Abstract
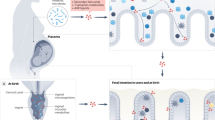
Recent studies have characterized how host genetics, prenatal environment and delivery mode can shape the newborn microbiome at birth. Following this, postnatal factors, such as antibiotic treatment, diet or environmental exposure, further modulate the development of the infant's microbiome and immune system, and exposure to a variety of microbial organisms during early life has long been hypothesized to exert a protective effect in the newborn. Furthermore, epidemiological studies have shown that factors that alter bacterial communities in infants during childhood increase the risk for several diseases, highlighting the importance of understanding early-life microbiome composition. In this review, we describe how prenatal and postnatal factors shape the development of both the microbiome and the immune system. We also discuss the prospects of microbiome-mediated therapeutics and the need for more effective approaches that can reconfigure bacterial communities from pathogenic to homeostatic configurations.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout

Marina Corral Spence/Nature Publishing Group

Marina Corral Spence/Nature Publishing Group

Marina Corral Spence/Nature Publishing Group
Similar content being viewed by others
References
Satokari, R., Grönroos, T., Laitinen, K., Salminen, S. & Isolauri, E. Bifidobacterium and Lactobacillus DNA in the human placenta. Lett. Appl. Microbiol. 48, 8–12 (2009).
Aagaard, K. et al. The placenta harbors a unique microbiome. Sci. Transl. Med. 6, 237ra265 (2014).
Oh, K.J. et al. Detection of ureaplasmas by the polymerase chain reaction in the amniotic fluid of patients with cervical insufficiency. J. Perinat. Med. 38, 261–268 (2010).
DiGiulio, D.B. et al. Microbial prevalence, diversity and abundance in amniotic fluid during preterm labor: a molecular and culture-based investigation. PLoS One 3, e3056 (2008).
Steel, J.H. et al. Bacteria and inflammatory cells in fetal membranes do not always cause preterm labor. Pediatr. Res. 57, 404–411 (2005).
Jiménez, E. et al. Isolation of commensal bacteria from umbilical cord blood of healthy neonates born by cesarean section. Curr. Microbiol. 51, 270–274 (2005).
Jiménez, E. et al. Is meconium from healthy newborns actually sterile? Res. Microbiol. 159, 187–193 (2008).
Hu, J. et al. Diversified microbiota of meconium is affected by maternal diabetes status. PLoS One 8, e78257 (2013).
Romero, R., Dey, S.K. & Fisher, S.J. Preterm labor: one syndrome, many causes. Science 345, 760–765 (2014).
Goldenberg, R.L., Culhane, J.F., Iams, J.D. & Romero, R. Epidemiology and causes of preterm birth. Lancet 371, 75–84 (2008).
Kliman, H.J. Comment on “the placenta harbors a unique microbiome”. Sci. Transl. Med. 6, 254le4 (2014).
Boggess, K.A. et al. Bacteremia shortly after placental separation during cesarean delivery. Obstet. Gynecol. 87, 779–784 (1996).
Salter, S.J. et al. Reagent and laboratory contamination can critically impact sequence-based microbiome analyses. BMC Biol. 12, 87 (2014).
Durack, D.T. Prevention of infective endocarditis. N. Engl. J. Med. 332, 38–44 (1995).
Palmer, C., Bik, E.M., DiGiulio, D.B., Relman, D.A. & Brown, P.O. Development of the human infant intestinal microbiota. PLoS Biol. 5, e177 (2007).
Bennet, R. & Nord, C.E. Development of the fecal anaerobic microflora after cesarean section and treatment with antibiotics in newborn infants. Infection 15, 332–336 (1987).
Dominguez-Bello, M.G. et al. Delivery mode shapes the acquisition and structure of the initial microbiota across multiple body habitats in newborns. Proc. Natl. Acad. Sci. USA 107, 11971–11975 (2010).
