coli and subcloned to the BamHI and HindIII sites of pLuc1430.
Luciferase Reporter Assay
C4-2B cells were seeded in six-well plates at 2 × 105 cells per well and grown for 2–3 days. For serum starvation, cells were pretreated in 0.1% serum medium for 24 h prior to cell transfection. When cells reached 70–80% confluence, 1 μg of each reporter constructs and 0.5 μg of pSV-β-galactosidase control vector were cotransfected. When needed, 0.5–1.0 μg of pAML1-ETO or 1 μg of empty vector was also transfected into the cells along with the reporter constructs and the pSV-β-galactosidase control. The transfected cells were then lysed in 1 × lysis buffer included in the Luciferase Assay System (Promega) at 24 h post transfection. The lysed supernatants were used for either luciferase assay or β-galactosidase assay as described,13 and the results were normalized to the activity of β-galactosidase.
The expression of AML1-ETO was confirmed by semi-quantitative RT-PCR following a reported procedure.24 The primers used were the following: forward, AML1-ES: 5′-GAGGGAAAAGCTTCACTCTG-3′ and reverse, ETO-EA: 5′-TCGGGTGAAATGTCATTGCC-3′, and the thermocycle setup included denaturation at 95C° for 30 s, annealing at 58C° for 30 s, and polymerization at 72C° for 90 s for 40 cycles.
RNA Extraction, cDNA Synthesis, and Comparative Quantitative PCR
Total RNA was extracted using the TRIzol Reagent (Invitrogen) following the manufacturer’s protocol and genomic DNA was digested by DNaseI (Ambion). cDNAs were synthesized with 2 μg of total RNA using the iScript cDNA Synthesis kit (Bio-Rad). Quantitative PCR analyses for Runx2 expression were performed using forward primer 5′-CCTCGGAGAGGTACCAGATG-3′ and reverse primer 5′-AGGTGAAACTCTTGCTGCA-3′, using the Stratagene Mx3000P PCR system. The reaction mix contained 13 μl of 2 × Brilliant SYBR Green QPCR Master Mix (Stratagene) or Faststart Universal SYBR Green Master Mix (Roche) with 1 μl of cDNA in a total volume of 25 μl. GAPDH was used for each sample as control for normalization.
Preparation of Whole-Cell Lysates of Human Prostatic Cell Lines or Mouse Prostate Tissues and Western Blot Analysis
Whole-cell lysates of prostatic cell lines and mouse prostatic tissues were prepared following the protocols described previously.8 The antibodies used for Western blot analysis were rabbit anti-PEBP2αA and goat anti-Actin (Santa Cruz Biotechnology), and rabbit anti-survivin (Novus Biologicals).
Human Prostate Cancer Specimens and Immunohistochemistry
Tissue microarrays were constructed using a manual arrayer (Beecher Instruments, WI, USA) from 59 cases of prostate cancer, all adenocarcinomas, diagnosed and graded at the Department of Pathology at University of Iowa Hospitals and Clinics, Iowa City, from 2001 to 2003. Core tissue biopsies (diameter 2.0 mm, height 3–4 mm) were taken from morphologically representative regions of formalin-fixed, paraffin-embedded blocks (45 × 20 mm) containing original tissue and arrayed into a regular recipient paraffin block.
An immunoperoxidase reaction for survivin and Runx2 proteins was performed on each case using the ABC method. Both primary antibodies for survivin (Lab Vision) and Runx2 (Santa Cruz Biotechnology) were diluted to 1:100 for the analysis. Antigen unmasking was accomplished using citrate buffer (pH 6.0) and using a heating procedure (microwave or preheated steamer). Slides were allowed to cool at room temperature. Nonspecific background staining was prevented by application of horse or goat serum. Sections were covered with primary antiserum, incubated, and rinsed. Counterstaining by 10% Harris hematoxylin was performed for 3 min. Negative-control slides were prepared by substituting rabbit immunoglobulin. Immunoreactivity for benign and malignant prostate epithelium, as well as stromal cells, was evaluated using both proportion and intensity scores graded from 0 to 3 for a total quantitative score range of 0–6. Intensity staining was graded as follows: 0=no staining; ranging to 3=strongly positive. Proportion staining was graded as follows: 0=no staining, 1=1–33% of cells positive, 2=33–66% of cells positive, and 3=over 66% of cells positive. Comparison of expression between malignant and benign epithelium, by assigned grade, and between epithelium and intervening stroma was performed for both proteins.
Statistical Analysis
All experiments were performed in triplicate and repeated at least three times. Statistical comparisons were made by unpaired, two-tailed t-test.
