Key Points
-
Metagenome-wide association studies (MWAS) of human disease are now possible, owing to advances in DNA sequencing and the development of reference gene catalogues and gene clustering methods.
-
MWAS have identified associations between the microbiome and several major diseases, despite the relatively small sample sizes that have been examined by these studies compared with genome-wide association studies (GWAS).
-
Common changes to the taxa and functions of the gut microbiota have emerged from MWAS of metabolic diseases.
-
To advance from the detection of associations to the demonstration that elements of the microbiota contribute to disease will require a range of validations, including mechanistic studies both in vivo and in vitro. However, even associations that are shown to be non-causal could form the basis of diagnostic markers.
-
In the future, MWAS may be further developed in numerous ways, including the use of multiomic data, the analysis of genetic variants in metagenomic data and the study of the non-bacterial components of the microbiome.
Abstract
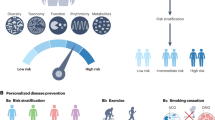
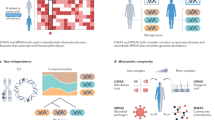
Metagenome-wide association studies (MWAS) have enabled the high-resolution investigation of associations between the human microbiome and several complex diseases, including type 2 diabetes, obesity, liver cirrhosis, colorectal cancer and rheumatoid arthritis. The associations that can be identified by MWAS are not limited to the identification of taxa that are more or less abundant, as is the case with taxonomic approaches, but additionally include the identification of microbial functions that are enriched or depleted. In this Review, we summarize recent findings from MWAS and discuss how these findings might inform the prevention, diagnosis and treatment of human disease in the future. Furthermore, we highlight the need to better characterize the biology of many of the bacteria that are found in the human microbiota as an essential step in understanding how bacterial strains that have been identified by MWAS are associated with disease.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Qin, J. et al. A human gut microbial gene catalogue established by metagenomic sequencing. Nature 464, 59–65 (2010). This study details the first gene catalogue of the human gut microbiome that is assembled from next-generation sequencing data.
Methé, B. A. et al. A framework for human microbiome research. Nature 486, 215–221 (2012).
Li, J. et al. An integrated catalog of reference genes in the human gut microbiome. Nat. Biotechnol. 32, 834–841 (2014). This study details a high-quality reference gene catalogue that is compiled from 1,267 samples across three continents, and identifies many differences in the gut microbiome between healthy Chinese and Danish individuals.
Clemente, J. C., Ursell, L. K., Parfrey, L. W. & Knight, R. The impact of the gut microbiota on human health: an integrative view. Cell 148, 1258–1270 (2012).
Sommer, F. & Bäckhed, F. The gut microbiota — masters of host development and physiology. Nat. Rev. Microbiol. 11, 227–238 (2013).
Marchesi, J. R. et al. The gut microbiota and host health: a new clinical frontier. Gut 65, 330–339 (2015).
Xu, Y. et al. Prevalence and control of diabetes in Chinese adults. JAMA 310, 948–959 (2013).
Larsen, N. et al. Gut microbiota in human adults with type 2 diabetes differs from non-diabetic adults. PLoS ONE 5, e9085 (2010).
Qin, J. et al. A metagenome-wide association study of gut microbiota in type 2 diabetes. Nature 490, 55–60 (2012). The first MWAS, which establishes the MLG method and identifies associations between the gut microbiome and type 2 diabetes.
Brubaker, P. L. & Anini, Y. Direct and indirect mechanisms regulating secretion of glucagon-like peptide-1 and glucagon-like peptide-2. Can. J. Physiol. Pharmacol. 81, 1005–1012 (2003).
Zhou, J. et al. Peptide YY and proglucagon mRNA expression patterns and regulation in the gut. Obesity (Silver Spring) 14, 683–689 (2006).
Sun, J. et al. Pancreatic β-cells limit autoimmune diabetes via an immunoregulatory antimicrobial peptide expressed under the influence of the gut microbiota. Immunity 43, 304–317 (2015).