Bäckhed, F. et al. Dynamics and stabilization of the human gut microbiome during the first year of life. Cell Host Microbe 17, 690–703 (2015).
Dominguez-Bello, M.G. et al. Partial restoration of the microbiota of cesarean-born infants via vaginal microbial transfer. Nat. Med. 22, 250–253 (2016).
Koenig, J.E. et al. Succession of microbial consortia in the developing infant gut microbiome. Proc. Natl. Acad. Sci. USA 108 Suppl 1, 4578–4585 (2011).
Lim, E.S. et al. Early-life dynamics of the human gut virome and bacterial microbiome in infants. Nat. Med. 21, 1228–1234 (2015).
Yatsunenko, T. et al. Human gut microbiome viewed across age and geography. Nature 486, 222–227 (2012).
Jakobsson, H.E. et al. Decreased gut microbiota diversity, delayed Bacteroidetes colonization and reduced TH1 responses in infants delivered by cesarean section. Gut 63, 559–566 (2014).
Kabeerdoss, J. et al. Development of the gut microbiota in southern Indian infants from birth to 6 months: a molecular analysis. J. Nutr. Sci. 2, e18 (2013).
Sevelsted, A., Stokholm, J., Bønnelykke, K. & Bisgaard, H. Cesarean section and chronic immune disorders. Pediatrics 135, e92–e98 (2015).
Huh, S.Y. et al. Delivery by cesarean section and risk of obesity in preschool age children: a prospective cohort study. Arch. Dis. Child. 97, 610–616 (2012).
Eggesbø, M., Botten, G., Stigum, H., Nafstad, P. & Magnus, P. Is delivery by cesarean section a risk factor for food allergy? J. Allergy Clin. Immunol. 112, 420–426 (2003).
Zhou, X. et al. The vaginal bacterial communities of Japanese women resemble those of women in other racial groups. FEMS Immunol. Med. Microbiol. 58, 169–181 (2010).
Gajer, P. et al. Temporal dynamics of the human vaginal microbiota. Sci. Transl. Med. 4, 132ra52 (2012).
Ravel, J. et al. Vaginal microbiome of reproductive-age women. Proc. Natl. Acad. Sci. USA 108 Suppl 1, 4680–4687 (2011).
Romero, R. et al. The composition and stability of the vaginal microbiota of normal pregnant women is different from that of nonpregnant women. Microbiome 2, 4 (2014).
Walther-António, M.R. et al. Pregnancy's stronghold on the vaginal microbiome. PLoS One 9, e98514 (2014).
Goplerud, C.P., Ohm, M.J. & Galask, R.P. Aerobic and anaerobic flora of the cervix during pregnancy and the puerperium. Am. J. Obstet. Gynecol. 126, 858–868 (1976).
Vásquez, A., Jakobsson, T., Ahrné, S., Forsum, U. & Molin, G. Vaginal lactobacillus flora of healthy Swedish women. J. Clin. Microbiol. 40, 2746–2749 (2002).
Redondo-Lopez, V., Cook, R.L. & Sobel, J.D. Emerging role of lactobacilli in the control and maintenance of the vaginal bacterial microflora. Rev. Infect. Dis. 12, 856–872 (1990).
Witkin, S.S. & Ledger, W.J. Complexities of the uniquely human vagina. Sci. Transl. Med. 4, 132fs11 (2012).
MacIntyre, D.A. et al. The vaginal microbiome during pregnancy and the postpartum period in a European population. Sci. Rep. 5, 8988 (2015).
Fettweis, J.M. et al. Differences in vaginal microbiome in African-American women versus women of European ancestry. Microbiology 160, 2272–2282 (2014).
Hicks, L.A. et al. US outpatient antibiotic-prescribing variation according to geography, patient population and provider specialty in 2011. Clin. Infect. Dis. 60, 1308–1316 (2015).
Hersh, A.L., Shapiro, D.J., Pavia, A.T. & Shah, S.S. Antibiotic prescribing in ambulatory pediatrics in the United States. Pediatrics 128, 1053–1061 (2011).