RESULTS
Modulation of Survivin Expression in C4-2B Cells and their Sensitivity to Apoptosis by AML1-ETO or Runx2 siRNA
Previously we described that survivin expression was elevated by BMP7 in serum-starved C4-2B cells, and that increased survivin expression was associated with antiapoptotic effect of BMP7.8 Thus we were motivated to further investigate the regulation of survivin expression in this cell line. Analysis of the upstream −2.8-kb fragment of the human survivin gene promoter showed the presence of multiple putative Runx2-binding sites within this region.12, 14 To investigate the possible involvement of Runx2 in survivin expression and subsequent cell apoptosis, we first examined the effect of AML1-ETO on C4-2B cells cultured under normal culture conditions. AML1-ETO is encoded by a fusion gene resulting from the translocation (t8:21), which is commonly associated with AML.21 It represses the expression of Runx domain targets by functioning as a constitutive transcription repressor.25 C4-2B cells were transfected with various amounts of the AML1-ETO expression vector and the levels of survivin in the transfected cells were determined. As illustrated in Figure 1a, while transfection with 0.5 μg of the plasmid did not cause significant reduction in survivin expression, the effect was robust when 1 μg of the plasmid DNA was used. The results also showed that pAML1-ETO transfection did result in the expression of the gene in the cells, and that Smad activation or the level of expression of the Runx2 protein was not affected by AML1-ETO expression to any significant extent (Figure 1a). Consistent with the downregulation of survivin expression, the apoptotic rate of the cells treated with 1 μg plasmid increased to 60% as compared with 10% of the untreated cells (Figure 1b).
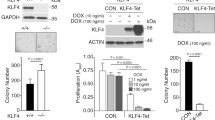
Downregulation of survivin expression and induction of cell apoptosis by AML1-ETO or Runx2 siRNA in C4-2B cells. (a) Effect of AML1-ETO on survivin expression. C4-2B cells at 50–70% confluence were transfected with 0.5 or 1 μg pAML1-ETO per well in six-well plates. The expression levels of survivin in the cells at 48 h post transfection were analyzed by Western blotting and compared with those in control cells transfected with the pCMV5 empty vector. Equal loading of the cells lysates was shown using actin as control. The expression of AML1-ETO in the transfected cells was determined by semi-quantative RT-PCR using GAPDH expression as control. (b) Induction of apoptosis by expression of AML1-ETO. C4-2B cells transfected with the indicated amount of pAML1-ETO were subjected to TUNEL assay. Compared with control cells or cells transfected with 0.5 μg pAML1-ETO, cells treated with 1 μg of pAML1-ETO had significantly higher apoptotic rate (P<0.05). (c) Reduction of survivin expression in cells treated with Runx2 siRNA (siRunx2) or survivin siRNA (siSurv). C4-2B cells at 30–50% confluence were transfected with siSurv, siRunx2, or control siRNA (siCtrl) at a final concentration of 50 nmol/ml. The protein levels of Runx2 and survivin were examined by Western blotting. Treatment of siRunx2 led to significantly decreased levels of Runx2 and survivin, while siSurv reduced the protein level of survivin but not Runx2. (d) Induction of cell apoptosis by siRunx2 or siSurv. The apoptotic rates of the cells at 4 days post transfection were determined by TUNEL assay.
Considering that AML1-ETO exerts repression on all tested Runx-dependent genes,26 and that Runx1 and Runx3 proteins are also expressed in C4-2B cell, we further used the RNAi technique to examine the specific effect of Runx2. As shown in Figure 1c, at 72 or 96 h post transfection of Runx2 siRNA, Runx2 level was significantly decreased and so was of survivin. In addition, cells transfected with Runx2 siRNA underwent apoptosis to an extent similar to that of cells transfected with survivin siRNA (Figure 1d). These results implied that Runx2 might be pivotal for survivin expression and therefore potentially critical to block apoptosis of C4-2B cells.
Runx2 Association with the Survivin Promoter in C4-2B Cells
Multiple putative Runx-binding sites on the −2.8-kb fragment of the survivin promoter were identified by the TFsearch programme,12 using default settings (Figure 2a). ChIP assays were performed in order to examine the association of Runx2 with the promoter. Highly specific primers could be designed to amplify two different regions each containing a fully consensus binding site. These were the −1953 to −1812 region containing the −1853 to −1848 binding site (region-I) and the −1485 to −1119 region encompassing the −1314 to −1309 binding site (region-II). Region-II also harbored an 89% homologous sequence at −1391 to −1386. Specific primers were used to amplify a region (−267 to −76) to serve as control for the binding study. The specificity of the primer sets was analyzed using the UCSC BLAT server27 and further confirmed by sequencing the PCR products. The results obtained with ChIP assays supported in vivo occupancy of the survivin promoter by Runx2 in C4-2B cells. For example, when cross-linking ChIP was performed on these cells cultured in full serum medium, interaction of Runx2 with region-I and II was clearly evident (Figure 2b). No significant occupancy of Runx2 at the control region was detected under the conditions used. Native ChIP was then performed to examine the relative affinities of the two binding regions detected by the cross-linking ChIP. Primarily, occupancy of region-I by Runx2 was detected in the native ChIP assay (Figure 2c), implying that interaction with this site was relatively stable, while that with region-II was not strong enough to survive the harsh sonication procedure. Since native ChIP was performed in a way similar to that of cross-linking ChIP, except for the fixation step, the results also indicated that chromosome shearing was sufficient to separate region-I and II.
