Article
|
Open Access
Featured
-
-
Article
| Open Access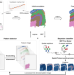 Pianno: a probabilistic framework automating semantic annotation for spatial transcriptomics
Pianno: a probabilistic framework automating semantic annotation for spatial transcriptomics
Recognising spatial spots’ biological identity in spatial transcriptomics remains a challenge. Here, authors introduce Pianno, a tool that helps annotate the biological structures or cell-type constructions across diverse tissues, offering new perspectives on understanding spatial transcriptomics.
- Yuqiu Zhou
- , Wei He
- & Ying Zhu
-
Article
| Open Access Developmental progression of DNA double-strand break repair deciphered by a single-allele resolution mutation classifier
Developmental progression of DNA double-strand break repair deciphered by a single-allele resolution mutation classifier
DNA double-strand breaks (DSBs) are repaired by a hierarchically regulated network of pathways. Here, authors develop ICP for deciphering somatic DSB repair patterns in multicellular organisms and discover developmental regulation in flies and mosquitoes, enabling tracking of mutant alleles and interhomolog copying of gene cassettes.
- Zhiqian Li
- , Lang You
- & Ethan Bier
-
Article
| Open Access AlphaPept: a modern and open framework for MS-based proteomics
AlphaPept: a modern and open framework for MS-based proteomics
Mass spectrometry-based proteomics faces the challenge of processing vast data amounts. Here, the authors introduce AlphaPept, an open-source, Python-based framework that offers high speed analysis and easy integration for large-scale proteome analysis.
- Maximilian T. Strauss
- , Isabell Bludau
- & Matthias Mann
-
Article
| Open Access Statistical method scDEED for detecting dubious 2D single-cell embeddings and optimizing t-SNE and UMAP hyperparameters
Statistical method scDEED for detecting dubious 2D single-cell embeddings and optimizing t-SNE and UMAP hyperparameters
2D visualisation of single-cell data is highly impacted by the hyperparameter setting of the 2D embedding method, such as t-SNE and UMAP. Here, authors develop a statistical method scDEED to detect dubious cell embeddings and optimise the hyperparameter setting for trustworthy visualisation.
- Lucy Xia
- , Christy Lee
- & Jingyi Jessica Li
-
Article
| Open Access scCASE: accurate and interpretable enhancement for single-cell chromatin accessibility sequencing data
scCASE: accurate and interpretable enhancement for single-cell chromatin accessibility sequencing data
Single-cell chromatin accessibility sequencing (scCAS) data suffers from high sparsity and dimensionality. Here, authors propose an accurate and interpretable computational framework for enhancing scCAS data that considers cell-to-cell similarity.
- Songming Tang
- , Xuejian Cui
- & Shengquan Chen
-
Article
| Open Access Phage-plasmids promote recombination and emergence of phages and plasmids
Phage-plasmids promote recombination and emergence of phages and plasmids
Phage-plasmids are mobile genetic elements that transfer horizontally between bacterial cells as viruses, and vertically within bacterial lineages as plasmids. Here, Pfeifer & Rocha show that phage-plasmids can mediate gene transfer across mobile elements within their hosts, and can act as intermediates in the conversion of one type of element into another.
- Eugen Pfeifer
- & Eduardo P. C. Rocha
-
Comment
| Open Access Fudging the volcano-plot without dredging the data
Fudging the volcano-plot without dredging the data
Selecting omic biomarkers using both their effect size and their differential status significance (i.e., selecting the “volcano-plot outer spray”) has long been equally biologically relevant and statistically troublesome. However, recent proposals are paving the way to resolving this dilemma.
- Thomas Burger
-
Article
| Open Access TFvelo: gene regulation inspired RNA velocity estimation
TFvelo: gene regulation inspired RNA velocity estimation
Most RNA velocity models extract dynamics from the phase delay between unspliced and spliced mRNA for each gene. Here, authors propose TFvelo, broadening RNA velocity beyond splicing information to include gene regulation. TFvelo accurately models genes dynamics and infers cell pseudo-time from RNA abundance data.
