Featured
-
-
Article
| Open Access Temporal and spatial dynamics of scaling-specific features of a gene regulatory network in Drosophila
Temporal and spatial dynamics of scaling-specific features of a gene regulatory network in Drosophila
How pattern formation is regulated relative to the size of an organism is unclear. Here, Wu et al.take data from gap gene expression in flies of different sizes together with simulations, identifying how scaling emerges dynamically and that local patterning influences global gene regulatory networks.
- Honggang Wu
- , Manu
- & Jun Ma
-
Article
| Open Access miRNA–target chimeras reveal miRNA 3′-end pairing as a major determinant of Argonaute target specificity
miRNA–target chimeras reveal miRNA 3′-end pairing as a major determinant of Argonaute target specificity
microRNAs (miRNAs) act as sequence-specific guides for Argonaute (AGO) proteins. By using a modified AGO HITS-CLIP strategy that enriches miRNAs ligated to their endogenous mRNA targets, here the authors show that miRNA 3' end pairing is a general determinant of AGO binding specificity.
- Michael J. Moore
- , Troels K. H. Scheel
- & Robert B. Darnell
-
Article
| Open Access Characterizing noise structure in single-cell RNA-seq distinguishes genuine from technical stochastic allelic expression
Characterizing noise structure in single-cell RNA-seq distinguishes genuine from technical stochastic allelic expression
Single-cell RNA-sequencing (scRNA-seq) can be applied to dissect the kinetics of gene expression and patterns of allele-specific expression. Here, Kim et al.report a generative statistical model that can separate biological variability from technical noise by quantifying technical noise using external RNA spike-ins.
- Jong Kyoung Kim
- , Aleksandra A. Kolodziejczyk
- & John C. Marioni
-
Article
| Open Access ChEC-seq kinetics discriminates transcription factor binding sites by DNA sequence and shape in vivo
ChEC-seq kinetics discriminates transcription factor binding sites by DNA sequence and shape in vivo
In chromatin endogenous cleavage (ChEC), micrococcal nuclease (MNase) is fused to a protein of interest and its cleavage is thus targeted to specific genomic loci in vivo. Here, the authors show that time-resolved ChEC-seq (high-throughput sequencing after ChEC) can detect DNA shape patterns regardless of motif strength.
- Gabriel E. Zentner
- , Sivakanthan Kasinathan
- & Steven Henikoff
-
Article
| Open Access Bayesian integration of genetics and epigenetics detects causal regulatory SNPs underlying expression variability
Bayesian integration of genetics and epigenetics detects causal regulatory SNPs underlying expression variability
Das et al. present a novel Bayesian approach called expression Quantitative Trait enhancer Loci (eQTeL), which effectively integrates genetic and epigenetic information to identify combination of regulatory genomic variants underlying expression variance. Using various functional data, the authors show the variants identified by eQTeL are likely to be causal.
- Avinash Das
- , Michael Morley
- & Sridhar Hannenhalli
-
Article
| Open Access Systematic analysis of somatic mutations impacting gene expression in 12 tumour types
Systematic analysis of somatic mutations impacting gene expression in 12 tumour types
Assessing functional impact of mutations in cancer on gene expression can improve our understanding of cancer biology and may identify potential therapeutic targets. Here, Ding et al. describe a novel statistical model named xseq for a systematic survey of how mutations impact transcriptome landscapes across 12 different tumour types.
- Jiarui Ding
- , Melissa K. McConechy
- & Sohrab P. Shah
-
Article
| Open Access Lower glycolysis carries a higher flux than any biochemically possible alternative
Lower glycolysis carries a higher flux than any biochemically possible alternative
The biochemical pathways of central carbon metabolism are highly conserved across all domains of life. Here, Courtet al. use a computational approach to test all possible pathways of glycolysis and gluconeogenesis and find that the existing trunk pathways may represent a maximal flux solution selected for during evolution.
- Steven J. Court
- , Bartlomiej Waclaw
- & Rosalind J. Allen
-
Article
| Open Access Global priorities for an effective information basis of biodiversity distributions
Global priorities for an effective information basis of biodiversity distributions
Comprehensive digital information on species distributions is crucial for research in ecology, evolution and conservation. Here, Meyer et al.find large gaps and biases in global vertebrate point records, especially in emerging economies, and identify key factors currently limiting information.
