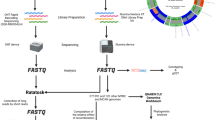
Abstract
Global spread and limited genetic variation are hallmarks of M. tuberculosis, the agent of human tuberculosis. In contrast, Mycobacterium canettii and related tubercle bacilli that also cause human tuberculosis and exhibit unusual smooth colony morphology are restricted to East Africa. Here, we sequenced and analyzed the whole genomes of five representative strains of smooth tubercle bacilli (STB) using Sanger (4–5× coverage), 454/Roche (13–18× coverage) and/or Illumina DNA sequencing (45–105× coverage). We show that STB isolates are highly recombinogenic and evolutionarily early branching, with larger genome sizes, higher rates of genetic variation, fewer molecular scars and distinct CRISPR-Cas systems relative to M. tuberculosis. Despite the differences, all tuberculosis-causing mycobacteria share a highly conserved core genome. Mouse infection experiments showed that STB strains are less persistent and virulent than M. tuberculosis. We conclude that M. tuberculosis emerged from an ancestral STB-like pool of mycobacteria by gain of persistence and virulence mechanisms, and we provide insights into the molecular events involved.
Similar content being viewed by others
Main
M. tuberculosis is a pervasive human pathogen currently estimated to infect 2 billion people throughout the world1. The bacterial population resulting from this massive spread is very large, yet the genetic diversity within the classical members of the M. tuberculosis complex (MTBC), comprising Mycobacterium africanum, Mycobacterium bovis, Mycobacterium microti, Mycobacterium pinnipedii and M. tuberculosis is very limited. Tuberculosis is therefore assumed to be a recent human disease2,3 linked to clonal expansion of its causative organism4,5,6.
In contrast to MTBC strains, STB clinical isolates, which have a distinctive smooth colony phenotype on culture media, named M. canettii and/or Mycobacterium prototuberculosis7,8,9,10, are less genetically restricted. Initial genotyping analysis suggested that these isolates possess higher diversity with traces of intraspecies horizontal gene transfer and might therefore represent early-branching lineages of tuberculosis-causing mycobacteria. Since their first isolation by Georges Canetti in 1969, less than 100 STB strains have been identified. All have been obtained from humans with tuberculosis, mostly from (or with connection to) East Africa8,11. Thus, a collection of a few tens of STB strains from a geographically restricted region seems to contain greater genetic diversity than the worldwide population of MTBC strains. This observation raises intriguing questions about the origin of tuberculosis and provided an opportunity to examine the molecular and evolutionary events involved in the emergence of M. tuberculosis. Here, we report the whole-genome sequence analysis of five diverse STB isolates and compare the physiopathological properties of these mycobacteria to those of M. tuberculosis, as well as the whole-genome shotgun sequences of four additional STB strains for secondary screening and confirmation purposes.
Results
Ancestral features of STB genomes
We applied multilocus sequence typing (MLST) based on 12 housekeeping genes to a panel of 55 available STB isolates and identified a total of 13 sequence types (Fig. 1 and Supplementary Tables 1 and 2). From analyses of the concatenated sequences, we inferred a highly reticulated phylogeny, suggestive of conflicting phylogenetic signals and possible horizontal gene transfer between the MLST target genes. We then selected five representative isolates for comprehensive genomic analysis. This selection included the original strain isolated by George Canetti of sequence type A (STB-A) and an isolate from the most prevalent group of sequence type D (STB-D)9, as well as strains from the most distant sequence types STB-L, STB-J and STB-K (Fig. 1 and Supplementary Fig. 1).