Arrieta, M.C., Stiemsma, L.T., Amenyogbe, N., Brown, E.M. & Finlay, B. The intestinal microbiome in early life: health and disease. Front. Immunol. 5, 427 (2014).
Kozyrskyj, A.L., Ernst, P. & Becker, A.B. Increased risk of childhood asthma from antibiotic use in early life. Chest 131, 1753–1759 (2007).
Risnes, K.R., Belanger, K., Murk, W. & Bracken, M.B. Antibiotic exposure by 6 months, and asthma and allergy at 6 years: findings in a cohort of 1,401 US children. Am. J. Epidemiol. 173, 310–318 (2011).
Hoskin-Parr, L., Teyhan, A., Blocker, A. & Henderson, A.J. Antibiotic exposure in the first 2 years of life and development of asthma and other allergic diseases by 7.5 years: a dose-dependent relationship. Pediatr. Allergy Immunol. 24, 762–771 (2013).
Bailey, L.C. et al. Association of antibiotics in infancy with early childhood obesity. JAMA Pediatr. 168, 1063–1069 (2014).
Mikkelsen, K.H., Knop, F.K., Frost, M., Hallas, J. & Pottegård, A. Use of antibiotics and risk of type 2 diabetes: a population-based case-control study. J. Clin. Endocrinol. Metab. 100, 3633–3640 (2015).
Metsälä, J. et al. Mother's and offspring's use of antibiotics, and infant allergy to cow's milk. Epidemiology 24, 303–309 (2013).
Kronman, M.P., Zaoutis, T.E., Haynes, K., Feng, R. & Coffin, S.E. Antibiotic exposure and IBD development among children: a population-based cohort study. Pediatrics 130, e794–e803 (2012).
Örtqvist, A.K. et al. Antibiotics in fetal and early life, and subsequent childhood asthma: nationwide population-based study with sibling analysis. Br. Med. J. 349, g6979 (2014).
Semic-Jusufagic, A. et al. Assessing the association of early-life antibiotic prescription with asthma exacerbations, impaired antiviral immunity and genetic variants in 17q21: a population-based birth-cohort study. Lancet Respir. Med. 2, 621–630 (2014).
Zaura, E. et al. Same exposure but two radically different responses to antibiotics: resilience of the salivary microbiome versus long-term microbial shifts in feces. MBio 6, e01693–e15 (2015).
Langdon, A., Crook, N. & Dantas, G. The effects of antibiotics on the microbiome throughout development and alternative approaches for therapeutic modulation. Genome Med. 8, 39 (2016).
Cox, L.M. et al. Altering the intestinal microbiota during a critical developmental window has lasting metabolic consequences. Cell 158, 705–721 (2014).
Nobel, Y.R. et al. Metabolic and metagenomic outcomes from early-life-pulsed antibiotic treatment. Nat. Commun. 6, 7486 (2015).
Ivanov, I.I. et al. Induction of intestinal TH17 cells by segmented filamentous bacteria. Cell 139, 485–498 (2009).
Stefka, A.T. et al. Commensal bacteria protect against food-allergen sensitization. Proc. Natl. Acad. Sci. USA 111, 13145–13150 (2014).
Noverr, M.C., Falkowski, N.R., McDonald, R.A., McKenzie, A.N. & Huffnagle, G.B. Development of allergic airway disease in mice following antibiotic therapy and fungal microbiota increase: role of host genetics, antigen and interleukin-13. Infect. Immun. 73, 30–38 (2005).
Gonzalez-Perez, G. et al. Maternal antibiotic treatment impacts development of the neonatal intestinal microbiome and antiviral immunity. J. Immunol. 196, 3768–3779 (2016).
Beura, L.K. et al. Normalizing the environment recapitulates adult human immune traits in laboratory mice. Nature 532, 512–516 (2016).
Sadeharju, K. et al. Maternal antibodies in breast milk protect the child from enterovirus infections. Pediatrics 119, 941–946 (2007).
