Article
|
Open Access
Featured
-
-
Article
| Open Access TAGET: a toolkit for analyzing full-length transcripts from long-read sequencing
TAGET: a toolkit for analyzing full-length transcripts from long-read sequencing
Accurate long-read RNA sequencing facilitates analysis of full-length transcripts. Here the authors develop an integrative toolkit, optimised for Iso-Seq data analysis, that includes transcript alignment, annotation, quantification and gene fusion detection.
- Yuchao Xia
- , Zijie Jin
- & Ruibin Xi
-
Article
| Open Access Deciphering the exact breakpoints of structural variations using long sequencing reads with DeBreak
Deciphering the exact breakpoints of structural variations using long sequencing reads with DeBreak
Long-read sequencing is promising for the detection of structural variants (SVs), which requires algorithms with high sensitivity and precision. Here, the authors develop DeBreak, an algorithm for comprehensive and accurate SV detection in long-read sequencing data across different platforms, which outperforms other SV callers.
- Yu Chen
- , Amy Y. Wang
- & Zechen Chong
-
Article
| Open Access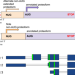 Thousands of human non-AUG extended proteoforms lack evidence of evolutionary selection among mammals
Thousands of human non-AUG extended proteoforms lack evidence of evolutionary selection among mammals
Analysis of a large number of Ribo-seq datasets and genomic alignments led to detection of novel non-AUG proteoforms. Unexpectedly the number of non-AUG proteoforms identified with Ribo-seq greatly exceeds those with strong phylogenetic support.
- Alla D. Fedorova
- , Stephen J. Kiniry
- & Pavel V. Baranov
-
Article
| Open Access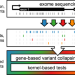 Identifying interpretable gene-biomarker associations with functionally informed kernel-based tests in 190,000 exomes
Identifying interpretable gene-biomarker associations with functionally informed kernel-based tests in 190,000 exomes
Genetic association studies for rare variants suffer from lack of power and thus there is a need for methods to improve rare variant discovery. Here, the authors present functionally informed association tests with increased statistical power to aid discovery and interpretation of rare variants.
- Remo Monti
- , Pia Rautenstrauch
- & Christoph Lippert
-
Article
| Open Access TP53-dependent toxicity of CRISPR/Cas9 cuts is differential across genomic loci and can confound genetic screening
TP53-dependent toxicity of CRISPR/Cas9 cuts is differential across genomic loci and can confound genetic screening
Toxicity of CRISPR/Cas9 induced DNA breaks depends on their repair mechanism, and on the chromatin environment at the cut site. Here the authors show that edits in active genes or regulatory elements can incur a higher toxicity via a TP53-dependent mechanism.
- Miguel M. Álvarez
- , Josep Biayna
- & Fran Supek
-
Article
| Open Access Leveraging omic features with F3UTER enables identification of unannotated 3’UTRs for synaptic genes
Leveraging omic features with F3UTER enables identification of unannotated 3’UTRs for synaptic genes
3’ untranslated regions (3’UTRs) play a crucial role in regulating gene expression, but our 3’UTR catalogue is incomplete. Here, the authors develop a machine learning-based framework to predict previously unannotated 3’UTRs in 39 human tissues.
- Siddharth Sethi
- , David Zhang
- & Juan A. Botia
-
Article
| Open Access Helical structure motifs made searchable for functional peptide design
Helical structure motifs made searchable for functional peptide design
Here, we present TP-DB; a pattern-based search engine based on 1.67 million helices from the Protein Database (PDB). We demonstrate the utility of TP-DB in identifying microbe-specific antigens, as well as the design of antimicrobial peptides and Protein-protein interaction blockers.
- Cheng-Yu Tsai
- , Emmanuel Oluwatobi Salawu
- & Lee-Wei Yang
-
Article
| Open Access R2DT is a framework for predicting and visualising RNA secondary structure using templates
R2DT is a framework for predicting and visualising RNA secondary structure using templates
Non-coding RNA function is poorly understood, partly due to the challenge of determining RNA secondary (2D) structure. Here, the authors present a framework for the reproducible prediction and visualization of the 2D structure of a wide array of RNAs, which enables linking RNA sequence to function.
- Blake A. Sweeney
- , David Hoksza
- & Anton I. Petrov
-
Article
| Open Access SARS-CoV-2 gene content and COVID-19 mutation impact by comparing 44 Sarbecovirus genomes
SARS-CoV-2 gene content and COVID-19 mutation impact by comparing 44 Sarbecovirus genomes
The SARS-CoV-2 gene set remains unresolved, hindering dissection of COVID-19 biology. Comparing 44 Sarbecovirus genomes provides a high-confidence protein-coding gene set. The study characterizes protein-level and nucleotide-level evolutionary constraints, and prioritizes functional mutations from the ongoing COVID-19 pandemic.
- Irwin Jungreis
- , Rachel Sealfon
- & Manolis Kellis
-
Article
| Open Access Uncovering transcriptional dark matter via gene annotation independent single-cell RNA sequencing analysis
Uncovering transcriptional dark matter via gene annotation independent single-cell RNA sequencing analysis
Conventional single-cell RNA sequencing analysis rely on genome annotations that may be incomplete or inaccurate especially for understudied organisms. Here the authors present a bioinformatic tool that leverages single-cell data to uncover biologically relevant transcripts beyond the best available genome annotation.