Karlsson, F. H. et al. Gut metagenome in European women with normal, impaired and diabetic glucose control. Nature 498, 99–103 (2013).
Pyra, K. A., Saha, D. C. & Reimer, R. A. Prebiotic fiber increases hepatic acetyl CoA carboxylase phosphorylation and suppresses glucose-dependent insulinotropic polypeptide secretion more effectively when used with metformin in obese rats. J. Nutr. 142, 213–220 (2012).
Forslund, K. et al. Disentangling type 2 diabetes and metformin treatment signatures in the human gut microbiota. Nature 528, 262–266 (2015).
Shin, N.-R. et al. An increase in the Akkermansia spp. population induced by metformin treatment improves glucose homeostasis in diet-induced obese mice. Gut 63, 727–735 (2014).
Lee, H. & Ko, G. Effect of metformin on metabolic improvement and gut microbiota. Appl. Environ. Microbiol. 80, 5935–5943 (2014).
Le Chatelier, E. et al. Richness of human gut microbiome correlates with metabolic markers. Nature 500, 541–546 (2013).
Cotillard, A. et al. Dietary intervention impact on gut microbial gene richness. Nature 500, 585–588 (2013).
Ley, R. E., Turnbaugh, P. J., Klein, S. & Gordon, J. I. Microbial ecology: human gut microbes associated with obesity. Nature 444, 1022–1023 (2006).
Turnbaugh, P. J. et al. A core gut microbiome in obese and lean twins. Nature 457, 480–484 (2009).
Jemal, A. et al. Global cancer statistics. CA Cancer J. Clin. 61, 69–90 (2011).
Ferlay, J. et al. Cancer incidence and mortality worldwide: sources, methods and major patterns in GLOBOCAN 2012. Int. J. Cancer 136, E359–E386 (2015).
Brenner, H., Kloor, M. & Pox, C. P. Colorectal cancer. Lancet 383, 1490–1502 (2014).
Kostic, A. D. et al. Genomic analysis identifies association of Fusobacterium with colorectal carcinoma. Genome Res. 22, 292–298 (2012).
Castellarin, M. et al. Fusobacterium nucleatum infection is prevalent in human colorectal carcinoma. Genome Res. 22, 299–306 (2012).
Kostic, A. D. et al. Fusobacterium nucleatum potentiates intestinal tumorigenesis and modulates the tumor-immune microenvironment. Cell Host Microbe 14, 207–215 (2013).
Gur, C. et al. Binding of the Fap2 protein of Fusobacterium nucleatum to human inhibitory receptor TIGIT protects tumors from immune cell attack. Immunity 42, 344–355 (2015).
Cho, I. & Blaser, M. J. The human microbiome: at the interface of health and disease. Nat. Rev. Genet. 13, 260–270 (2012).
Weir, T. L. et al. Stool microbiome and metabolome differences between colorectal cancer patients and healthy adults. PLoS ONE 8, e70803 (2013).
Zeller, G. et al. Potential of fecal microbiota for early-stage detection of colorectal cancer. Mol. Syst. Biol. 10, 766 (2014).
Feng, Q. et al. Gut microbiome development along the colorectal adenoma–carcinoma sequence. Nat. Commun. 6, 6528 (2015).
Yu, J. et al. Metagenomic analysis of faecal microbiome as a tool towards targeted non-invasive biomarkers for colorectal cancer. Gut http://dx.doi.org/10.1136/gutjnl-2015-309800 (2015).
Marchesi, J. R. et al. Towards the human colorectal cancer microbiome. PLoS ONE 6, e20447 (2011).
Nakatsu, G. et al. Gut mucosal microbiome across stages of colorectal carcinogenesis. Nat. Commun. 6, 8727 (2015).