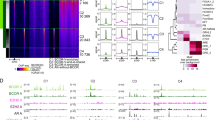
In vivo occupancy of the human survivin gene promoter by Runx2 in C4-2B cells cultured in full serum medium. (a) A line diagram of the survivin promoter (−2.8 kb) indicating the positions of the various potential Runx2-binding sites. The sequence is numbered relative to the ATG translation initiation site and each site is marked by the number of the first nucleotide. The number in the parentheses indicates the weighted score of sequence conservation at that site calculated.12 Underscored region-I and II were analyzed by ChIP assays for which a region marked as Ctrl served as negative control for the binding assays. The nucleotide sequence encompassing region-I and II is presented underscoring the theoretical Runx- and Smad-interacting elements. Runx sites are indicated in bold and underlined with the score of conservation in the parenthesis. The unmatched nucleotide is italicized. Smad elements,29, 34 in italics, are also underlined. (b) Detection of Runx2 binding to region-I and II by the cross-linking ChIP assay. The immunoprecipitates were subjected to PCR analysis to determine the binding affinity. (c) Detection of Runx2 binding to region-I by native ChIP assay.
Runx2-Dependence of the Effect of BMP7 on Survivin Expression
In cells cultured in 0.1% serum medium, BMP7 treatment upregulated survivin expression and protected the cells from apoptosis, which was consistent with our previous results.13 However, in the presence of optimal AML1-ETO, survivin expression was greatly inhibited (Figure 3a) and protective effect of BMP7 was counteracted (Figure 3b), suggesting involvement of Runx proteins. We performed cross-linking ChIP assays with C4-2B cells, which were treated with BMP7 for 1 day with those of the untreated cells. The ChIP results showed that the extent of interaction of Runx2 with either region-I or II was below the level of detection in the serum-starved cells (Figure 3c). However, when these cells were treated with BMP7, occupation by Runx2 of region-II but not I could be demonstrated. Since the protein level of Runx2 was determined to be similar in both untreated cells and 1 day BMP7-treated cells (data not shown), the results indicated that exposure to BMP7 enhanced the association of Runx2 with region-II. Considering that Runx2 was found to interact with both regions under normal culture conditions, these results also indicated that regulation of survivin transcription might be different under varying cellular environment.
Effect of BMP7 on survivin expression and binding of Runx2 to survivin promoter in serum-starved C4-2B cells. (a) The effect of BMP7 on survivin expression. C4-2B cells were pretreated with medium containing 0.1% serum for 24 h followed by supplementation of BMP7 with or without transfection of pAML1-ETO. The expression levels of survivin protein in the cells at 48 h post transfection were compared with that of the corresponding controls. Equal loading of the cells lysates was determined using actin levels. Semi-quantitative RT-PCR was used to determine the expression of the transfected AML1-ETO gene. (b) Suppression of the antiapoptotic effect of BMP7 by AML1-ETO. C4-2B cells cultured in 0.1% serum with or without BMP7 treatment were transfected with the indicated amount of pAML1-ETO and subjected to TUNEL assay. BMP7 treatment significantly reduced cell apoptosis (P<0.05). However, cells treated with 1 μg of pAML1-ETO showed significantly higher apoptotic rate (P<0.01) compared with control cells or cells transfected with 0.5 μg pAML1-ETO, indicating that AML-ETO had a counteracting effect and the effect was dose-dependent. (c) Increased occupancy of region-II of the human survivin promoter by Runx2 in BMP7-treated cells. Serum-starved C4-2B cells were cultured with or without BMP7 and cross-linking ChIP was performed to examine the Runx2 binding to region-I (upper panel) and region-II (lower panel) of the survivin promoter, respectively. Runx2 binding was only evident for region-II of the promoter in cells stimulated with BMP7. (d) Luciferase reporter constructs that contained various lengths of survivin promoter (pLuc1430, pLuc1270, and pLuc649) are shown on the upper panel. The promoter pLuc1430 retained the Runx2-binding sites of the region-II as marked. Effect of BMP7 on the activity of the promoter constructs in serum-starved C4-2B cells is illustrated in the lower panel. The activity of each construct was analyzed in serum-starved cells with or without supplementation of BMP7. The pAML1-ETO plasmids were also transfected as indicated to examine the effect of Runx2 binding on the promoter activity. The cells were cotransfected with β-galactosidase and all activities are presented relative to the activity of pLuc1430 after normalization with respect to β-galactosidase activity. For pLuc1430 plasmid, P-values determined were as follows: ±BMP7 (<0.01); +BMP7, ±0.5 μg AML1-ETO (<0.01); and +BMP7, ±1 μg AML1-ETO (<0.001). For the pLuc 1270 plasmid, the corresponding values were <0.01, not significant, and <0.05, and for pLuc649, not significant. Transfection with ptkLuc (a construct with a minimal promoter) was used as a negative control in the assays.
