Abstract
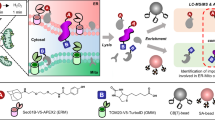
APEX is an engineered peroxidase that functions as an electron microscopy tag and a promiscuous labeling enzyme for live-cell proteomics. Because limited sensitivity precludes applications requiring low APEX expression, we used yeast-display evolution to improve its catalytic efficiency. APEX2 is far more active in cells, enabling the use of electron microscopy to resolve the submitochondrial localization of calcium uptake regulatory protein MICU1. APEX2 also permits superior enrichment of endogenous mitochondrial and endoplasmic reticulum membrane proteins.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Accession codes
Change history
10 December 2014
In the version of this article initially published online, the asterisks in Figure 2a were incorrectly depicted as exclamation marks, and some ordinate labels in the left graph of Figure 2e were obscured. The errors have been corrected for the print, PDF and HTML versions of this article.
References
Porstmann, B., Porstmann, T., Nugel, E. & Evers, U. J. Immunol. Methods 79, 27–37 (1985).
Li, J., Wang, Y., Chiu, S.-L. & Cline, H.T. Front. Neural Circuits 4, 6 (2010).
Hopkins, C., Gibson, A., Stinchcombe, J. & Futter, C. Methods Enzymol. 327, 35–45 (2000).
Martell, J.D. et al. Nat. Biotechnol. 30, 1143–1148 (2012).
Rhee, H.-W. et al. Science 339, 1328–1331 (2013).
Hung, V. et al. Mol. Cell 55, 332–341 (2014).
Mandelman, D., Schwarz, F.P., Li, H. & Poulos, T.L. Protein Sci. 7, 2089–2098 (1998).
Ebert, P.S., Hess, R.A., Frykholm, B.C. & Tschudy, D.P. Biochem. Biophys. Res. Commun. 88, 1382–1390 (1979).
Sharp, K.H., Moody, P.C.E., Brown, K.A. & Raven, E.L. Biochemistry 43, 8644–8651 (2004).
Nicell, J.A. & Wright, H. Enzyme Microb. Technol. 21, 302–310 (1997).
Arnao, M.B., Acosta, M. & del Río, J.A. & García-Cánovas, F. Biochim. Biophys. Acta 1038, 85–89 (1990).
Baughman, J.M. et al. Nature 476, 341–345 (2011).
De Stefani, D., Raffaello, A., Teardo, E., Szabò, I. & Rizzuto, R. Nature 476, 336–340 (2011).
Kamer, K.J. & Mootha, V.K. EMBO Rep. 15, 299–307 (2014).
Perocchi, F. et al. Nature 467, 291–296 (2010).
Csordás, G. et al. Cell Metab. 17, 976–987 (2013).
Hoffman, N.E. et al. Cell Rep. 5, 1576–1588 (2013).
Vander Heiden, M.G. et al. Proc. Natl. Acad. Sci. USA 97, 4666–4671 (2000).
Plovanich, M. et al. PLoS ONE 8, e55785 (2013).
Shu, X. et al. PLoS Biol. 9, e1001041 (2011).
Gaietta, G. et al. Science 296, 503–507 (2002).
Grabenbauer, M. et al. Nat. Methods 2, 857–862 (2005).
Horstmann, H., Vasileva, M. & Kuner, T. PLoS ONE 8, e64764 (2013).
Lad, L., Mewies, M. & Raven, E.L. Biochemistry 41, 13774–13781 (2002).
Hiner, A.N. et al. Biochem. J. 348, 321–328 (2000).
Mallilankaraman, K. et al. Cell 151, 630–644 (2012).
Pagliarini, D.J. et al. Cell 134, 112–123 (2008).
Sanjana, N.E. et al. Nat. Protoc. 7, 171–192 (2012).
Sancak, Y. et al. Cell 141, 290–303 (2010).
Wen, W., Meinkoth, J.L., Tsien, R.Y. & Taylor, S.S. Cell 82, 463–473 (1995).