Tjalsma, H., Boleij, A., Marchesi, J. R. & Dutilh, B. E. A bacterial driver-passenger model for colorectal cancer: beyond the usual suspects. Nat. Rev. Microbiol. 10, 575–582 (2012).
Louis, P., Hold, G. L. & Flint, H. J. The gut microbiota, bacterial metabolites and colorectal cancer. Nat. Rev. Microbiol. 12, 661–672 (2014).
Iida, N. et al. Commensal bacteria control cancer response to therapy by modulating the tumor microenvironment. Science 342, 967–970 (2013).
Viaud, S. et al. The intestinal microbiota modulates the anticancer immune effects of cyclophosphamide. Science 342, 971–976 (2013).
Schwabe, R. F. & Jobin, C. The microbiome and cancer. Nat. Rev. Cancer 13, 800–812 (2013).
Vetizou, M. et al. Anticancer immunotherapy by CTLA-4 blockade relies on the gut microbiota. Science 350, 1079–1084 (2015).
Sivan, A. et al. Commensal Bifidobacterium promotes antitumor immunity and facilitates anti-PD-L1 efficacy. Science 350, 1084–1089 (2015).
McInnes, I. B. & Schett, G. The pathogenesis of rheumatoid arthritis. N. Engl. J. Med. 365, 2205–2219 (2011).
Demoruelle, M. K., Deane, K. D. & Holers, V. M. When and where does inflammation begin in rheumatoid arthritis? Curr. Opin. Rheumatol. 26, 2264–2271 (2014).
Zhang, X. et al. The oral and gut microbiomes are perturbed in rheumatoid arthritis and partly normalized after treatment. Nat. Med. 21, 895–905 (2015). This study extends MWAS to the oral microbiome, and identifies potential markers in the oral and gut microbiomes for rheumatoid arthritis and its treatment by drugs.
Scher, J. U. et al. Expansion of intestinal Prevotella copri correlates with enhanced susceptibility to arthritis. eLife 2, e01202 (2013).
Scher, J. U. et al. Periodontal disease and the oral microbiota in new-onset rheumatoid arthritis. Arthritis Rheum. 64, 3083–3094 (2012).
Deane, K. D. & El-Gabalawy, H. Pathogenesis and prevention of rheumatic disease: focus on preclinical RA and SLE. Nat. Rev. Rheumatol. 10, 212–228 (2014).
Karlsson, F. H. et al. Symptomatic atherosclerosis is associated with an altered gut metagenome. Nat. Commun. 3, 1245 (2012).
Fu, J. et al. The gut microbiome contributes to a substantial proportion of the variation in blood lipids. Circ. Res. 117, 817–824 (2015).
Qin, N. et al. Alterations of the human gut microbiome in liver cirrhosis. Nature 513, 859–864 (2014).
Ding, T. & Schloss, P. D. Dynamics and associations of microbial community types across the human body. Nature 509, 357–360 (2014).
Bäckhed, F. et al. Dynamics and stabilization of the human gut microbiome during the first year of life. Cell Host Microbe 17, 690–703 (2015). This study details the largest cohort of infants for which gut microbiomes have been longitudinally profiled for one year from birth; draft genomes were assembled from each sample through the binning of contigs instead of genes.
Sonnenburg, J. L. et al. Glycan foraging in vivo by an intestine-adapted bacterial symbiont. Science 307, 1955–1959 (2005).
Koropatkin, N. M., Cameron, E. A. & Martens, E. C. How glycan metabolism shapes the human gut microbiota. Nat. Rev. Microbiol. 10, 323–335 (2012).
Kashnap, P. et al. Genetically dictated change in host mucus carbohydrate landscape exerts a diet-dependent effect on gut microbiota. Proc. Natl Acad. Sci. USA 110, 17059–17064 (2013).
Lee, S. M. et al. Bacterial colonization factors control specificity and stability of the gut microbiota. Nature 501, 426–429 (2013). This study identifies glycan-utilizing colonization factors in Bacteroides spp. that are responsible for saturable colonization of individual Bacteroides species in mice.