- Carsten Meyer
- , Holger Kreft
- & Walter Jetz
-
Article
| Open Access Systematic chromatin state comparison of epigenomes associated with diverse properties including sex and tissue type
Systematic chromatin state comparison of epigenomes associated with diverse properties including sex and tissue type
In contrast to genetic information, epigenetic state varies greatly under different conditions. Here, the authors develop ChromDiff and apply it to ENCODE and Roadmap Epigenomics datasets to find chromatin state differences associated with sex, tissue, and developmental age.
- Angela Yen
- & Manolis Kellis
-
Article
| Open Access A draft network of ligand–receptor-mediated multicellular signalling in human
A draft network of ligand–receptor-mediated multicellular signalling in human
Cell-to-cell communication relies upon interactions between secreted ligands and cell surface receptors. Here, Ramilowski et al.present a draft cell-to-cell communication network based on expression of ligand-receptor pairs in 144 different human cell types.
- Jordan A. Ramilowski
- , Tatyana Goldberg
- & Alistair R. R. Forrest
-
Article
| Open Access Context influences on TALE–DNA binding revealed by quantitative profiling
Context influences on TALE–DNA binding revealed by quantitative profiling
TALE proteins are popular tools for genome engineering because they can recognize specific DNA sequences, however off-target effects are a routine problem. Here Rogers and Barrera et al. comprehensively map TALE–DNA interactions to develop a computational model to predict binding specificity.
- Julia M. Rogers
- , Luis A. Barrera
- & Martha L. Bulyk
-
Article
| Open Access Pharmacogenomic and clinical data link non-pharmacokinetic metabolic dysregulation to drug side effect pathogenesis
Pharmacogenomic and clinical data link non-pharmacokinetic metabolic dysregulation to drug side effect pathogenesis
Adverse drug reactions are an important clinical problem. Here the authors combine information about drug-induced gene expression changes and genetic variability of patients with a genome-scale metabolic model to identify drug-induced changes in cellular metabolism that may be linked to drug side effects.
- Daniel C. Zielinski
- , Fabian V. Filipp
- & Bernhard O. Palsson
-
Article
| Open Access Extreme multifunctional proteins identified from a human protein interaction network
Extreme multifunctional proteins identified from a human protein interaction network
Proteins are sometimes implicated in separate and seemingly unrelated processes, so called moonlighting functions. Here the authors use bioinformatics tools to identify extreme multifunctional proteins and define a signature of extreme multifunctionality.
- Charles E. Chapple
- , Benoit Robisson
- & Christine Brun
-
Article |
 Evolutionary-guided de novo structure prediction of self-associated transmembrane helical proteins with near-atomic accuracy
Evolutionary-guided de novo structure prediction of self-associated transmembrane helical proteins with near-atomic accuracy
While the transmembrane regions of single-pass transmembrane proteins play critical roles in receptor signalling, they remain difficult to characterize structurally. Here the authors present a computational approach for accurate structure prediction of associated single-pass transmembrane helical proteins.
- Y. Wang
- & P. Barth
-
Article
| Open Access Planctomycetes do possess a peptidoglycan cell wall
Planctomycetes do possess a peptidoglycan cell wall
Planctomycetes appear to differ from all other bacteria in their cellular organization and their apparent lack of a peptidoglycan (PG) cell wall. Here Jeske et al. show that Planctomycetes do possess a typical PG cell wall and that their cellular architecture resembles that of Gram-negative bacteria.
- Olga Jeske
- , Margarete Schüler
- & Christian Jogler
-
Article |
 A network approach for identifying and delimiting biogeographical regions
A network approach for identifying and delimiting biogeographical regions
There is currently no consensus on how best to identify and delimit biogeographical regions. Here the authors develop a network-based approach incorporating complex presence–absence patterns that can successfully identify commonly recognized biogeographical regions, and apply it to two large-scale data sets of plants and amphibians.
- Daril A. Vilhena
- & Alexandre Antonelli
-
Article
| Open Access Combinatorial code governing cellular responses to complex stimuli
Combinatorial code governing cellular responses to complex stimuli
Cells constantly integrate information from multiple stimuli. By considering every possible means by which two stimuli can interact, Cappuccio et al. define 10 interaction modes and demonstrate their preferential use by dendritic cells responding to different combinations of microbial and host inflammatory cues.