The effect of Runx2 on the survivin promoter was further investigated with luciferase reporter assays. A luciferase reporter construct, pLuc1430,20 was used in our previous study to assess the effect of Smad signaling on surviving-promoter activity.13 To further evaluate the sequences immediately downstream of −1430, we constructed pLuc1270 in which the potential Runx- and Smad- binding sites of region-II were deleted (Figure 3d, upper part). A shorter construct, pLuc649, was also used in this study. As illustrated in the lower part of Figure 3d, the transcriptional activity of pLuc1430 increased over 200% after 24 h of BMP7 treatment in serum-starved cells, while under similar conditions pLuc1270 displayed relatively much smaller response, and pLuc649 practically failed to respond to BMP7. This observation suggested that the −1430 to −1271 region might be important for the effect of BMP on survivin transcription. As Smad-interacting sequences are juxtaposed in this region and since we showed earlier that activated Smad was indeed involved in BMP-induced survivin expression,13 we introduced AML1-ETO to the luciferase reporter assays to examine the role of Runx2 in response to BMP7 treatment in the serum-starved cells (Figure 3d). When AML1-ETO was present, the effect of BMP7 on the promoter activity of pLuc1430 was indeed strongly inhibited in a dose-dependent manner. AML1-ETO had no significant negative effect on the activity of pLuc1270 at 0.5 μg, although there was a slight inhibitory effect noted at 1 μg concentration, and the activity of pLuc649 was not affected by AML1-ETO. These results suggested a lack of functionally prominent Runx interaction sites downstream of −1270 on the survivin promoter.
Induction of Runx2 Expression by BMP7 in C4-2B Cells
It was reported that Runx2 expression could be positively influenced by BMP2 or BMP7 in mouse chondrocyte and myoblast cells,28, 29, 30, 31 and in human pancreatic cancer cells.32 Interestingly, in differentiating fibroblasts Runx2 was reported to bind to a repressive element within its own promoter, to autoregulate gene transcription.30, 31 We examined the effect of BMP7 on Runx2 expression in serum-starved C4-2B cells by quantitative PCR. During the first 4 h post BMP7 treatment, the Runx2 mRNA level peaked at 1 h and was 3.2-fold of that of the control, and then appeared to decline to almost normal levels (Figure 4a). It appeared from Western blot analysis that at this 1-h time point there was an increase in the level of Runx2 protein (Figure 4b). The effect of BMP7 dose on Runx2 protein expression was also examined at 1 h post exposure; the effect appeared to be dose dependent, with the most robust effect observed at 50 ng/ml (Figure 4b).
Stimulation of Runx2 expression by BMP7 in serum-starved C4-2B cells. (a) Examination of Runx2 mRNA levels by quantitative real-time PCR at early time points after BMP7 treatment. The fold increase was determined by 2-Δ Ct analysis and normalized to GAPDH expression. The increase in Runx2 at 1 h post treatment was significant (P<0.05). (b) The dose-dependent effect by BMP7. BMP7 protein at the indicated concentrations was supplemented to C4-2B for an hour after 24 h of serum starvation and Runx2 protein levels were analyzed by Western blotting. (c, d) Examination of changes in Runx2 mRNA or protein levels, respectively, at 2-day intervals during the 6 days of BMP7 treatment.
In our previous study, we found that the elevated survivin protein level continued to persist up to 6 days of BMP7 treatment in starved C4-2B cells.8 Hence we continued to investigate Runx2 expression every two days up to 6 days post BMP7 treatment. The mRNA level of Runx2 was highest (2.7-fold compared with that of the control cells) on day 2, and then declined to become similar to that of the control cells by day 6 (Figure 4c). The protein level of Runx2 was not significantly different from the controls at day 2 following exposure to BMP7, but increased maximally at day 4 (Figure 4d). Thus, there appeared to be a complex regulation of Runx2 expression by BMP7 in the C4-2B cells. The effect might also be complicated by constant adjustment of the cells to the stress from serum starvation as well by the factors directly related to Runx2 protein modification, stability, degradation as well as autoregulation of Runx2 gene expression. Although an overall cell survival is relatively strong in the serum-starved condition in the presence of BMP7 as compared with that in the absence of BMP7, there was still progressive loss of cells in both situations over the period of observation (6 days). In the population of control cells that continued to survive under serum starvation, a trend of increased Runx2 protein expression level was also noted (Figure 4d).