Motta, J.-P. et al. Hydrogen sulfide protects from colitis and restores intestinal microbiota biofilm and mucus production. Inflamm. Bowel Dis. 21, 1006–1017 (2015).
Benson, A. K. et al. Individuality in gut microbiota composition is a complex polygenic trait shaped by multiple environmental and host genetic factors. Proc. Natl Acad. Sci. USA 107, 18933–18938 (2010).
Benson, A. K. Host genetic architecture and the landscape of microbiome composition: humans weigh in. Genome Biol. 16, 203 (2015).
Goodrich, J. K. et al. Human genetics shape the gut microbiome. Cell 159, 789–799 (2014).
van Opstal, E. J. & Bordenstein, S. R. Rethinking heritability of the microbiome. Science 349, 1172–1173 (2015).
Rausch, P. et al. Colonic mucosa-associated microbiota is influenced by an interaction of Crohn disease and FUT2 (Secretor) genotype. Proc. Natl Acad. Sci. USA 108, 19030–19035 (2011).
Pickard, J. M. et al. Rapid fucosylation of intestinal epithelium sustains host–commensal symbiosis in sickness. Nature 514, 638–641 (2014).
Goto, Y. et al. Innate lymphoid cells regulate intestinal epithelial cell glycosylation. Science 345, 1254009 (2014).
Franzosa, E. A. et al. Sequencing and beyond: integrating molecular 'omics' for microbial community profiling. Nat. Rev. Microbiol. 13, 360–372 (2015).
Sridharan, G. V. et al. Prediction and quantification of bioactive microbiota metabolites in the mouse gut. Nat. Commun. 5, 5492 (2014).
Shoaie, S. et al. Quantifying diet-induced metabolic changes of the human gut microbiome. Cell. Metab. 22, 320–331 (2015).
Xu, J. et al. Structural modulation of gut microbiota during alleviation of type 2 diabetes with a Chinese herbal formula. ISME J. 9, 552–562 (2014).
Lukens, J. R. et al. Dietary modulation of the microbiome affects autoinflammatory disease. Nature 516, 246–249 (2014).
Hsiao, E. Y. et al. Microbiota modulate behavioral and physiological abnormalities associated with neurodevelopmental disorders. Cell 155, 1451–1463 (2013).
Atarashi, K. et al. Treg induction by a rationally selected mixture of Clostridia strains from the human microbiota. Nature 500, 232–236 (2013).
Ridaura, V. K. et al. Gut microbiota from twins discordant for obesity modulate metabolism in mice. Science 341, 1241214 (2013). This study shows that co-housing mice that have received microbial transplants from an obese twin with mice that have received microbial transplants from a lean twin prevents the development of obesity-associated phenotypes.
Xiao, L. et al. A catalog of the mouse gut metagenome. Nat. Biotechnol. 33, 1103–1108 (2015). This paper details the first gene catalogue for the gut microbiome of laboratory mice, which reports differences from the human gut microbiome as well as between mice.
Bäckhed, F., Manchester, J. K., Semenkovich, C. F. & Gordon, J. I. Mechanisms underlying the resistance to diet-induced obesity in germ-free mice. Proc. Natl Acad. Sci. USA 104, 979–984 (2007).
Mukherji, A., Kobiita, A., Ye, T. & Chambon, P. Homeostasis in intestinal epithelium is orchestrated by the circadian clock and microbiota cues transduced by TLRs. Cell 153, 812–827 (2013).
Olszak, T. et al. Microbial exposure during early life has persistent effects on natural killer T cell function. Science 336, 489–493 (2012).
Braniste, V. et al. The gut microbiota influences blood–brain barrier permeability in mice. Sci. Transl Med. 6, 263ra158 (2014).