WHO Collaborative Study Team on the Role of Breast-feeding on the Prevention of Infant Mortality. Effect of breast-feeding on infant and child mortality due to infectious diseases in less developed countries: a pooled analysis. Lancet 355, 451–455 (2000).
Harder, T., Bergmann, R., Kallischnigg, G. & Plagemann, A. Duration of breast-feeding and risk of overweight: a meta-analysis. Am. J. Epidemiol. 162, 397–403 (2005).
Weng, S.F., Redsell, S.A., Swift, J.A., Yang, M. & Glazebrook, C.P. Systematic review and meta-analyses of risk factors for childhood overweight identifiable during infancy. Arch. Dis. Child. 97, 1019–1026 (2012).
Greer, F.R., Sicherer, S.H. & Burks, A.W. Effects of early nutritional interventions on the development of atopic disease in infants and children: the role of maternal dietary restriction, breast-feeding, timing of introduction of complementary foods and hydrolyzed formulas. Pediatrics 121, 183–191 (2008).
Schwarz, E.B. et al. Duration of lactation and risk factors for maternal cardiovascular disease. Obstet. Gynecol. 113, 974–982 (2009).
Stuebe, A.M., Rich-Edwards, J.W., Willett, W.C., Manson, J.E. & Michels, K.B. Duration of lactation and incidence of type 2 diabetes. J. Am. Med. Assoc. 294, 2601–2610 (2005).
Guilbert, T.W., Stern, D.A., Morgan, W.J., Martinez, F.D. & Wright, A.L. Effect of breast-feeding on lung function in childhood, and modulation by maternal asthma and atopy. Am. J. Respir. Crit. Care Med. 176, 843–848 (2007).
Dogaru, C.M. et al. Breast-feeding and lung function at school age: does maternal asthma modify the effect? Am. J. Respir. Crit. Care Med. 185, 874–880 (2012).
Brandtzaeg, P. The mucosal immune system and its integration with the mammary glands. J. Pediatr. 156 Suppl, S8–S15 (2010).
Lönnerdal, B. Nutritional and physiologic significance of human milk proteins. Am. J. Clin. Nutr. 77, 1537S–1543S (2003).
Yoshioka, H., Iseki, K. & Fujita, K. Development and differences of intestinal flora in the neonatal period in breast-fed and bottle-fed infants. Pediatrics 72, 317–321 (1983).
Engfer, M.B., Stahl, B., Finke, B., Sawatzki, G. & Daniel, H. Human milk oligosaccharides are resistant to enzymatic hydrolysis in the upper gastrointestinal tract. Am. J. Clin. Nutr. 71, 1589–1596 (2000).
Martín, R. et al. Human milk is a source of lactic acid bacteria for the infant gut. J. Pediatr. 143, 754–758 (2003).
Heikkilä, M.P. & Saris, P.E. Inhibition of Staphylococcus aureus by the commensal bacteria of human milk. J. Appl. Microbiol. 95, 471–478 (2003).
Pantoja-Feliciano, I.G. et al. Biphasic assembly of the murine intestinal microbiota during early development. ISME J. 7, 1112–1115 (2013).
Braegger, C. et al. Supplementation of infant formula with probiotics and/or prebiotics: a systematic review and comment by the ESPGHAN committee on nutrition. J. Pediatr. Gastroenterol. Nutr. 52, 238–250 (2011).
Bergmann, H., Rodríguez, J.M., Salminen, S. & Szajewska, H. Probiotics in human milk and probiotic supplementation in infant nutrition: a workshop report. Br. J. Nutr. 112, 1119–1128 (2014).
Deshpande, G., Rao, S. & Patole, S. Probiotics in neonatal intensive care—back to the future. Aust. N. Z. J. Obstet. Gynaecol. 55, 210–217 (2015).
Cuello-Garcia, C.A. et al. Probiotics for the prevention of allergy: a systematic review and meta-analysis of randomized controlled trials. J. Allergy Clin. Immunol. 136, 952–961 (2015).