Expression of Runx2 in Prostate Cancer Cells and Tissues
Considering the important role of Runx2 on survivin expression and cell survival, we examined Runx2 expression in various human prostatic cell lines. As shown in Figure 5a, expression in immortalized ‘normal’ MLC and benign BPH-1 prostate cell lines was practically undetected, whereas all prostate cancer cell lines examined, in particular C4-2B and LNCaP, showed robust expression.
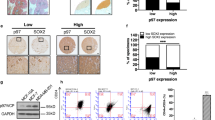
Analysis of Runx2 and survivin expression in prostate cancer cells. (A) Protein expression in various human prostate cancer cell lines in comparison with immortalized normal prostate epithelial (MLC) and benign hyperplastic (BPH-1) cells. For each cell line approximately 40 μg of whole-cell lysates were assayed by Western blotting and actin served as loading control. (B) Examples of immunostaining in benign and malignant human prostate cancer tissues: (a, b) Survivin (case 1), strong staining of malignant epithelium relative to adjacent benign epithelium and stroma; (c, d) Survivin (case 2), similarly strong staining of malignant epithelium relative to benign epithelium and stroma; (e, f) Runx2 (case 1), moderate staining of malignant epithelium with weak stromal staining; (g, h) Runx2 (case 2), comparable moderate staining of malignant epithelium, with weak stromal staining. Benign epithelium displayed low-to-moderate staining: (a, c, e, g) magnification × 25 and (b, d, f, h) magnification × 100. (C) Correlation of Runx2 expression with tumor growth in the conditional Pten deletion mouse model of prostate cancer. Runx2 expression in the anterior prostate tissues of the knockout mice (KO) and in the corresponding normal prostate lobe of the littermate controls (c) were analyzed by Western blotting. The tissues were obtained from mice at age ranging from 1.6 to 11 months. (D) Detection of increased Runx2 expression by Western blotting in tumors from all the lobes from an 11-month old mouse as compared with its littermate control. AP, VP, DLP denote anterior, ventral, and dorsolateral lobes, respectively.
By immunohistochemistry we demonstrated the expression of both survivin and Runx2 proteins in malignant and benign human prostate epithelium. Survivin expression was found predominantly in the cytoplasm and was qualitatively and significantly higher in the cancerous epithelium (Figure 5b). Additionally, our data implied a possible trend of decreased expression with higher tumor grade (Table 1). A trend of increased expression of Runx2 (nuclear) in the malignant epithelium relative to the benign epithelium was also noted (Figure 5b), although the analysis could not point to any remarkable changes with respect to Gleason grades (Table 1). It would be necessary to confirm these initial immunohistochemical findings by other more quantitative means, such as Western blot analysis, whenever such means are available. Given the limited cohort size statistical analysis was not performed at this time point. Previously, we showed that survivin expression is progressively increased with growth of mouse prostate tumors.13 To determine whether a similar change occurs for Runx2, we examined Runx2 expression in prostatic tumors of the conditional Pten deletion mouse model. Western blot analysis showed that Runx2 levels in anterior prostate (AP) were gradually increased with the tumor growth in the mutant mice ranging in age from 1.6 to 11 months, while the levels remained consistently low in normal tissues of normal littermate controls (Figure 5c), similar to what was observed for BMP7 and survivin.8, 13 Additionally, each prostatic lobe obtained from the 11-month-old, Pten-knockout mouse was examined. In all three lobes, anterior, ventral, and dorsolateral, Runx2 expression was remarkably higher in tumorous tissues as compared with that in the corresponding tissues in the littermate controls, although the expression levels appeared to vary from lobe to lobe (Figure 5d). It is recognized that epithelial cells in the various lobes of the mouse prostate differ in the height of the epithelium, amount of epithelial infolding, and the size of the nuclei.33 The changes between normal and tumor-bearing lobes, however, remain striking. This type of distinctive changes in Runx2 expression, on the other hand, was not apparent from the human prostate cancer specimens. The apparent differences between human and mouse prostate cancer specimens could be due to multiple parameters, such as, variations between species, increased homogeneity of cancer cells in the genetically engineered mouse model over the complex heterogeneity of sporadic human cancer, and the differences in the techniques used, namely, Western blots for mouse tissues and immunohistochemistry for human tissues.