- Jiachen Li
- , Xiaoyong Pan
- & Hong-Bin Shen
-
Article
| Open Access The impacts of active and self-supervised learning on efficient annotation of single-cell expression data
The impacts of active and self-supervised learning on efficient annotation of single-cell expression data
Cell type annotation for single-cell data is challenging. Here, authors explore active and self-supervised learning and introduce adaptive reweighting as a tailored heuristic, demonstrating competitive performance and showing that incorporating prior knowledge enhances cell type annotation accuracy.
- Michael J. Geuenich
- , Dae-won Gong
- & Kieran R. Campbell
-
Article
| Open Access ContScout: sensitive detection and removal of contamination from annotated genomes
ContScout: sensitive detection and removal of contamination from annotated genomes
It is unclear whether naturally evolved de novo proteins have stable, folded structures. Here, systematic identification and structural modeling of de novo genes, this study reveals that a small subset of these proteins may have well-folded structures, and were likely born with these structures.
- Balázs Bálint
- , Zsolt Merényi
- & László G. Nagy
-
Article
| Open Access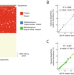 Longitudinal quantification of Bifidobacterium longum subsp. infantis reveals late colonization in the infant gut independent of maternal milk HMO composition
Longitudinal quantification of Bifidobacterium longum subsp. infantis reveals late colonization in the infant gut independent of maternal milk HMO composition
Here, the authors develop a high-throughput method to quantify Bifidobacterium longum subsp. infantis (BL. infantis), a proficient HMO-utilizer, from metagenomic sequencing, and applied it to a longitudinal cohort consisting of 21 mother-infant dyads, suggesting BL. infantis colonization to start late in the breast-feeding period.
- Dena Ennis
- , Shimrit Shmorak
- & Moran Yassour
-
Article
| Open Access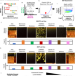 Anti-correlated feature selection prevents false discovery of subpopulations in scRNAseq
Anti-correlated feature selection prevents false discovery of subpopulations in scRNAseq
Typical single-cell RNAseq pipelines will subcluster homogeneous cells. Here, authors present a computational algorithm for accurately identifying cell-type marker genes in single-cell data analysis with a low false discovery rate.
- Scott R. Tyler
- , Daniel Lozano-Ojalvo
- & Eric E. Schadt
-
Article
| Open Access Human whole-exome genotype data for Alzheimer’s disease
Human whole-exome genotype data for Alzheimer’s disease
The heterogeneity of whole-exome sequencing (WES) data generation methods presents a challenge to joint analysis. Here, the authors present a bioinformatics strategy to generate high-quality data from processing diversely generated WES samples, as applied in the Alzheimer’s Disease Sequencing Project.
- Yuk Yee Leung
- , Adam C. Naj
- & Li-San Wang
-
Article
| Open Access ACIDES: on-line monitoring of forward genetic screens for protein engineering
ACIDES: on-line monitoring of forward genetic screens for protein engineering
Screening mutated proteins is a versatile strategy in protein research, producing massive datasets when combined with NGS. Here, authors present ACIDES to estimate mutated protein fitness and aid protein engineering pipelines in a range of applications, including gene therapy.
- Takahiro Nemoto
- , Tommaso Ocari
- & Ulisse Ferrari
-
Article
| Open Access vcfdist: accurately benchmarking phased small variant calls in human genomes
vcfdist: accurately benchmarking phased small variant calls in human genomes
Accurately benchmarking small variant calling accuracy is critical for the continued improvement of human genome sequencing. Here, the authors show that current approaches are biased towards certain variant representations and develop a new approach to ensure consistent and accurate benchmarking, regardless of the original variant representations.
- Tim Dunn
- & Satish Narayanasamy
-
Article
| Open Access Spatial transcriptomics deconvolution at single-cell resolution using Redeconve
Spatial transcriptomics deconvolution at single-cell resolution using Redeconve
Computational deconvolution with single-cell RNA sequencing data as a reference is pivotal for interpreting spatial transcriptomics data. Here, authors present Redeconve, which improves the resolution by more than 100-fold with higher accuracy and speed.
- Zixiang Zhou
- , Yunshan Zhong
- & Xianwen Ren
-
Article
| Open Access CurveCurator: a recalibrated F-statistic to assess, classify, and explore significance of dose–response curves
CurveCurator: a recalibrated F-statistic to assess, classify, and explore significance of dose–response curves
Dose-response curves are ubiquitous in pharmacology and biology, yet potency and effect size are often estimated even when there is no response. Here, authors present a statistical framework to assess curve significance and demonstrate how this aids drug mode of action analysis in large public datasets.