Panduru, M., Panduru, N.M., Sălăvăstru, C.M. & Tiplica, G.S. Probiotics and primary prevention of atopic dermatitis: a meta-analysis of randomized controlled studies. J. Eur. Acad. Dermatol. Venereol. 29, 232–242 (2015).
Foolad, N., Brezinski, E.A., Chase, E.P. & Armstrong, A.W. Effect of nutrient supplementation on atopic dermatitis in children: a systematic review of probiotics, prebiotics, formula and fatty acids. JAMA Dermatol. 149, 350–355 (2013).
Doege, K. et al. Impact of maternal supplementation with probiotics during pregnancy on atopic eczema in childhood—a meta-analysis. Br. J. Nutr. 107, 1–6 (2012).
Kim, S.-O. et al. Effects of probiotics for the treatment of atopic dermatitis: a meta-analysis of randomized controlled trials. Ann. Allergy Asthma Immunol. 113, 217–226 (2014).
Pelucchi, C. et al. Probiotics supplementation during pregnancy or infancy for the prevention of atopic dermatitis: a meta-analysis. Epidemiology 23, 402–414 (2012).
Elazab, N. et al. Probiotic administration in early life, atopy and asthma: a meta-analysis of clinical trials. Pediatrics 132, e666–e676 (2013).
Savino, F. et al. Lactobacillus reuteri DSM 17938 in infantile colic: a randomized, double-blind, placebo-controlled trial. Pediatrics 126, e526–e533 (2010).
Brunser, O. et al. Effects of probiotic- or prebiotic-supplemented milk formulas on fecal microbiota composition of infants. Asia Pac. J. Clin. Nutr. 15, 368–376 (2006).
Mah, K.W. et al. Effect of a milk formula containing probiotics on the fecal microbiota of Asian infants at risk of atopic diseases. Pediatr. Res. 62, 674–679 (2007).
Maldonado, J. et al. Human milk probiotic Lactobacillus fermentum CECT5716 reduces the incidence of gastrointestinal and upper respiratory tract infections in infants. J. Pediatr. Gastroenterol. Nutr. 54, 55–61 (2012).
Vandenplas, Y., Zakharova, I. & Dmitrieva, Y. Oligosaccharides in infant formula: more evidence to validate the role of prebiotics. Br. J. Nutr. 113, 1339–1344 (2015).
Krebs, N.F. et al. Effects of different complementary feeding regimens on iron status and enteric microbiota in breast-fed infants. J. Pediatr. 163, 416–423 (2013).
Andersen, A.D., Mølbak, L., Michaelsen, K.F. & Lauritzen, L. Molecular fingerprints of the human fecal microbiota from 9 to 18 months old and the effect of fish oil supplementation. J. Pediatr. Gastroenterol. Nutr. 53, 303–309 (2011).
Zimmermann, M.B. et al. The effects of iron fortification on the gut microbiota in African children: a randomized controlled trial in Cote d'Ivoire. Am. J. Clin. Nutr. 92, 1406–1415 (2010).
Clemente, J.C. et al. The microbiome of uncontacted Amerindians. Sci. Adv. 1, e1500183 (2015).
Flores, G.E. et al. Temporal variability is a personalized feature of the human microbiome. Genome Biol. 15, 531 (2014).
Hesselmar, B. et al. Pacifier cleaning practices and risk of allergy development. Pediatrics 131, e1829–e1837 (2013).
Song, S.J. et al. Cohabiting family members share microbiota with one another and with their dogs. eLife 2, e00458 (2013).
Faith, J.J. et al. The long-term stability of the human gut microbiota. Science 341, 1237439 (2013).
Virtanen, S.M. et al. Microbial exposure in infancy and subsequent appearance of type 1 diabetes mellitus–associated autoantibodies: a cohort study. JAMA Pediatr. 168, 755–763 (2014).