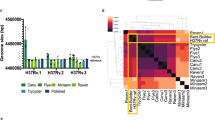
(a) Multilocus sequence typing of 56 STB and 10 MTBC reference isolates. Phylogenetic positions based on the split decomposition analysis of the concatenated sequences of 12 housekeeping gene segments are represented. The scale bar represents Hamming distance. Numbers indicate the percent of bootstrap support for the splits obtained after 1,000 replicates. Arrows and stars indicate isolates selected for complete genome sequence and genome shotgun analyses, respectively. (b) Pairwise linear genomic comparisons of M. tuberculosis H37Rv, M. bovis AF2122/97, five selected STB strains and two non-tuberculous mycobacterial species, M. marinum M and M. smegmatis MC2155. Red and blue lines indicate collinear blocks of DNA-DNA similarity and inverted matches, respectively. (c) Numbers of SNPs in pairwise comparisons between the indicated genomes. (d) Network phylogeny inferred among the five STB isolates subjected to complete genome sequence analysis and six selected MTBC (M. tuberculosis and M. africanum) genome sequences by NeighborNet analysis, based on pairwise alignments of whole-genome SNP data. An asterisk indicates 90% bootstrap support, whereas all other nodes had 100% support, 1,000 iterations. (e) Histogram showing the respective numbers of SNPs between the aligned M. tuberculosis H37Rv reference and M. bovis or STB genomes (depicted in b).
Comparison of these five STB genomes with those of M. tuberculosis H37Rv12 and other MTBC members13 showed very similar overall organization between STB and MTBC strains, with a high percentage of syntenic genes, ranging from 93% for STB-K to 96% for STB-A, compared to only 77% between Mycobacterium marinum, one of the phylogenetically closest non-tuberculous mycobacterial species14, and M. tuberculosis H37Rv. No major chromosomal rearrangements or plasmids were detected (Fig. 1). Pairwise analyses between the conserved STB and MTBC genome sequences showed that all combinations had average nucleotide identities of at least 97.3%, above the 95% threshold proposed for classification into the same species15. However, the genomes of the STB strains were 10–115 kb larger than those of the MTBC members and thus represent the largest genomes known for tubercle bacilli, although they are still much smaller than those of M. marinum (6.6 Mb)16 and the other most closely related non-tuberculous species Mycobacterium kansasii (6.4 Mb)17. Excluding repetitive sequences such as PE_PGRS- and PPE_MPTR-encoding regions, which account for ∼8% of the coding capacity of M. tuberculosis12, STB and MTBC strains shared >89.3% of their genomes, representing a core genome for tubercle bacilli of >3.938 Mb. This core comprises 96.3% of the 774 M. tuberculosis H37Rv genes predicted to be essential for in vitro growth and all 194 genes required for mycobacterial survival during mouse infection18,19,20, further reflecting the close affiliations of STB strains and M. tuberculosis. The accessory genomes of individual STB strains harbor from 124 (STB-A) to 366 (STB-J and STB-K) genes not present in MTBC members that enlarge the known pan-genome of tubercle bacilli by 890 predicted coding sequences, representing a supplement of more than 20% relative to the gene pool of M. tuberculosis (Supplementary Fig. 2a,b and Supplementary Table 3). Notably, only nine of these predicted coding sequences were common to all five STB genomes analyzed (Supplementary Fig. 2c and Supplementary Table 3). Conversely, 51 genes partially overlapping with genomic islands21 present in MTBC members were not found in any of the STB strains (Supplementary Table 4). These genes encode derivatives of mobile elements, such as the phiRv1 and phiRv2 prophage-like regions (24 coding sequences), 3 transposases, 5 unique members of a glycine-rich protein family (for example, PE_PGRS33; Supplementary Fig. 3a) and 19 other hypothetical proteins (Supplementary Fig. 3b). It is noteworthy that Rv1989c-Rv1990c from one such MTBC-specific region showed around 90% identity with proteins encoded on a plasmid from Mycobacterium gilvum and Mycobacterium sp. KMS, raising intriguing questions about possible transmission routes by which the corresponding genes were introduced into the MTBC genomes. Several other MTBC-specific hypothetical proteins had no or only weak amino-acid similarity with other mycobacterial proteins (Supplementary Table 4), suggesting horizontal gene transfer into the MTBC lineage from distant donors after its separation from the STB lineages.