- Florian P. Bayer
- , Manuel Gander
- & Matthew The
-
Article
| Open Access PhenoSV: interpretable phenotype-aware model for the prioritization of genes affected by structural variants
PhenoSV: interpretable phenotype-aware model for the prioritization of genes affected by structural variants
Here, authors present PhenoSV, a phenotype-aware machine-learning model for the functional interpretation of various types of structural variants (SVs) and genes within or outside SVs, facilitating the extraction of biological insights from coding and noncoding SVs.
- Zhuoran Xu
- , Quan Li
- & Kai Wang
-
Article
| Open Access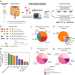 Detection of isoforms and genomic alterations by high-throughput full-length single-cell RNA sequencing in ovarian cancer
Detection of isoforms and genomic alterations by high-throughput full-length single-cell RNA sequencing in ovarian cancer
Long-read single-cell RNA sequencing is capable of detecting isoform-level gene expression and genomic alterations such as mutations and gene fusions, thereby providing cell-specific genotype-phenotype information. Here, the authors use long-read scRNA-seq on metastatic ovarian cancer samples and detect cell-type specific isoforms and gene fusions that may otherwise be misclassified in short-read data.
- Arthur Dondi
- , Ulrike Lischetti
- & Niko Beerenwinkel
-
Article
| Open Access SPACEL: deep learning-based characterization of spatial transcriptome architectures
SPACEL: deep learning-based characterization of spatial transcriptome architectures
Spatial transcriptomics (ST) technologies detect transcript distribution in space. Here, authors present a deep learning based method SPACEL for cell type deconvolution, spatial domain identification and 3D alignment, showcasing it as a valuable toolkit for ST data analysis
- Hao Xu
- , Shuyan Wang
- & Kun Qu
-
Article
| Open Access A statistical framework for differential pseudotime analysis with multiple single-cell RNA-seq samples
A statistical framework for differential pseudotime analysis with multiple single-cell RNA-seq samples
Pseudotime analysis is prevalent in single-cell RNA-seq, but it remains challenging to perform it across multiple samples and experimental conditions. Here, the authors develop Lamian, a computational framework for multi-sample pseudotime analysis that adjusts for biological and technical variation to detect gene program changes along cell trajectories and across conditions.
- Wenpin Hou
- , Zhicheng Ji
- & Hongkai Ji
-
Article
| Open Access A deep population reference panel of tandem repeat variation
A deep population reference panel of tandem repeat variation
Tandem repeats (TRs) comprise some of the most polymorphic regions of the human genome but are difficult to study. Here, the authors develop an ensemble-based genotyping method and characterize 1.7 million TRs across 3,550 humans from diverse populations.
- Helyaneh Ziaei Jam
- , Yang Li
- & Melissa Gymrek
-
Review Article
| Open AccessThe promise of data science for health research in Africa
In this Review article, the authors discuss emerging efforts to build ethical governance frameworks for data science health research in Africa and the opportunities to advance these through investments by African governments and institutions, international funding organizations and collaborations for research and capacity development.
- Clement A. Adebamowo
- , Shawneequa Callier
- & Sally N. Adebamowo
-
Article
| Open Access SiGra: single-cell spatial elucidation through an image-augmented graph transformer
SiGra: single-cell spatial elucidation through an image-augmented graph transformer
Recent advances have pushed spatial transcriptomics to subcellular resolution. Here, the authors propose SiGra, a graph artificial intelligence model designed for high-throughput spatial molecular imaging.
- Ziyang Tang
- , Zuotian Li
- & Qianqian Song
-
Article
| Open Access Placental growth factor exerts a dual function for cardiomyogenesis and vasculogenesis during heart development
Placental growth factor exerts a dual function for cardiomyogenesis and vasculogenesis during heart development
Growth factors play key roles during heart development. Here they show that PLGF has both autocrine and paracrine roles during cardiomyogenesis and vasculogenesis, suggesting it may have therapeutic potential for heart disease.