Erny, D. et al. Host microbiota constantly control maturation and function of microglia in the CNS. Nat. Neurosci. 18, 965–977 (2015).
Lacombe, A. et al. Lowbush wild blueberries have the potential to modify gut microbiota and xenobiotic metabolism in the rat colon. PLoS ONE 8, e67497 (2013).
Hildebrand, F. et al. A comparative analysis of the intestinal metagenomes present in guinea pigs (Cavia porcellus) and humans (Homo sapiens). BMC Genomics 13, 514 (2012).
Cabreiro, F. & Gems, D. Worms need microbes too: microbiota, health and aging in Caenorhabditis elegans. EMBO Mol. Med. 5, 1300–1310 (2013).
Turnbaugh, P. J. et al. An obesity-associated gut microbiome with increased capacity for energy harvest. Nature 444, 1027–1031 (2006).
Lukovac, S. et al. Differential modulation by Akkermansia muciniphila and Faecalibacterium prausnitzii of host peripheral lipid metabolism and histone acetylation in mouse gut organoids. mBio 5, e01438-14 (2014).
Kim, K., Lee, S. & Ryu, C.-M. Interspecific bacterial sensing through airborne signals modulates locomotion and drug resistance. Nat. Commun. 4, 1809 (2013).
Cuskin, F. et al. Human gut Bacteroidetes can utilize yeast mannan through a selfish mechanism. Nature 517, 165–169 (2015).
Stowell, S. R. et al. Microbial glycan microarrays define key features of host-microbial interactions. Nat. Chem. Biol. 10, 470–476 (2014).
Wang, X. et al. Cloning and variation of ground state intestinal stem cells. Nature 522, 173–178 (2015).
Joice, R., Yasuda, K., Shafquat, A., Morgan, X. C. & Huttenhower, C. Determining microbial products and identifying molecular targets in the human microbiome. Cell. Metab. 20, 731–741 (2014).
Dobkin, J. F., Saha, J. R., Butler, V. P., Neu, H. C. & Lindenbaum, J. Inactivation of digoxin by Eubacterium lentum, an anaerobe of the human gut flora. Trans. Assoc. Am. Physicians 95, 22–29 (1982).
Haiser, H. J. et al. Predicting and manipulating cardiac drug inactivation by the human gut bacterium Eggerthella lenta. Science 341, 295–298 (2013).
Albertsen, M. et al. Genome sequences of rare, uncultured bacteria obtained by differential coverage binning of multiple metagenomes. Nat. Biotechnol. 31, 533–538 (2013).
Nielsen, H. B. et al. Identification and assembly of genomes and genetic elements in complex metagenomic samples without using reference genomes. Nat. Biotechnol. 32, 822–828 (2014). This study reports the use of MGSs to assemble genomes, 238 of which met the Human Microbiome Project (HMP) high-quality draft genome standard.
Kuleshov, V. et al. Synthetic long-read sequencing reveals intraspecies diversity in the human microbiome. Nat. Biotechnol. 34, 64–69 (2015).
McLean, J. S. et al. Candidate phylum TM6 genome recovered from a hospital sink biofilm provides genomic insights into this uncultivated phylum. Proc. Natl Acad. Sci. USA 110, E2390–E2399 (2013).
Rinke, C. et al. Insights into the phylogeny and coding potential of microbial dark matter. Nature 499, 431–437 (2013).
Reyes, A. et al. Viruses in the faecal microbiota of monozygotic twins and their mothers. Nature 466, 334–338 (2010).
Smillie, C. S. et al. Ecology drives a global network of gene exchange connecting the human microbiome. Nature 480, 241–244 (2011).
Minot, S. et al. The human gut virome: inter-individual variation and dynamic response to diet. Genome Res. 21, 1616–1625 (2011).
Modi, S. R., Lee, H. H., Spina, C. S. & Collins, J. J. Antibiotic treatment expands the resistance reservoir and ecological network of the phage metagenome. Nature 499, 219–222 (2013).