- Antonio Cappuccio
- , Raphaël Zollinger
- & Vassili Soumelis
-
Article |
 Calibrating genomic and allelic coverage bias in single-cell sequencing
Calibrating genomic and allelic coverage bias in single-cell sequencing
Artifacts caused by whole-genome amplification bias are a recurrent challenge in single-cell sequencing analysis. Here, the authors develop statistical models and demonstrate an efficient strategy for controlling amplification errors by a joint analysis of single cell genomes.
- Cheng-Zhong Zhang
- , Viktor A. Adalsteinsson
- & J. Christopher Love
-
Article |
 Changing cell behaviours during beetle embryogenesis correlates with slowing of segmentation
Changing cell behaviours during beetle embryogenesis correlates with slowing of segmentation
Sequential segmentation in development is best described in vertebrates, where it relies on cell proliferation and shows regular periodicity. Here, the authors show that in the flour beetle segments are added with irregular rate and their elongation during periods of fast growth relies mostly on cell movements.
- A. Nakamoto
- , S. D. Hester
- & T. A. Williams
-
Article
| Open Access Decoding the regulatory landscape of melanoma reveals TEADS as regulators of the invasive cell state
Decoding the regulatory landscape of melanoma reveals TEADS as regulators of the invasive cell state
The key regulators that allow transition from proliferative to invasive phenotype in melanoma cells have not been identified yet. The authors perform chromatin and transcriptome profiling followed by comprehensive bioinformatics analysis identifying new candidate regulators for two distinct cell states of melanoma.
- Annelien Verfaillie
- , Hana Imrichova
- & Stein Aerts
-
Article
| Open Access Ptch1 and Gli regulate Shh signalling dynamics via multiple mechanisms
Ptch1 and Gli regulate Shh signalling dynamics via multiple mechanisms
Gradients of the secreted morphogen Sonic Hedgehog (Shh) pattern the neural tube in vertebrates. Cohen et al.quantify Shh signalling in developing mice, and by constructing a computational model of the process, identify mechanisms by which the dynamics of Shh signalling are regulated.
- Michael Cohen
- , Anna Kicheva
- & James Briscoe
-
Article |
 Notch1–Dll4 signalling and mechanical force regulate leader cell formation during collective cell migration
Notch1–Dll4 signalling and mechanical force regulate leader cell formation during collective cell migration
Many forms of collective cell migration are directed by specialized leader cells that have a distinct protrusive phenotype. Riahi et al.show that lateral inhibition mediated by Notch1–Dll4 signalling determines the stochastic emergence of leader cells in epithelial monolayers.
- Reza Riahi
- , Jian Sun
- & Pak Kin Wong
-
Article |
 Hippos stem from the longest sequence of terrestrial cetartiodactyl evolution in Africa
Hippos stem from the longest sequence of terrestrial cetartiodactyl evolution in Africa
The evolutionary origin of Hippopotamidae, the family of hippos, is poorly understood. Here, the authors describe a new fossil from Kenya that unambiguously roots Hippopotamidae into the group that includes the first large terrestrial mammals to invade Africa, more than 30 million years ago.
- Fabrice Lihoreau
- , Jean-Renaud Boisserie
- & Stéphane Ducrocq
-
Article
| Open Access Sensory integration dynamics in a hierarchical network explains choice probabilities in cortical area MT
Sensory integration dynamics in a hierarchical network explains choice probabilities in cortical area MT
The activity of sensory neurons can be correlated with perceptual decisions and this effect may provide insights into how sensory information is processed during perceptual tasks. Here the authors develop a network model of sensory and decision-making areas and propose that the dynamics across the network hierarchy explains the choice probabilities.
- Klaus Wimmer
- , Albert Compte
- & Jaime de la Rocha
-
Article
| Open Access Prospective errors determine motor learning
Prospective errors determine motor learning
Motor learning is characterized by diverse cognitive processes, which lack a unified theoretical framework. Here, Takiyama et al.present a model demonstrating that motor learning is determined by prospective errors, which they test in a specially designed visuomotor adaptation task.
- Ken Takiyama
- , Masaya Hirashima
- & Daichi Nozaki
-
Article
| Open Access The ligand binding mechanism to purine nucleoside phosphorylase elucidated via molecular dynamics and machine learning
The ligand binding mechanism to purine nucleoside phosphorylase elucidated via molecular dynamics and machine learning
Understanding the dynamics of enzyme-substrate complexation provides an insight into potential drugs, but intermediate states are difficult to observe experimentally. Here, the authors use simulations and machine learning to analyse the binding of transition state inhibitors to purine nucleoside phosphorylase.