- Nevin Witman
- , Chikai Zhou
- & Makoto Sahara
-
Article
| Open Access Removal of false positives in metagenomics-based taxonomy profiling via targeting Type IIB restriction sites
Removal of false positives in metagenomics-based taxonomy profiling via targeting Type IIB restriction sites
Here, leveraging species-specific Type IIB restriction endonuclease digestion sites as reference instead of universal markers or whole microbial genomes, the authors introduce MAP2B, a metagenomic profiler, showing it can significantly remove false-positive identification and generate highly accurate taxonomic profiling results.
- Zheng Sun
- , Jiang Liu
- & Yang-Yu Liu
-
Article
| Open Access iCLOTS: open-source, artificial intelligence-enabled software for analyses of blood cells in microfluidic and microscopy-based assays
iCLOTS: open-source, artificial intelligence-enabled software for analyses of blood cells in microfluidic and microscopy-based assays
Microscopy has undoubtedly advanced biomedical research, but novel hypotheses are often lost to a lack of analytical tools. Here authors propose iCLOTS, a freely-available software that allows researchers to apply image processing and artificial intelligence algorithms to their own data.
- Meredith E. Fay
- , Oluwamayokun Oshinowo
- & Wilbur A. Lam
-
Article
| Open Access Systematic review of cnidarian microbiomes reveals insights into the structure, specificity, and fidelity of marine associations
Systematic review of cnidarian microbiomes reveals insights into the structure, specificity, and fidelity of marine associations
This study unified cnidarian microbiome data from 186 studies (~ 6.5 billion sequence reads), providing novel insights into cnidarian microbial communities and highlighting key bacteria across sub-phylum, geography, depth and microhabitat. Understanding factors governing microbiome health will support ongoing and future coral preservation efforts.
- M. McCauley
- , T. L. Goulet
- & S. Loesgen
-
Article
| Open Access Trackable and scalable LC-MS metabolomics data processing using asari
Trackable and scalable LC-MS metabolomics data processing using asari
Reproducible and scalable data processing is key to the progress of metabolomics. Here, the authors present a software tool that offers predictable metabolomics feature detection and improved computational performance in large datasets.
- Shuzhao Li
- , Amnah Siddiqa
- & Shujian Zheng
-
Article
| Open Access DNA 5-methylcytosine detection and methylation phasing using PacBio circular consensus sequencing
DNA 5-methylcytosine detection and methylation phasing using PacBio circular consensus sequencing
Existing methods for detecting DNA methylation (5mC) are less accurate and robust. Here, the authors develop a deep learning tool ccsmeth and a Nextflow pipeline ccsmethphase for genome-wide 5mCpG detection and phasing with high accuracy from CCS reads in human.
- Peng Ni
- , Fan Nie
- & Jianxin Wang
-
Article
| Open Access Histone exchange sensors reveal variant specific dynamics in mouse embryonic stem cells
Histone exchange sensors reveal variant specific dynamics in mouse embryonic stem cells
Eviction of histones from nucleosomes and their exchange with newly synthesized or alternative variants is a central epigenetic determinant. Here the authors implement a molecular sensor that reports on steady-state exchange of histones in mESC and mice revealing dependency between deposition of histone variant H3.3 and exchange of H3.1 and H2B in both open and closed chromatin.
- Marko Dunjić
- , Felix Jonas
- & Yonatan Stelzer
-
Article
| Open Access A high-throughput test enables specific detection of hepatocellular carcinoma
A high-throughput test enables specific detection of hepatocellular carcinoma
DNA methylation analysis is a promising method to detect liver cancer. Here, the authors develop a 5 CpG site signature which can detect HCC at high specificity across multiple cohorts.
- David Cheishvili
- , Chifat Wong
- & Mamun Al Mahtab
-
Article
| Open Access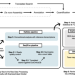 ExpressAnalyst: A unified platform for RNA-sequencing analysis in non-model species
ExpressAnalyst: A unified platform for RNA-sequencing analysis in non-model species
RNA-sequencing data analysis is difficult for non-model species that have no reference genome. ExpressAnalyst enables RNA-sequencing analysis for any eukaryotic species in less than 24 h, on a laptop, and without any programming.