We also identified prominent horizontal gene transfer–related differences in clustered, regularly interspaced short palindromic repeats (CRISPR)-associated protein (CRISPR-Cas) systems between STB and MTBC strains. These systems may confer adaptive immunity against phages and plasmids in bacteria and archaea via repeat-spacer–derived RNAs22. The genomes of STB-A and STB-D contain a single CRISPR-Cas locus encoding a system of major type III-A that is similar to that of MTBC genomes but with a few crispr spacers in common7,10 and substantially lower sequence similarity for their Cas proteins (down to 75%) than is seen for the core proteins (98–100%) (Fig. 2). The same genomic region in the more distant STB-J, STB-K and STB-L strains is occupied by a completely different CRISPR-Cas system of a rare type I-C variant (Fig. 2), most closely related to those of environmental actinobacteria, such as Gordonia amarae, or purple sulfur bacteria from the Thioalkalivibrio genus. Furthermore, in STB-K, the presence of a second CRISPR-Cas module of a different type I-C was identified 260 kb upstream of the other locus (Fig. 2), encoding Cas proteins that were most similar to those of Moorella or Thiorhodovibrio species. Finally, screening of whole-genome shotgun–derived sequences from strains STB-E, STB-G, STB-H and STB-I, located at well-distributed intermediate positions on the STB MLST-based network (Fig. 1), showed the existence of yet another type I-E module in STB-G and STB-I that was most closely related to those of environmental actinobacteria, such as members of the Saccharomonospora genus, whereas, in the two remaining STB-E and STB-H strains, a type I-C variant similar to those in strains STB-J, STB-K and STB-L was found (Fig. 2a). As CRISPR-Cas systems have not been identified in non-tuberculous mycobacterial species, these different systems were most likely acquired by independent horizontal gene transfer events that occurred after the divergence of the STB and MTBC lineages. Although it is not known whether the CRISPR systems in tubercle bacilli are functional, their disparate origins suggest that the distinct respective crispr spacer sets might not necessarily reflect genetic records of recent encounters of tubercle bacilli with distinct phage transgressors but could also represent older traces of interaction of donor organisms encoding the respective CRISPR-Cas systems with non-mycobacterial phages. The identification of a 55-kb prophage region in the whole-genome shotgun–derived sequence of STB-I that is large enough to encode a potentially complete virion23 (Fig. 2), which to our knowledge represents the first such finding in tubercle bacilli, provides a promising future model for testing the functionality of mycobacterial CRISPR-Cas systems on adaptive immunity against phages.
(a) Gene content of different CRISPR-Cas systems in MTBC and STB strains. Spacers are colored according to sequence similarity. Percentages of protein sequence identity are indicated between the type III-A systems of M. tuberculosis H37Rv and STB-A and STB-D. The various combinations of identities between ubiquitous proteins (for example, Cas2) of different CRISPR-Cas types are much lower (below 40%) and are not indicated. A star indicates a potential csb1 pseudogene in the system of STB-H. A broken line denotes the ends of sequence contigs in repeat zones of the type I-E systems of STB-G and STB-I. Mutual identity, mutual protein sequence identity; tnp, transposon. cas, csm, csb, csx and cse represent various cas gene families. (b) Schematic of a 55-kb spanning genomic region that encodes a putative prophage in STB-I. STB-I genomic positions are marked on the horizontal lines. The brackets enclose a portion homologous to a prophage region in the M. marinum genome. Predicted coding sequences are shown above and below the genomic positions, corresponding to transcription to the right and left, respectively. Colors define the features of the predicted encoded products: gray, phage protein without database match or homologous to non-mycobacteriophage proteins of unknown function; blue, phage protein homologous to other mycobacteriophage proteins of unknown function (names of homologs are shown in blue, except for the portion homologous to the M. marinum prophage region); black, STB-I coding sequences and tRNA genes (conserved in other STB strains and M. tuberculosis H37Rv) flanking the phage insertion site corresponding to the lysT gene (Lys tRNA); all other colors, phage proteins with a predicted function (indicated in black text). A gray box on the second horizontal line indicates a sequence contig break. Functional annotations of the predicted genes were made on the basis of comparisons of the encoded products via the Genbank database, detection of protein domain signatures and expert annotation of 374 other mycobacteriophage genomes retrieved from the PhagesDB database.