- Sergio Decherchi
- , Anna Berteotti
- & Andrea Cavalli
-
Article
| Open Access Human-to-mosquito transmission efficiency increases as malaria is controlled
Human-to-mosquito transmission efficiency increases as malaria is controlled
Understanding the epidemiology of malaria transmission between humans and mosquitoes is crucial for successful disease control. Analysing data from an 18-year malaria control programme, Churcher et al. show that decreased parasite prevalence in humans can be found concurrently with an increase in transmission efficiency.
- Thomas S. Churcher
- , Jean-François Trape
- & Anna Cohuet
-
Article |
 Biological interpretation of genome-wide association studies using predicted gene functions
Biological interpretation of genome-wide association studies using predicted gene functions
Identifying which genes and pathways explain genetic associations is challenging. Here, the authors present DEPICT, a tool for gene prioritization, pathway analysis and tissue/cell-type enrichment analysis that can be used to generate testable hypotheses from genetic association studies.
- Tune H. Pers
- , Juha M. Karjalainen
- & Lude Franke
-
Article |
 The statistical geometry of transcriptome divergence in cell-type evolution and cancer
The statistical geometry of transcriptome divergence in cell-type evolution and cancer
Body plan complexity is associated with the number of different cell types, yet the processes that create this diversity are unclear. Here the authors use transcriptomics to test the hypothesis that unlike cancer cells, novel normal cell types arise through sub-specialization of an ancestral cell type.
- Cong Liang
- , Alistair R.R. Forrest
- & Günter P. Wagner
-
Article |
 Amino acid coevolution reveals three-dimensional structure and functional domains of insect odorant receptors
Amino acid coevolution reveals three-dimensional structure and functional domains of insect odorant receptors
The structure of insect odorant receptors (ORs) has remained elusive due to their lack of homology to other proteins and the inability to obtain OR crystals. Here, the authors use amino acid evolutionary covariation patterns to fold these proteins de novoand generate the first three-dimensional models of insect ORs.
- Thomas A. Hopf
- , Satoshi Morinaga
- & Richard Benton
-
Article
| Open Access Visualizing cellular imaging data using PhenoPlot
Visualizing cellular imaging data using PhenoPlot
Cellular imaging studies can generate large volumes of complex phenotypic data; however, presenting this information in a form that quickly conveys trends in the data set remains a challenge. Sailem et al.present a tool which translates such data into easily interpretable cell-like glyphs.
- Heba Z. Sailem
- , Julia E. Sero
- & Chris Bakal
-
Article
| Open Access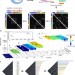 High-quality genome (re)assembly using chromosomal contact data
High-quality genome (re)assembly using chromosomal contact data
The correct assembly of genomes from sequencing data remains a challenge due to difficulties in correctly assigning the location of repeated DNA elements. Here the authors describe GRAAL, an algorithm that utilizes genome-wide chromosome contact data within a probabilistic framework to produce accurate genome assemblies.
- Hervé Marie-Nelly
- , Martial Marbouty
- & Romain Koszul
-
Article |
 microTSS: accurate microRNA transcription start site identification reveals a significant number of divergent pri-miRNAs
microTSS: accurate microRNA transcription start site identification reveals a significant number of divergent pri-miRNAs
microRNAs are short non-coding RNAs that post-transcriptionally regulate gene expression for which the identification of promoter and primary transcripts (pri-miRNAs) has been difficult. Here the authors describe microTSS, an algorithm that supports the precise identification of intergenic pri-miRNA transcription start sites.
- Georgios Georgakilas
- , Ioannis S. Vlachos
- & Artemis G. Hatzigeorgiou
-
Article |
 Dynamic analyses of alternative polyadenylation from RNA-seq reveal a 3′-UTR landscape across seven tumour types
Dynamic analyses of alternative polyadenylation from RNA-seq reveal a 3′-UTR landscape across seven tumour types
Alternative polyadenylation (APA) has been implicated in diverse physiological and pathological conditions including cancer. The authors present a new algorithm, DaPars, for APA analysis using available RNA-seq data and suggest CstF64 as a master regulator of 3′-UTR shortening across multiple tumour types.