Ownby, D.R., Johnson, C.C. & Peterson, E.L. Exposure to dogs and cats in the first year of life and risk of allergic sensitization at 6 to 7 years of age. J. Am. Med. Assoc. 288, 963–972 (2002).
Fujimura, K.E. et al. House dust exposure mediates gut microbiome Lactobacillus enrichment and airway immune defense against allergens and virus infection. Proc. Natl. Acad. Sci. USA 111, 805–810 (2014).
Sjögren, Y.M., Jenmalm, M.C., Böttcher, M.F., Björkstén, B. & Sverremark-Ekström, E. Altered early infant gut microbiota in children developing allergy up to 5 years of age. Clin. Exp. Allergy 39, 518–526 (2009).
Haahtela, T. et al. Hunt for the origin of allergy—comparing the Finnish and Russian Karelia. Clin. Exp. Allergy 45, 891–901 (2015).
von Hertzen, L. et al. Microbial content of drinking water in Finnish and Russian Karelia—implications for atopy prevalence. Allergy 62, 288–292 (2007).
Normand, A.C. et al. Airborne cultivable microflora and microbial transfer in farm buildings and rural dwellings. Occup. Environ. Med. 68, 849–855 (2011).
Frayling, T.M. et al. A common variant in the FTO gene is associated with body mass index, and predisposes to childhood and adult obesity. Science 316, 889–894 (2007).
Herbert, A. et al. A common genetic variant is associated with adult and childhood obesity. Science 312, 279–283 (2006).
Turnbaugh, P.J. et al. A core gut microbiome in obese and lean twins. Nature 457, 480–484 (2009).
Qin, J. et al. A metagenome-wide association study of gut microbiota in type 2 diabetes. Nature 490, 55–60 (2012).
Karlsson, F.H. et al. Gut metagenome in European women with normal, impaired and diabetic glucose control. Nature 498, 99–103 (2013).
Knights, D. et al. Complex host genetics influence the microbiome in inflammatory bowel disease. Genome Med. 6, 107 (2014).
Goodrich, J.K. et al. Human genetics shape the gut microbiome. Cell 159, 789–799 (2014).
Chervonsky, A.V. Influence of microbial environment on autoimmunity. Nat. Immunol. 11, 28–35 (2010).
Hooper, L.V., Littman, D.R. & Macpherson, A.J. Interactions between the microbiota and the immune system. Science 336, 1268–1273 (2012).
Ajslev, T.A., Andersen, C.S., Gamborg, M., Sørensen, T.I. & Jess, T. Childhood overweight after establishment of the gut microbiota: the role of delivery mode, prepregnancy weight and early administration of antibiotics. Int. J. Obes. 35, 522–529 (2011).
Jostins, L. et al. Host–microbe interactions have shaped the genetic architecture of inflammatory bowel disease. Nature 491, 119–124 (2012).
Nestle, F.O., Kaplan, D.H. & Barker, J. Psoriasis. N. Engl. J. Med. 361, 496–509 (2009).
Adkins, B. & Du, R.Q. Newborn mice develop balanced TH1 and TH2 primary effector responses in vivo but are biased to TH2 secondary responses. J. Immunol. 160, 4217–4224 (1998).
Bach, J.F. The effect of infections on susceptibility to autoimmune and allergic diseases. N. Engl. J. Med. 347, 911–920 (2002).
Liu, A.H. & Leung, D.Y. Renaissance of the hygiene hypothesis. J. Allergy Clin. Immunol. 117, 1063–1066 (2006).
Lotz, M. et al. Postnatal acquisition of endotoxin tolerance in intestinal epithelial cells. J. Exp. Med. 203, 973–984 (2006).
Vatanen, T. et al. Variation in microbiome LPS immunogenicity contributes to autoimmunity in humans. Cell 165, 842–853 (2016).
An, D. et al. Sphingolipids from a symbiotic microbe regulate homeostasis of host intestinal natural killer T cells. Cell 156, 123–133 (2014).