Progressive genome downsizing is a hallmark of mycobacterial pathogen evolution17,24. Therefore, the larger genome sizes of STB compared with MTBC strains argue for the ancestral status of the STB lineages. Further evidence for the ancestral nature of STB genome structures comes from inspection of interrupted coding sequences (ICDSs), thought to reflect molecular scars inherited during the pseudogenization of the MTBC genomes25,26. Among the 81 reported ICDSs in MTBC strains, most were found to be also interrupted in STB strains and more distantly related mycobacteria, suggesting that they are evolutionary ancient mycobacterial scars (Supplementary Table 5). However, we identified four ICDS orthologs—for example, pks8 belonging to a multigene family encoding polyketide synthases that are involved in the biosynthesis of important cell envelope lipids16,27—which were intact in the genomes of STB strains (in one case, rv3741c-rv3742c, the region was absent from STB-J) and in the M. marinum and/or M. kansasii outgroup genomes (Supplementary Fig. 4). Thus, these scars occurred in the most recent common ancestor of the MTBC lineage after divergence from STB-like progenitors. The opposite situation, that is, where ICDSs shared by the STB genomes corresponded to intact coding sequences in MTBC strains, was not observed, further supporting the ancestral status of the STB genome structure. In addition, we detected four independent loci (narX, pks5, pknH and lppV) where a likely ancestral gene organization, present both in the mycobacterial outgroups M. marinum and/or M. kansasii and in STB strains, was rearranged to result in a single hybrid gene and loss of intervening gene(s) in MTBC genomes (Supplementary Fig. 5), similar to what has been observed for pknH in M. africanum28.
Ancient branching of STB lineages is also consistent with the much higher number of SNPs detected in STB genomes compared to those of MTBC strains. Pairwise comparisons of STB-A, STB-D, STB-J, STB-K and STB-L genome sequences with the M. tuberculosis H37Rv reference uncovered 16,168–61,228 SNPs (Fig. 1). This number is within the range of 9,525–65,744 SNPs observed in the group of STB strains alone and up to 25-fold higher than the 741–2,437 SNPs previously observed in members of the MTBC lineage13,29,30. Consistent with MLST data (here and in ref. 9), a NeighborNet analysis based on pairwise comparisons of the genome-wide SNP data showed that MTBC strains form a single compact group within a much larger reticulated network of STB genotypes (Fig. 1). Consistently, this reticulation was even increased when whole-genome shotgun–derived sequence data from four additional STB strains were included (Supplementary Fig. 6), further confirming the MLST-derived phylogeny at the genome level. The Phi test for recombination was highly significant (P = 1 × 10−6). Notably, the relative compactness of the MTBC branch was additionally confirmed by the structure of the phylogenetic tree, obtained after exclusion of the genome portions affected by recombination and/or horizontal gene transfer (Fig. 3a). These results thus show that the worldwide MTBC population represents a genetically homogeneous subset branching from the larger diversity of recombinogenic STB isolates. Taken together with independent lines of evidence pointing to earlier branching, these findings suggest that the STB lineages diverged from the common ancestor of all tubercle bacilli well before the successful clonal radiation of MTBC strains began.
(a) Phylogenetic tree inferred using the Neighbor-Joining algorithm on nucleotide p distances, after concatenation of the sequence alignments of 2,047 genes of the predicted clonal portion of the STB-MTBC core genome (after exclusion of the genes affected by recombination and gapped regions). (b) SNP distribution among STB and MTBC aligned genome segments, showing probable recombination regions involving genes rv1936-rv1937 between STB-J and M. tuberculosis. Each of the three panels shows a comparison of two STB or M. bovis strains (top and bottom) relative to M. tuberculosis H37Rv (middle). Red lines indicate individual SNPs identified between pairwise-compared genomes. Thicker or uneven red lines result from multiple SNPs in close proximity or shifts due to small indels. Note the SNP-free identical genome segments in STB-J and H37Rv (boxed) that conflict with their distant respective positions on the clonal core genome–based tree.