- Zheng Xia
- , Lawrence A. Donehower
- & Wei Li
-
Article |
 Prediction and quantification of bioactive microbiota metabolites in the mouse gut
Prediction and quantification of bioactive microbiota metabolites in the mouse gut
Metabolites produced by the gut microbiota can potentially affect our physiology. Here, the authors present a metabolomics strategy that models microbiota metabolism as a reaction network and uses pathway analysis to facilitate identification and characterization of microbial metabolites.
- Gautham V. Sridharan
- , Kyungoh Choi
- & Arul Jayaraman
-
Article |
 Quantitative profiling of peptides from RNAs classified as noncoding
Quantitative profiling of peptides from RNAs classified as noncoding
A large portion of the transcribed genome—such as introns and noncoding RNAs—is believed to not be translated into protein products. Here, the authors provide evidence for the existence of regulated peptide products that are translated from transcribed sequences generally characterized as noncoding.
- Sudhakaran Prabakaran
- , Martin Hemberg
- & Judith A. Steen
-
Article
| Open Access CpG island-mediated global gene regulatory modes in mouse embryonic stem cells
CpG island-mediated global gene regulatory modes in mouse embryonic stem cells
CpG islands are high GC content DNA elements that surround the majority of transcriptional start sites in eukaryotes. Here, the authors analyse over 200 genomic data sets to provide new insight into global CpG islands-dependent regulatory mechanisms in differentiated and pluripotent stem cells.
- Samuel Beck
- , Bum-Kyu Lee
- & Jonghwan Kim
-
Article |
 Nuclear stability and transcriptional directionality separate functionally distinct RNA species
Nuclear stability and transcriptional directionality separate functionally distinct RNA species
Despite our growing understanding of their complexity, different types of RNA are still classified using technical rather than functional criteria. Andersson et al.show that categorization of RNAs based on stability and direction of transcription is an effective means of functional classification.
- Robin Andersson
- , Peter Refsing Andersen
- & Albin Sandelin
-
Article |
 MS-GF+ makes progress towards a universal database search tool for proteomics
MS-GF+ makes progress towards a universal database search tool for proteomics
The development of software tools to analyse large mass spectrometry data sets lags behind the increase in diversity of the data. Here the authors develop MS-GF+, a database search tool that outperforms other popular tools in identifying peptides from a variety of data sets.
- Sangtae Kim
- & Pavel A. Pevzner
-
Article
| Open Access On the absence of intrahelical DNA dynamics on the μs to ms timescale
On the absence of intrahelical DNA dynamics on the μs to ms timescale
No experimental evidence exists for intra-helical motion of DNA at the μs timescale, which has been attributed to technical difficulties in observing motion in this time range. Here, the authors demonstrate, using extensive molecular dynamics simulations and experimental analysis, that such motion is effectively absent from a B-DNA duplex.
- Rodrigo Galindo-Murillo
- , Daniel R. Roe
- & Thomas E. Cheatham III
-
Article |
 Protein design with a comprehensive statistical energy function and boosted by experimental selection for foldability
Protein design with a comprehensive statistical energy function and boosted by experimental selection for foldability
Methods to design proteins de novo can give insights into how amino acids fold into particular structures and aid in protein engineering. Here, Xiong et al. compare a novel statistical energy function with established methods and use it to generate four de novoproteins.
- Peng Xiong
- , Meng Wang
- & Haiyan Liu
-
Article
| Open Access Concomitant Notch activation and p53 deletion trigger epithelial-to-mesenchymal transition and metastasis in mouse gut
Concomitant Notch activation and p53 deletion trigger epithelial-to-mesenchymal transition and metastasis in mouse gut
Metastasizing tumour cells undergo epithelial-to-mesenchymal transition. Using both bioinformatic and in vivo approaches, Chanrion et al.identify combined Notch activation and p53 inactivation as a potent inducer of this transition, and apply this to create a highly metastatic tumour model in mice.
- Maia Chanrion
- , Inna Kuperstein
- & Sylvie Robine
-
Article
| Open Access An exact arithmetic toolbox for a consistent and reproducible structural analysis of metabolic network models
An exact arithmetic toolbox for a consistent and reproducible structural analysis of metabolic network models
Current tools to analyse constraint-based models of metabolic networks have limited accuracy due to their use of floating-point arithmetic. Here the authors present MONGOOSE, a new computational tool that analyses such models in exact arithmetic, providing improved accuracy and reproducibility.