- Peng Liu
- , Jessica Ewald
- & Jianguo Xia
-
Article
| Open Access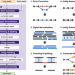 Reference-free assembly of long-read transcriptome sequencing data with RNA-Bloom2
Reference-free assembly of long-read transcriptome sequencing data with RNA-Bloom2
Most existing long-read transcriptome assembly methods rely on reference genomes and transcript annotations, while reference-free methods remain scarce. Here, Nip et al. introduce RNA-Bloom2, a reference-free method that requires substantially less memory and runtime than other reference-free methods.
- Ka Ming Nip
- , Saber Hafezqorani
- & Inanc Birol
-
Article
| Open Access Explainable multi-task learning for multi-modality biological data analysis
Explainable multi-task learning for multi-modality biological data analysis
Multimodal biological data is challenging to analyze. Here, the authors develop UnitedNet, an explainable deep neural network for analyzing single-cell multimodal biological data and estimating relationships between gene expression and other modalities with cell-type specificity.
- Xin Tang
- , Jiawei Zhang
- & Jia Liu
-
Article
| Open Access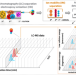 PeakDecoder enables machine learning-based metabolite annotation and accurate profiling in multidimensional mass spectrometry measurements
PeakDecoder enables machine learning-based metabolite annotation and accurate profiling in multidimensional mass spectrometry measurements
Alternative algorithms exploiting advantages of multidimensional mass spectrometry in untargeted metabolomics are needed. Here, the authors develop and demonstrate PeakDecoder for confident and accurate metabolite profiling in 116 microbial sample runs and using a library built from 64 standards.
- Aivett Bilbao
- , Nathalie Munoz
- & Kristin E. Burnum-Johnson
-
Article
| Open Access Vaccination of SARS-CoV-2-infected individuals expands a broad range of clonally diverse affinity-matured B cell lineages
Vaccination of SARS-CoV-2-infected individuals expands a broad range of clonally diverse affinity-matured B cell lineages
Here, the authors isolated and characterized genetic features of spike-specific monoclonal antibodies. They show how the antibodies evolve from infection to after vaccination and conclude that highly polyclonal repertoires of affinity-matured memory B cells are efficiently recalled by vaccination.
- Mark Chernyshev
- , Mrunal Sakharkar
- & Gunilla B. Karlsson Hedestam
-
Article
| Open Access Systematic comparison of tools used for m6A mapping from nanopore direct RNA sequencing
Systematic comparison of tools used for m6A mapping from nanopore direct RNA sequencing
Direct RNA sequencing using nanopore platform can be used to detect N6-methyladenosine (m6A) modifications on mRNAs. Here the authors systematically compare tools used for m6A detection from nanopore direct sequencing.
- Zhen-Dong Zhong
- , Ying-Yuan Xie
- & Guan-Zheng Luo
-
Article
| Open Access Integrated analysis of genomic and transcriptomic data for the discovery of splice-associated variants in cancer
Integrated analysis of genomic and transcriptomic data for the discovery of splice-associated variants in cancer
Analysing the regulatory consequences of mutations and splice variants at large scale in cancer requires efficient computational tools. Here, the authors develop RegTools, a software package that can identify splice-associated variants from large-scale genomics and transcriptomics data with efficiency and flexibility.
- Kelsy C. Cotto
- , Yang-Yang Feng
- & Malachi Griffith
-
Article
| Open Access Determining protein structures in cellular lamella at pseudo-atomic resolution by GisSPA
Determining protein structures in cellular lamella at pseudo-atomic resolution by GisSPA
High-resolution in situ protein structure can be solved by cryo-ET, which requires several days of data collection. Here Cheng et al. report GisSPA, a program that may enable determining sub-4 Å resolution structures on cellular lamellae within one day of data collection.
- Jing Cheng
- , Tong Liu
- & Xinzheng Zhang
-
Article
| Open Access Deciphering the exact breakpoints of structural variations using long sequencing reads with DeBreak
Deciphering the exact breakpoints of structural variations using long sequencing reads with DeBreak
Long-read sequencing is promising for the detection of structural variants (SVs), which requires algorithms with high sensitivity and precision. Here, the authors develop DeBreak, an algorithm for comprehensive and accurate SV detection in long-read sequencing data across different platforms, which outperforms other SV callers.