Norman, J. M. et al. Disease-specific alterations in the enteric virome in inflammatory bowel disease. Cell 160, 447–460 (2015).
Schloissnig, S. et al. Genomic variation landscape of the human gut microbiome. Nature 493, 45–50 (2012). This study represents the first analysis of genomic variations, such as SNPs, in the gut microbiome.
Hu, Y. et al. Metagenome-wide analysis of antibiotic resistance genes in a large cohort of human gut microbiota. Nat. Commun. 4, 2151 (2013).
Greenblum, S., Carr, R. & Borenstein, E. Extensive strain-level copy-number variation across human gut microbiome species. Cell 160, 583–594 (2015).
Aagaard, K. et al. The placenta harbors a unique microbiome. Sci. Transl Med. 6, 237ra65 (2014).
Oh, J. et al. Biogeography and individuality shape function in the human skin metagenome. Nature 514, 59–64 (2014).
Kultima, J. R. et al. MOCAT: a metagenomics assembly and gene prediction toolkit. PLoS ONE 7, e47656 (2012).
Human Microbiome Project Consortium. Structure, function and diversity of the healthy human microbiome. Nature 486, 207–214 (2012).
Arumugam, M. et al. Enterotypes of the human gut microbiome. Nature 473, 174–180 (2011).
Sunagawa, S. et al. Metagenomic species profiling using universal phylogenetic marker genes. Nat. Methods 10, 1196–1199 (2013).
Mende, D. R., Sunagawa, S., Zeller, G. & Bork, P. Accurate and universal delineation of prokaryotic species. Nat. Methods 10, 881–884 (2013).
Dupont, C. L. et al. Genomic insights to SAR86, an abundant and uncultivated marine bacterial lineage. ISME J. 6, 1186–1199 (2012).
Freitas, T. A., Li, P.-E., Scholz, M. B. & Chain, P. S. Accurate read-based metagenome characterization using a hierarchical suite of unique signatures. Nucleic Acids Res. 43, e69 (2015).
Spanogiannopoulos, P., Bess, E. N., Carmody, R. N. & Turnbaugh, P. J. The microbial pharmacists within us: a metagenomic view of xenobiotic metabolism. Nat. Rev. Microbiol. 14, 273–287 (2016).
Gill, S. & Panda, S. A. Smartphone app reveals erratic diurnal eating patterns in humans that can be modulated for health benefits. Cell. Metab. 22, 789–798 (2015).
Kovatcheva-Datchary, P. et al. Dietary fiber-induced improvement in glucose metabolism is associated with increased abundance of Prevotella. Cell. Metab. 22, 971–982 (2015).
Zeevi, D. et al. Personalized nutrition by prediction of glycemic responses. Cell 163, 1079–1094 (2015). This study shows that the composition of the gut microbiota, integrated with other parameters, can be used to predict the blood glucose level of an individual after a certain meal and facilitate dietary interventions.
Olle, B. Medicines from microbiota. Nat. Biotechnol. 31, 309–315 (2013).
Ling, L. L. et al. A new antibiotic kills pathogens without detectable resistance. Nature 517, 455–459 (2015).
Wang, Z. et al. Non-lethal inhibition of gut microbial trimethylamine production for the treatment of atherosclerosis. Cell 163, 1585–1595 (2015). This study identifies a chemical analogue of choline that shows success in inhibiting the production of trimethylamine by the gut microbiota.
Acknowledgements
This study was supported by the Natural Science Foundation of China (grants 30890032, 30725008 and 30811130531), the Shenzhen Municipal Government of China (grants JSGG20140702161403250, DRC-SZ[2015]162 and CXB201108250098A), the Danish Strategic Research Council (grant 2106-07-0021) and the Ole RØmer grant from the Danish Natural Science Research Council and Solexa project (272-07-0196). The authors thank their colleagues at BGI, Shenzhen, China, especially J. Li, Z. Lan, S. Liang, H. Xie, D. Zhang, X. Luo, M. Arumugam and K. Kristiansen, for their help in the preparation of this Review. The authors also thank Y. Xie at Michigan State University, East Lansing, USA, for helpful discussions regarding this manuscript.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
J.W. is the CEO of iCarbonX.