- Leonid Chindelevitch
- , Jason Trigg
- & Bonnie Berger
-
Article |
 Assessing technical performance in differential gene expression experiments with external spike-in RNA control ratio mixtures
Assessing technical performance in differential gene expression experiments with external spike-in RNA control ratio mixtures
Differential gene expression experiments yield quantitative insight into biological activity and may be important in disease classification and treatment. Here, the authors analyse external spike-in RNA controls to provide a standard method to assess and compare experiment performance.
- Sarah A. Munro
- , Steven P. Lund
- & Marc Salit
-
Article
| Open Access Warped linear mixed models for the genetic analysis of transformed phenotypes
Warped linear mixed models for the genetic analysis of transformed phenotypes
Linear mixed models (LMMs) provide a powerful method for studying genotype–phenotype associations. Here the authors present a LMM application that estimates an optimal transformation from observed data and increases the accuracy of heritability estimation and phenotype prediction.
- Nicolo Fusi
- , Christoph Lippert
- & Oliver Stegle
-
Article |
 Improved nucleosome-positioning algorithm iNPS for accurate nucleosome positioning from sequencing data
Improved nucleosome-positioning algorithm iNPS for accurate nucleosome positioning from sequencing data
Changes in nucleosome positioning often underlie the reprogramming of gene expression during differentiation. Here Chen et al. describe a novel algorithm - iNPS - that outperforms current methods in accurately determine nucleosome position genome-wide.
- Weizhong Chen
- , Yi Liu
- & Jing-Dong Jackie Han
-
Article |
 Ribosomal DNA copy number is coupled with gene expression variation and mitochondrial abundance in humans
Ribosomal DNA copy number is coupled with gene expression variation and mitochondrial abundance in humans
The functional consequences of naturally occurring variation in ribosomal DNA (rDNA) copy number are poorly understood. Here the authors estimate rDNA copy number and mitochondrial DNA abundance in humans using whole-genome short-read DNA sequencing and characterize global regulatory mechanisms for cellular homeostasis and adaptation.
- John G. Gibbons
- , Alan T. Branco
- & Bernardo Lemos
-
Article
| Open Access Genome dynamics of the human embryonic kidney 293 lineage in response to cell biology manipulations
Genome dynamics of the human embryonic kidney 293 lineage in response to cell biology manipulations
The human embryonic kidney 293 (HEK293) cell lineage is widely used in cell biology and biotechnology. Here, the authors apply whole genome resequencing methods to characterise genomic variation in six HEK293 cell lines and suggest that this variation could affect experiments using these cell lines.
- Yao-Cheng Lin
- , Morgane Boone
- & Nico Callewaert
-
Article
| Open Access A genome-wide map of hyper-edited RNA reveals numerous new sites
A genome-wide map of hyper-edited RNA reveals numerous new sites
Common methods to detect adenosine-to-inosine RNA editing sites rely on mapping short RNA reads to the genome while allowing only a limited number of mismatches. Here, Porath et al. present a novel RNA-seq based approach to identify hyper-edited reads that significantly expands the RNA editome.
- Hagit T. Porath
- , Shai Carmi
- & Erez Y. Levanon
Browse broader subjects
Browse narrower subjects
- Biochemical reaction networks
- Cellular signalling networks
- Classification and taxonomy
- Communication and replication
- Computational models
- Computational neuroscience
- Computational platforms and environments
- Data acquisition
- Data integration
- Data mining
- Data processing
- Data publication and archiving
- Databases
- Functional clustering
- Gene ontology
- Gene regulatory networks
- Genome informatics
- Hardware and infrastructure
- High-throughput screening
- Image processing
- Literature mining
- Machine learning
- Microarrays
- Network topology
- Phylogeny
- Power law
- Predictive medicine
- Probabilistic data networks
- Programming language
- Protein analysis
- Protein design
- Protein folding
- Protein function predictions
- Protein structure predictions
- Proteome informatics
- Quality control
- Scale invariance
- Sequence annotation
- Software
- Standards
- Statistical methods
- Virtual drug screening

 A comprehensive assessment of somatic mutation detection in cancer using whole-genome sequencing
A comprehensive assessment of somatic mutation detection in cancer using whole-genome sequencing