DISCUSSION
The survivin gene promoter could be a cancer-specific promoter as expression of survivin is documented in nearly all human tumors, with minimal or no expression in most of normal adult tissues with the exception of hematopoietic cells.10 The basal transcriptional requirements of survivin gene expression have been defined to include Sp1 sites.20 With ChIP assays we show that Runx2 can interact with at least two regions (region-I and II) in the survivin promoter in C4-2B cells cultured under normal conditions, with the association with region-I being relatively stronger. When the cells are deprived of serum, these regions remain free of bound Runx2. However, upon BMP7 stimulation, Runx2 localization to region-II, but not to region-I, is rescued in serum-starved cells. Thus, there seems to be differential utilization of upstream promoter sequences for survivin transcription in vivo in response to BMP signaling. The elevated survivin expression is unlikely to be related to cell cycle as no significant changes in the number of cells at the G2–M phase were apparent when the cells were treated with BMP7.13 The scenario projects a complex and dynamic Runx2 interaction with the survivin promoter for regulation of this important gene activity in cancer cells.
In analysis of reporter gene activity, a putative role for region-II is indicated as well. BMP7 substantially enhanced the promoter activity of pLuc1430, which retained the full complements of region-II. However, when a truncated construct (pLuc1270) that lacks the stretch of upstream 160 nucleotide sequences, including the Runx-binding sites of region-II, was used, the BMP7 effect was significantly reduced. It should be noted that region-II not only harbors Runx-binding sites, but also carries multiple Smad-binding elements, and, in fact, we described earlier that transcription of the pLuc1430 promoter was Smad-dependent with respect to the BMP effect.13 It was reported that Runx proteins may indeed recruit Smads to specific subnuclear foci that are coupled to active transcription.34 Furthermore, in BMP2-induced osteoblast differentiation, Smad and Runx2 appeared to be structurally coupled in rendering a biological signal.19 Therefore, it is likely that in BMP-induced activation of survivin gene transcription in the malignant epithelial cells, such an interaction between Runx2 and Smad may also be occurring for recruitment of the complex to the appropriate responsive elements. The details of these interactions and their sequence specificity, however, remain to be elucidated.
BMP7 enhancement of Runx2 binding to the survivin promoter may be influenced by multiple other factors. For example, the ability of Runx2 to bind DNA has been reported to be influenced by both phosphorylation31, 35, 36 and acetylation.31 There are factors, such as HDAC4, that may bind to the Runt domain of Runx2 to inhibit its activity.37 Finally, the biological function of Runx2 is dependent on the mechanisms that facilitate nuclear translocation of this and related molecules.
Besides the dynamics of the recruitment of Runx2 to various sites on the survivin promoter in cancer cells under diverse cellular environments, we propose that the molecular basis of overexpression of survivin may also be linked to Runx2 levels in cancer. Runx2 mRNA level is reported to be significantly increased in human pancreatic ductal adenocarcinoma tissues32 and esophageal squamous cell carcinoma.38 Runx2 expression has also been implicated in bone metastasis of breast cancer,39 and in human prostate cancer tissues and cells lines the expression of Runx2 protein has been reported.40 Here we describe elevated expression of Runx2 protein in human prostate cancer cell lines relative to non-tumorigenic cell lines. Most remarkably, in a relatively homogeneous model system of prostate tumor, namely the conditional Pten deletion mice,6, 7 we demonstrate a strong correlation between increased Runx2 levels with growth of the tumor, an observation that is very similar to what we described for survivin protein levels in the same model.13 Compared with this cancer of the mouse model, human prostate cancer is, however, highly heterogeneous, with diversity in the cellular types. Still, consistent with previous observations with a variety of cancers, we confirm that there is an overall and apparently significant elevation in the level of survivin protein expression in the human prostate cancer specimens relative to either benign epithelium or the surrounding stroma. Runx2 expression is also readily detected in these cancer tissues. Both malignant and benign epithelia appear to express Runx2 significantly more than the stroma, and a tendency of higher expression in malignant relative to benign tissues is noted. Perhaps, to reach or maintain a threshold level is what is needed to amplify Runx2 activity in the cancer cells. It would be important now to critically examine intracellular Runx2 protein modifications, stability, and parameters of nuclear translocation to correlate with the potential functional attributes of this important transcription factor in neoplastic cells.
Finally, we show that in serum-starved C4-2B cancer cells, BMP7 treatment enhances Runx2 level, an effect that is likely to increase the responsiveness of the cells to BMP signaling to diverse biological functions, including gain in the potential for survival. Since there is detectable early transcriptional upregulation of Runx2, as well as late increases in the level of this protein, the observed BMP7 effect may be both direct and indirect. For instance, BMP2 is described to inhibit Smurf-mediated ubiquitination and degradation of Runx2 by stimulating p300-mediated Runx2 acetylation.31 It is possible that, in addition to its action on Runx2 transcription, BMP7 may, perhaps, influence Runx2 level in a similar way. Taken together, our results indicate that among various factors that determine cancer-selective activation of the survivin promoter, Runx2 is an important molecular entity and that the Runx2 threshold or activation may be linked to tumor cell survival and cancer progression.