Glossary
- Microbiome
-
The ensemble of microbial genomes and products at a given site.
- Microbiota
-
The ecological community of microorganisms at a given site.
- 16S rRNA gene amplicon sequencing
-
Amplification and sequencing of the variable regions in 16S ribosomal RNA genes for the taxonomic profiling of bacteria and archaea in a sample.
- Dysbiosis
-
An imbalance of the microbiota at a body site that is caused by an overgrowth of pathogenic microorganisms or a lack of commensal micoorganisms.
- Contigs
-
Contiguous DNA sequences that are assembled from shorter, overlapping sequencing reads.
- Short-chain fatty acids
-
(SCFAs). Fatty acids that have fewer than six carbon atoms. In the context of the microbiome, SCFAs usually refer to acetate, propionate and butyrate, which are produced by various species of bacteria.
- Metformin
-
A biguanide drug that is commonly prescribed as a treatment for type 2 diabetes.
- Supervised machine learning
-
Machine learning in which the training data are labelled (for example, as cases or controls). Using the training data, the algorithm learns to classify new data according to these labels.
- Area under the receiver operating characteristic curve
-
(AUC). The area under a receiver operating characteristic (ROC) curve of true-positive rates versus false-positive rates, which depicts the performance of a binary classifier. AUCs typically range between 0.5 and 1, corresponding to a random and a perfect classification, respectively.
- Dyslipidaemia
-
An abnormal amount of lipids in the blood.
- Adenomas
-
Benign tumours that are formed from glands or that have characteristics of glands.
- Periodontitis
-
Inflammation of the tissue that surrounds the teeth, which leads to the progressive loss of the alveolar bone and the loosening or loss of teeth.
- Rheumatoid factor
-
An autoantibody against the constant region (known as the fragment crystallisable (Fc) region) of immunoglobulin G.
- Anti-cyclic citrullinated peptide autoantibodies
-
Autoantibodies against proteins that contain the modified amino acid citrulline. Cyclic citrullinated peptides are used to clinically detect these antibodies.
- Guilt by association
-
A concept from genome-wide association studies (GWAS) that describes associations in such studies as 'guilt' of a gene for a trait of interest, which means that the gene is of interest for further investigation.
- Specific pathogen-free mice
-
Laboratory mice that are free of particular pathogens that could interfere with experiments. The excluded pathogens include both viral and bacterial pathogens.
- Organoids
-
Organ-like structures that are grown in the laboratory.
- Mobile genetic elements
-
DNA sequences that can be transferred between genomes or between loci of the same genome.
Rights and permissions
About this article
Cite this article
Wang, J., Jia, H. Metagenome-wide association studies: fine-mining the microbiome. Nat Rev Microbiol 14, 508–522 (2016). https://doi.org/10.1038/nrmicro.2016.83
Published:
Issue Date:
DOI: https://doi.org/10.1038/nrmicro.2016.83
This article is cited by
-
Challenges and opportunities in sharing microbiome data and analyses
Nature Microbiology (2023)
-
Removal of false positives in metagenomics-based taxonomy profiling via targeting Type IIB restriction sites
Nature Communications (2023)
-
Environmental Effects on Bee Microbiota
Microbial Ecology (2023)
-
Cultivation-independent genomes greatly expand taxonomic-profiling capabilities of mOTUs across various environments
Microbiome (2022)
-
Sequencing introduced false positive rare taxa lead to biased microbial community diversity, assembly, and interaction interpretation in amplicon studies
Environmental Microbiome (2022)