Gevers, D. et al. The treatment-naive microbiome in new-onset Crohn's disease. Cell Host Microbe 15, 382–392 (2014).
Arrieta, M.C. et al. Early-infancy microbial and metabolic alterations affect risk of childhood asthma. Sci. Transl. Med. 7, 307ra152 (2015).
Langille, M.G. et al. Predictive functional profiling of microbial communities using 16S rRNA marker gene sequences. Nat. Biotechnol. 31, 814–821 (2013).
Bunyavanich, S. et al. Early-life gut microbiome composition is associated with milk allergy resolution. J. Allergy Clin. Immunol. http://dx.doi.org/10.1016/j.jaci.2016.03.041 (2016).
Smith, M.I. et al. Gut microbiomes of Malawian twin pairs discordant for kwashiorkor. Science 339, 548–554 (2013).
Noval Rivas, M. et al. A microbiota signature associated with experimental food allergy promotes allergic sensitization and anaphylaxis. J. Allergy Clin. Immunol. 131, 201–212 (2013).
Muniyappa, P., Gulati, R., Mohr, F. & Hupertz, V. Use and safety of rifaximin in children with inflammatory bowel disease. J. Pediatr. Gastroenterol. Nutr. 49, 400–404 (2009).
Turner, D., Levine, A., Kolho, K.L., Shaoul, R. & Ledder, O. Combination of oral antibiotics may be effective in severe pediatric ulcerative colitis: a preliminary report. J. Crohn's Colitis 8, 1464–1470 (2014).
Cotten, C.M. et al.; NICHD Neonatal Research Network. Prolonged duration of initial empirical antibiotic treatment is associated with increased rates of necrotizing enterocolitis and death for extremely low-birth-weight infants. Pediatrics 123, 58–66 (2009).
Faden, D. & Faden, H.S. The high rate of adverse drug events in children receiving prolonged outpatient parenteral antibiotic therapy for osteomyelitis. Pediatr. Infect. Dis. J. 28, 539–541 (2009).
Buchanan, E. et al. The use of exclusive enteral nutrition for induction of remission in children with Crohn's disease demonstrates that disease phenotype does not influence clinical remission. Aliment. Pharmacol. Ther. 30, 501–507 (2009).
Quince, C. et al. Extensive modulation of the fecal metagenome in children with Crohn's disease during exclusive enteral nutrition. Am. J. Gastroenterol. 110, 1718–1729, quiz 1730 (2015).
Khoruts, A., Dicksved, J., Jansson, J.K. & Sadowsky, M.J. Changes in the composition of the human fecal microbiome after bacteriotherapy for recurrent Clostridium difficile–associated diarrhea. J. Clin. Gastroenterol. 44, 354–360 (2010).
Youngster, I. et al. Oral, capsulized, frozen fecal microbiota transplantation for relapsing Clostridium difficile infection. J. Am. Med. Assoc. 312, 1772–1778 (2014).
Kunde, S. et al. Safety, tolerability and clinical response after fecal transplantation in children and young adults with ulcerative colitis. J. Pediatr. Gastroenterol. Nutr. 56, 597–601 (2013).
Grinspan, A.M. & Kelly, C.R. Fecal microbiota transplantation for ulcerative colitis: not just yet. Gastroenterology 149, 15–18 (2015).
Martinez, F.D. The human microbiome. Early-life determinant of health outcomes. Ann. Am. Thorac. Soc. 11 Suppl 1, S7–S12 (2014).
Furusawa, Y. et al. Commensal-microbe-derived butyrate induces the differentiation of colonic regulatory T cells. Nature 504, 446–450 (2013).
Sander, L.E. et al. Detection of prokaryotic mRNA signifies microbial viability and promotes immunity. Nature 474, 385–389 (2011).
Franzosa, E.A. et al. Relating the metatranscriptome and metagenome of the human gut. Proc. Natl. Acad. Sci. USA 111, E2329–E2338 (2014).