References
Celeste AJ, Iannazzi JA, Taylor RC, et al. Identification of transforming growth factor beta family members present in bone-inductive protein purified from bovine bone. Proc Natl Acad Sci USA 1990;87:9843–9847.
Ducy P, Karsenty G . The family of bone morphogenetic proteins. Kidney Int 2000;57:2207–2214.
Bobinac D, Maric I, Zoricic S, et al. Expression of bone morphogenetic proteins in human metastatic prostate and breast cancer. Croat Med J 2005;46:389–396.
Masuda H, Fukabori Y, Nakano K, et al. Increased expression of bone morphogenetic protein-7 in bone metastatic prostate cancer. Prostate 2003;54:268–274.
Masuda H, Fukabori Y, Nakano K, et al. Expression of bone morphogenetic protein-7 (BMP-7) in human prostate. Prostate 2004;59:101–106.
Wang S, Gao J, Lei Q, et al. Prostate-specific deletion of the murine Pten tumor suppressor gene leads to metastatic prostate cancer. Cancer Cell 2003;4:209–221.
Liao CP, Zhong C, Saribekyan G, et al. Mouse models of prostate adenocarcinoma with the capacity to monitor spontaneous carcinogenesis by bioluminescence or fluorescence. Cancer Res 2007;67:7525–7533.
Yang S, Zhong C, Frenkel B, et al. Diverse biological effect and Smad signaling of bone morphogenetic protein 7 in prostate tumor cells. Cancer Res 2005;65:5769–5777.
Yang S, Pham LK, Liao CP, et al. A novel bone morphogenetic protein signaling in heterotypic cell interactions in prostate cancer. Cancer Res 2008;68:198–205.
Altieri DC . Survivin, cancer networks and pathway-directed drug discovery. Nat Rev Cancer 2008;8:61–70.
Aoki Y, Feldman GM, Tosato G . Inhibition of STAT3 signaling induces apoptosis and decreases survivin expression in primary effusion lymphoma. Blood 2003;101:1535–1542.
Heinemeyer T, Wingender E, Reuter I, et al. Databases on transcriptional regulation: TRANSFAC, TRRD and COMPEL. Nucleic Acids Res 1998;26:362–367.
Yang S, Lim M, Pham LK, et al. Bone morphogenetic protein 7 protects prostate cancer cells from stress-induced apoptosis via both Smad and c-Jun NH2-terminal kinase pathways. Cancer Res 2006;66:4285–4290.
Meyers S, Downing JR, Hiebert SW . Identification of AML-1 and the (8;21) translocation protein (AML-1/ETO) as sequence-specific DNA-binding proteins: the runt homology domain is required for DNA binding and protein–protein interactions. Mol Cell Biol 1993;13:6336–6345.
Fowler M, Borazanci E, McGhee L, et al. RUNX1 (AML-1) and RUNX2 (AML-3) cooperate with prostate-derived Ets factor to activate transcription from the PSA upstream regulatory region. J Cell Biochem 2006;97:1–17.
Blyth K, Cameron ER, Neil JC . The RUNX genes: gain or loss of function in cancer. Nat Rev Cancer 2005;5:376–387.
Hanai J, Chen LF, Kanno T, et al. Interaction and functional cooperation of PEBP2/CBF with Smads. Synergistic induction of the immunoglobulin germline Calpha promoter. J Biol Chem 1999;274:31577–31582.
Javed A, Afzal F, Bae JS, et al. Specific residues of RUNX2 are obligatory for formation of BMP2-induced RUNX2–SMAD complex to promote osteoblast differentiation. Cells Tissues Organs 2009;189:133–137.
Javed A, Bae JS, Afzal F, et al. Structural coupling of Smad and Runx2 for execution of the BMP2 osteogenic signal. J Biol Chem 2008;283:8412–8422.
Li F, Altieri DC . Transcriptional analysis of human survivin gene expression. Biochem J 1999;344:305–311.
Meyers S, Lenny N, Hiebert SW . The t(8;21) fusion protein interferes with AML-1B-dependent transcriptional activation. Mol Cell Biol 1995;15:1974–1982.
Pratap J, Javed A, Languino LR, et al. The Runx2 osteogenic transcription factor regulates matrix metalloproteinase 9 in bone metastatic cancer cells and controls cell invasion. Mol Cell Biol 2005;25:8581–8591.
Roca H, Franceschi RT . Analysis of transcription factor interactions in osteoblasts using competitive chromatin immunoprecipitation. Nucleic Acids Res 2008;36:1723–1730.
Yan M, Kanbe E, Peterson LF, et al. A previously unidentified alternatively spliced isoform of t(8;21) transcript promotes leukemogenesis. Nat Med 2006;12:945–949.