- Yu Chen
- , Amy Y. Wang
- & Zechen Chong
-
Article
| Open Access Region-specific denoising identifies spatial co-expression patterns and intra-tissue heterogeneity in spatially resolved transcriptomics data
Region-specific denoising identifies spatial co-expression patterns and intra-tissue heterogeneity in spatially resolved transcriptomics data
Spatially resolved transcriptomics is a relatively new technique that maps transcriptional information within a tissue. Here the authors present MIST, which detects molecular regions from spatially resolved transcriptomics and denoises the missing gene expression values by region-specific imputation.
- Linhua Wang
- , Mirjana Maletic-Savatic
- & Zhandong Liu
-
Article
| Open Access A flexible cross-platform single-cell data processing pipeline
A flexible cross-platform single-cell data processing pipeline
As the throughput of single-cell RNA-seq studies increases, there is a need for tools that can make the data analysis steps more streamlined and convenient. Here, the authors develop UniverSC, a tool that unifies single-cell RNA-seq analysis workflows and also facilitates their use for non-experts.
- Kai Battenberg
- , S. Thomas Kelly
- & Aki Minoda
-
Article
| Open Access Metabolite annotation from knowns to unknowns through knowledge-guided multi-layer metabolic networking
Metabolite annotation from knowns to unknowns through knowledge-guided multi-layer metabolic networking
Unknown metabolite annotation is a grand challenge in untargeted metabolomics. Here, the authors develop knowledge-guided multi-layer networking (KGMN) to enable global metabolite annotation from knowns to unknowns in untargeted metabolomics.
- Zhiwei Zhou
- , Mingdu Luo
- & Zheng-Jiang Zhu
-
Article
| Open Access Inferring differential subcellular localisation in comparative spatial proteomics using BANDLE
Inferring differential subcellular localisation in comparative spatial proteomics using BANDLE
Changes in protein subcellular localization can be determined using mass spectrometry. Here, the authors present a statistical approach to determine relocalising proteins from spatial proteomics experiments.
- Oliver M. Crook
- , Colin T. R. Davies
- & Kathryn S. Lilley
-
Article
| Open Access A method for multiplexed full-length single-molecule sequencing of the human mitochondrial genome
A method for multiplexed full-length single-molecule sequencing of the human mitochondrial genome
Accurate analysis of mitochondrial DNA is important for mitochondrial disease clinical research and diagnostics. Here, authors present a method using Cas9 cleavage, nanopore sequencing and a custom pipeline to identify pathogenic variants, deletions and accurately quantify heteroplasmy to below 1%.
- Ieva Keraite
- , Philipp Becker
- & Ivo Glynne Gut
-
Article
| Open Access Batch effects removal for microbiome data via conditional quantile regression
Batch effects removal for microbiome data via conditional quantile regression
Here, the authors present ConQuR, a conditional quantile regression method that removes microbiome batch effects through non-parametric modeling of complex microbial read counts, while preserving the signals of interest.
- Wodan Ling
- , Jiuyao Lu
- & Michael C. Wu
-
Article
| Open Access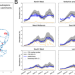 An analysis of 45 large-scale wastewater sites in England to estimate SARS-CoV-2 community prevalence
An analysis of 45 large-scale wastewater sites in England to estimate SARS-CoV-2 community prevalence
Wastewater surveillance could provide a means of monitoring SARS-CoV-2 prevalence that does not rely on testing individuals. Here, the authors report results from England’s national wastewater surveillance program, use it to estimate prevalence, and compare estimates with those from population-based prevalence surveys.
- Mario Morvan
- , Anna Lo Jacomo
- & Leon Danon
-
Article
| Open Access Benchmarking of analysis strategies for data-independent acquisition proteomics using a large-scale dataset comprising inter-patient heterogeneity
Benchmarking of analysis strategies for data-independent acquisition proteomics using a large-scale dataset comprising inter-patient heterogeneity
Data independent acquisition (DIA) has been gaining momentum in clinical proteomics. Here, the authors create a benchmark dataset comprising inter-patient heterogeneity to compare popular DIA data analysis workflows for identifying differentially abundant proteins.
- Klemens Fröhlich
- , Eva Brombacher
- & Oliver Schilling
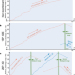
 Data-driven recombination detection in viral genomes
Data-driven recombination detection in viral genomes