Sellitto, M. et al. Proof of concept of microbiome–metabolome analysis and delayed gluten exposure on celiac disease autoimmunity in genetically at-risk infants. PLoS One 7, e33387 (2012).
Stewart, C.J. et al. Preterm gut microbiota and metabolome following discharge from intensive care. Sci. Rep. 5, 17141 (2015).
Kersulyte, D. et al. Differences in genotypes of Helicobacter pylori from different human populations. J. Bacteriol. 182, 3210–3218 (2000).
Palm, N.W. et al. Immunoglobulin A coating identifies colitogenic bacteria in inflammatory bowel disease. Cell 158, 1000–1010 (2014).
Shen, N. & Clemente, J.C. Engineering the microbiome: a novel approach to immunotherapy for allergic and immune diseases. Curr. Allergy Asthma Rep. 15, 39 (2015).
David, L.A. et al. Diet rapidly and reproducibly alters the human gut microbiome. Nature 505, 559–563 (2014).
Saari, A., Virta, L.J., Sankilampi, U., Dunkel, L. & Saxen, H. Antibiotic exposure in infancy and risk of being overweight in the first 24 months of life. Pediatrics 135, 617–626 (2015).
Schwartz, B.S. et al. Antibiotic use and childhood body mass index trajectory. Int. J. Obes. (Lond) 40, 615–621 (2016).
Human Microbiome Project Consortium. Structure, function and diversity of the healthy human microbiome. Nature 486, 207–214 (2012).
Luo, C. et al. ConStrains identifies microbial strains in metagenomic data sets. Nat. Biotechnol. 33, 1045–1052 (2015).
Cleary, B. et al. Detection of low-abundance bacterial strains in metagenomic data sets by eigengenome partitioning. Nat. Biotechnol. 33, 1053–1060 (2015).
Maurice, C.F., Haiser, H.J. & Turnbaugh, P.J. Xenobiotics shape the physiology and gene expression of the active human gut microbiome. Cell 152, 39–50 (2013).
Peris-Bondia, F., Latorre, A., Artacho, A., Moya, A. & D'Auria, G. The active human gut microbiota differs from the total microbiota. PLoS One 6, e22448 (2011).
Ghannoum, M.A. et al. Characterization of the oral fungal microbiome (mycobiome) in healthy individuals. PLoS Pathog. 6, e1000713 (2010).
Gaitanis, G., Magiatis, P., Hantschke, M., Bassukas, I.D. & Velegraki, A. The Malassezia genus in skin and systemic diseases. Clin. Microbiol. Rev. 25, 106–141 (2012).
Chang, F.Y. et al. Analysis of the serum levels of fungi-specific immunoglobulin E in patients with allergic diseases. Int. Arch. Allergy Immunol. 154, 49–56 (2011).
Acknowledgements
J.C.C. was supported by the SUCCESS philanthropic grant (GCO14-0560) and by the Crohn's and Colitis Foundation of America (CCFA) (362048). We thank J.-F. Colombel for suggestions on Figure 3.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Tamburini, S., Shen, N., Wu, H. et al. The microbiome in early life: implications for health outcomes. Nat Med 22, 713–722 (2016). https://doi.org/10.1038/nm.4142
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nm.4142
This article is cited by
-
Can the Regime Type (Democracy versus Autocracy) Explain the COVID-19 Recovery Rates?
Chinese Political Science Review (2024)
-
Influence of gut microbiome on health and development of penaeid shrimps
Aquatic Sciences (2024)
-
Comparative Analysis of Outpatient Antibiotic Prescribing in Early Life: A Population-Based Study Across Birth Cohorts in Denmark and Germany
Infectious Diseases and Therapy (2024)
-
Development of the gut microbiota during early life in premature and term infants
Gut Pathogens (2023)
-
Dietary supplementation of laminarin improves the reproductive performance of sows and the growth of suckling piglets
Journal of Animal Science and Biotechnology (2023)