Wildonger J, Mann RS . The t(8;21) translocation converts AML1 into a constitutive transcriptional repressor. Development 2005;132:2263–2272.
Ning YM, Robins DM . AML3/CBFalpha1 is required for androgen-specific activation of the enhancer of the mouse sex-limited protein (Slp) gene. J Biol Chem 1999;274:30624–30630.
Kent WJ . BLAT—the BLAST-like alignment tool. Genome Res 2002;12:656–664.
Lee KS, Kim HJ, Li QL, et al. Runx2 is a common target of transforming growth factor beta1 and bone morphogenetic protein 2, and cooperation between Runx2 and Smad5 induces osteoblast-specific gene expression in the pluripotent mesenchymal precursor cell line C2C12. Mol Cell Biol 2000;20:8783–8792.
Leboy P, Grasso-Knight G, D’Angelo M, et al. Smad–Runx interactions during chondrocyte maturation. J Bone Joint Surg Am 2001;83-A Suppl 1(Pt 1):S15–S22.
Tou L, Quibria N, Alexander JM . Transcriptional regulation of the human Runx2/Cbfa1 gene promoter by bone morphogenetic protein-7. Mol Cell Endocrinol 2003;205:121–129.
Jeon EJ, Lee KY, Choi NS, et al. Bone morphogenetic protein-2 stimulates Runx2 acetylation. J Biol Chem 2006;281:16502–16511.
Kayed H, Jiang X, Keleg S, et al. Regulation and functional role of the Runt-related transcription factor-2 in pancreatic cancer. Br J Cancer 2007;97:1106–1115.
Roy-Burman P, Wu H, Powell WC, et al. Genetically defined mouse models that mimic natural aspects of human prostate cancer development. Endocr Relat Cancer 2004;11:225–254.
Zaidi SK, Sullivan AJ, van Wijnen AJ, et al. Integration of Runx and Smad regulatory signals at transcriptionally active subnuclear sites. Proc Natl Acad Sci USA 2002;99:8048–8053.
Selvamurugan N, Shimizu E, Lee M, et al. Identification and characterization of Runx2 phosphorylation sites involved in matrix metalloproteinase-13 promoter activation. FEBS Lett 2009;583:1141–1146.
Wee HJ, Huang G, Shigesada K, et al. Serine phosphorylation of RUNX2 with novel potential functions as negative regulatory mechanisms. EMBO Rep 2002;3:967–974.
Vega RB, Matsuda K, Oh J, et al. Histone deacetylase 4 controls chondrocyte hypertrophy during skeletogenesis. Cell 2004;119:555–566.
Tonomoto Y, Tachibana M, Dhar DK, et al. Differential expression of RUNX genes in human esophageal squamous cell carcinoma: downregulation of RUNX3 worsens patient prognosis. Oncology 2007;73:346–356.
Barnes GL, Hebert KE, Kamal M, et al. Fidelity of Runx2 activity in breast cancer cells is required for the generation of metastases-associated osteolytic disease. Cancer Res 2004;64:4506–4513.
Brubaker KD, Vessella RL, Brown LG, et al. Prostate cancer expression of runt-domain transcription factor Runx2, a key regulator of osteoblast differentiation and function. Prostate 2003;56:13–22.
Acknowledgements
This study was supported by NIH grants RO1 CA113392 and NIH RO1 CA59705 (to P Roy-Burman). We thank Dr Dario C Altieri, Dr A Hari Reddi, and Dr Baruch Frankel for the gift of various reagents used in this work, and Dr Young-Ho Lee and all the members of the Roy-Burman laboratory for their support in various aspects of this study.
Author information
Authors and Affiliations
Corresponding author
Additional information
DISCLOSURE/Conflict of interest
The authors declare no conflict of interest.
Rights and permissions
About this article
Cite this article
Lim, M., Zhong, C., Yang, S. et al. Runx2 regulates survivin expression in prostate cancer cells. Lab Invest 90, 222–233 (2010). https://doi.org/10.1038/labinvest.2009.128
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/labinvest.2009.128
Keywords
This article is cited by
-
HDACs control RUNX2 expression in cancer cells through redundant and cell context-dependent mechanisms
Journal of Experimental & Clinical Cancer Research (2019)
-
Impact of RUNX2 on drug-resistant human pancreatic cancer cells with p53 mutations
BMC Cancer (2018)
-
Runx2, an inducer of osteoblast and chondrocyte differentiation
Histochemistry and Cell Biology (2018)
-
MicroRNA-466 inhibits tumor growth and bone metastasis in prostate cancer by direct regulation of osteogenic transcription factor RUNX2
Cell Death & Disease (2017)
-
Role of Runx2 phosphorylation in prostate cancer and association with metastatic disease
Oncogene (2016)