- Michael F. Z. Wang
- , Madhav Mantri
- & Iwijn De Vlaminck
-
Article
| Open Access MVP predicts the pathogenicity of missense variants by deep learning
MVP predicts the pathogenicity of missense variants by deep learning
Accurate prediction of variant pathogenicity is essential to understanding genetic risks in disease. Here, the authors present a deep neural network method for prediction of missense variant pathogenicity, MVP, and demonstrate its utility in prioritizing de novo variants contributing to developmental disorders.
- Hongjian Qi
- , Haicang Zhang
- & Yufeng Shen
-
Article
| Open Access Donkey genomes provide new insights into domestication and selection for coat color
Donkey genomes provide new insights into domestication and selection for coat color
A new donkey reference genome and comparisons with wild asses yields insights into the evolutionary history of donkey domestication and identifies a genetic variant that results in the non-Dun coat colours of domestic donkeys.
- Changfa Wang
- , Haijing Li
- & Jifeng Zhong
-
Article
| Open Access A possible universal role for mRNA secondary structure in bacterial translation revealed using a synthetic operon
A possible universal role for mRNA secondary structure in bacterial translation revealed using a synthetic operon
The mechanisms for regulating translation re-initiation in bacteria remain poorly understood. Here, the authors screened a library of synthetic operons and identified a ribosome termination structure that modulates re-initiation efficiency and which is conserved across bacteria.
- Yonatan Chemla
- , Michael Peeri
- & Lital Alfonta
-
Article
| Open Access Improved haplotype inference by exploiting long-range linking and allelic imbalance in RNA-seq datasets
Improved haplotype inference by exploiting long-range linking and allelic imbalance in RNA-seq datasets
Haplotype reconstruction of distant genetic variants is problematic in short-read sequencing. Here, the authors describe HapTree-X, a probabilistic framework that uses differential allele-specific expression to better reconstruct paternal haplotypes from diploid and polyploid genomes.
- Emily Berger
- , Deniz Yorukoglu
- & Bonnie Berger
-
Article
| Open Access Full-length transcriptome reconstruction reveals a large diversity of RNA and protein isoforms in rat hippocampus
Full-length transcriptome reconstruction reveals a large diversity of RNA and protein isoforms in rat hippocampus
It is challenging to characterize diverse transcript isoforms by short-read sequencing. Here the authors report full-length transcriptomes in rat hippocampus by hybrid-sequencing, predict isoform-specific translational status, and reconstruct open reading frames validated by mass spectrometry.
- Xi Wang
- , Xintian You
- & Wei Chen
-
Article
| Open Access Multi-platform discovery of haplotype-resolved structural variation in human genomes
Multi-platform discovery of haplotype-resolved structural variation in human genomes
Structural variants (SVs) in human genomes contribute diversity and diseases. Here, the authors use a multi-platform strategy to generate haplotype-resolved SVs for three human parent–child trios.
- Mark J. P. Chaisson
- , Ashley D. Sanders
- & Charles Lee
-
Article
| Open Access Biological relevance of computationally predicted pathogenicity of noncoding variants
Biological relevance of computationally predicted pathogenicity of noncoding variants
Researchers can make use of a variety of computational tools to prioritize genetic variants and predict their pathogenicity. Here, the authors evaluate the performance of six of these tools in three typical biological tasks and find generally low concordance of predictions and experimental confirmation.
- Li Liu
- , Maxwell D. Sanderford
- & Sudhir Kumar
-
Article
| Open Access Germline pathogenic variants of 11 breast cancer genes in 7,051 Japanese patients and 11,241 controls
Germline pathogenic variants of 11 breast cancer genes in 7,051 Japanese patients and 11,241 controls
Association between variants in 11 different genes and breast cancer risk has been established and sequencing of these genes is recommended to provide personalized diagnosis, therapy, and surveillance for the high-risk patients and their relatives. Here the authors analyse the frequency of germline pathogenic mutations in these genes specifically in a Japanese population.
- Yukihide Momozawa
- , Yusuke Iwasaki
- & Michiaki Kubo
-
Article
| Open Access The North American bullfrog draft genome provides insight into hormonal regulation of long noncoding RNA
The North American bullfrog draft genome provides insight into hormonal regulation of long noncoding RNA
The globally-distributed Ranidae (true frogs) are the largest frog family. Here, Hammond et al. present a draft genome of the North American bullfrog, Rana (Lithobates) catesbeiana, as a foundation for future understanding of true frog genetics as amphibian species face difficult environmental challenges.
- S. Austin Hammond
- , René L. Warren
- & Inanc Birol
-
Article
| Open Access Annotating pathogenic non-coding variants in genic regions
Annotating pathogenic non-coding variants in genic regions
While non-coding synonymous and intronic variants are often not under strong selective constraint, they can be pathogenic through affecting splicing or transcription. Here, the authors develop a score that uses sequence context alterations to predict pathogenicity of synonymous and non-coding genetic variants, and provide a web server of pre-computed scores.
- Sahar Gelfman
- , Quanli Wang
- & David B. Goldstein
-
Article
| Open Access Extension of human lncRNA transcripts by RACE coupled with long-read high-throughput sequencing (RACE-Seq)
Extension of human lncRNA transcripts by RACE coupled with long-read high-throughput sequencing (RACE-Seq)
Long non-coding RNAs are increasingly recognised to be important factors in regulating cellular processes and comprise a large faction of the transcriptome, however most are uncharacterised. Here the authors present RACE-Seq, a tool to improve and extend the annotation of low-expression transcripts.
- Julien Lagarde
- , Barbara Uszczynska-Ratajczak
- & Jennifer Harrow

 De novo genome assembly depicts the immune genomic characteristics of cattle
De novo genome assembly depicts the immune genomic characteristics of cattle