Seth, R.B., Sun, L., Ea, C.-K. & Chen, Z.J. Cell 122, 669–682 (2005).
Szczesna-Skorupa, E. Proc. Natl. Acad. Sci. USA 95, 14793–14798 (1998).
Chao, G. et al. Nat. Protoc. 1, 755–768 (2006).
Colby, D.W. et al. Methods Enzymol. 388, 348–358 (2004).
Liu, D.S., Loh, K.H., Lam, S.S., White, K.A. & Ting, A.Y. PLoS ONE 8, e52823 (2013).
Richards, M.K. & Marletta, M.A. Biochemistry 33, 14723–14732 (1994).
Kery, V., Elleder, D. & Kraus, J.P. Arch. Biochem. Biophys. 316, 24–29 (1995).
Delcarte, J. et al. J. Chromatogr. B Analyt. Technol. Biomed. Life Sci. 786, 229–236 (2003).
Barrows, T.P. & Poulos, T.L. Biochemistry 44, 14062–14068 (2005).
Berry, E.A. & Trumpower, B.L. Anal. Biochem. 161, 1–15 (1987).
Noble, R.W. & Gibson, Q.H. J. Biol. Chem. 245, 2409–2413 (1970).
Acknowledgements
We acknowledge funding from the US National Institutes of Health (DP1 OD003961 to A.Y.T.; P41 GM103412, R01GM086197 to M.H.E.; 5R01GM077465-08 to V.K.M.), the Howard Hughes Medical Institute (V.K.M.), and the Howard Hughes Medical Institute Collaborative Initiative Award (A.Y.T.). S.S.L. and J.D.M. were supported by US National Science Foundation Graduate Research Fellowships and National Defense Science and Engineering Graduate Fellowships. N. Watson acquired EM images of MICU1-APEX2. V. Hung provided APEX2-Stx17 and ATP5J-APEX2 EM images. K. Cox (Massachusetts Institute of Technology) provided some plasmids. FACS experiments were performed at the Koch Institute Swanson Biotechnology Center Flow Cytometry Facility. Color bright-field imaging was performed at the Koch Institute Microcopy Core Facility. We thank C. Drennan for use of her Cary300 spectrophotometer. D. McSwiggen assisted with enzyme purification. We thank T. Poulos, K. White, and the laboratory of D. Wittrup for advice.
Author information
Authors and Affiliations
Contributions
S.S.L., J.D.M. and A.Y.T. designed the research, analyzed the data and wrote the paper. All authors edited the paper. K.J.K. and V.K.M. prepared MICU1 stable cells and performed calcium uptake assays. S.S.L. and J.D.M. performed EM sample preparation, and T.J.D. and M.H.E. performed EM imaging. J.D.M. performed enzyme kinetic assays and analysis. S.S.L. performed all other experiments.
Corresponding author
Ethics declarations
Competing interests
The Massachusetts Institute of Technology has submitted a patent application on the peroxidase technology. J.D.M. and A.Y.T. are authors on this patent.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–16, Supplementary Tables 1 and 2 and Supplementary Discussion (PDF 5389 kb)
Rights and permissions
About this article
Cite this article
Lam, S., Martell, J., Kamer, K. et al. Directed evolution of APEX2 for electron microscopy and proximity labeling. Nat Methods 12, 51–54 (2015). https://doi.org/10.1038/nmeth.3179
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth.3179
This article is cited by
-
Profiling phagosome proteins identifies PD-L1 as a fungal-binding receptor
Nature (2024)
-
The rapid proximity labeling system PhastID identifies ATP6AP1 as an unconventional GEF for Rheb
Cell Research (2024)
-
Mitochondrial proteome research: the road ahead
Nature Reviews Molecular Cell Biology (2024)
-
A platform to induce and mature biomolecular condensates using chemicals and light
Nature Chemical Biology (2024)
-
Using the heme peroxidase APEX2 to probe intracellular H2O2 flux and diffusion
Nature Communications (2024)