Impact of selection and recombination
To compare the effect of selection on the evolution of the STB and MTBC genomes, we calculated global ratios of nonsynonymous versus synonymous SNPs (dN/dS). The genome-wide dN/dS ratio is unusually high in MTBC strains, which has been suggested to reflect relaxed purifying selection against nonsynonymous changes that are in general slightly deleterious31. The dN/dS ratios in different gene categories among the STB strains were only about a third of those found in the MTBC strains (Table 1) and are thus compatible with a much longer time during which STB strains were exposed to purifying selection, given the time dependence of dN/dS ratios for closely related bacteria32,33,34 and assuming that purifying selection pressures were the same for STB strains as for MTBC ones.
As an important exception, the sequences encoding the protective human CD4+ and CD8+ T-cell antigens and the epitopes of M. tuberculosis have been described to be under purifying selection, suggesting that MTBC members do not use T-cell antigen variation to escape human immune responses but might instead benefit from recognition by T cells30. Similarly, we found that the dN/dS ratios based on pairwise concatenated codon alignments35,36 of the 65 T-cell antigen–encoding STB genes conserved across all STB genomes were on average lower than those of the 2,300 genes classified as nonessential and similar or slightly lower than those of the 710 essential genes18 conserved among all STB genomes (Table 1). Overall, similar results were also obtained when only the epitope-encoding regions of the T-cell antigens were considered. Thus, similar to the subset of essential proteins, human T-cell antigens tend to be more conserved in STB relative to the rest of the proteome. Following the argument of Comas and colleagues30, this sequence conservation suggests that the STB and MTBC lineages might have inherited a common strategy of immune subversion of the human host that predates the clonal emergence of the MTBC lineage. However, there may be alternative explanations, as most of the antigen-encoding regions with low dN/dS ratios are also highly conserved in the environmental facultative pathogens M. marinum and/or M. kansasii and/or in other mycobacteria. For example, the 6-kDa early secreted antigenic target (ESAT-6, EsxA, Rv3875) and the 34.6-kD secreted antigen 85B (Ag85B, FbpB, Rv1886c), that both show 100% amino-acid conservation in MTBC and STB strains, have corresponding orthologs in M. marinum that show 91% (ESAT-6) and 89% (Ag85B) amino-acid identity, which is above the average overall pairwise identity of 85.2% (ref. 16). The conservation of these proteins might thus also be explained by their role in host-pathogen interaction, such as phagosomal rupture37, or mycobacterial cell envelope stability38 and/or other functions that are not necessarily linked to interactions with human T cells.
Extensive recombination among STB genomes, as identified by our comparative genome analysis, might be partly responsible for the discrepancy in dN/dS ratios between the MTBC and STB groups, as it could more efficiently oppose fixation of slightly deleterious mutations than in the more clonal MTBC population39. Consistent with this contention, strong variations in the local distribution of SNPs were observed throughout the aligned STB and MTBC genomes, suggestive of numerous recombination events. Approximately one-third of the core genome alignment consists of zones with significantly lower or higher SNP density compared to expectations for predicted recombination-free nucleotide differences between each pair of genomes. Stringent selection of informative regions in the predicted recombination-free blocks led to the identification of a minimal clonal backbone of 1,794,643 characters (∼33% of the core genome), which was used to infer a phylogenetic tree (Fig. 3a). Inspection of the genomic regions with unexpected SNP densities allowed us to identify >110 blocks of up to 14 kb that each included from 1 to 12 complete genes (Supplementary Table 6), with homoplasic SNP distributions (relative to the tree), indicative of likely interstrain recombination events among STB strains and/or between STB and MTBC strains (Fig. 3b and Supplementary Fig. 7a,b). The extensive impact of recombination was independently confirmed by the finding that ∼8–15% of the protein-coding sequence alignments from the core genome had mosaic structures indicative of interstrain intragenic recombination events. In contrast, the influence of exogenous importation from more distant mycobacterial species on core genome sequence diversity seemed to be minimal, as inferred by the detection of only a few regions with unexpectedly high SNP densities in STB strains, yielding BLAST best hits closer to non-tuberculous mycobacteria than to STB and MTBC strains (Supplementary Fig. 7c).
Notably, the gene blocks in M. tuberculosis, whose sequences perfectly match those of one or more STB strain, showed SNPs in the orthologous region in M. bovis and/or other MTBC strains (Fig. 3b), suggesting that gene flux between M. tuberculosis and the pool of STB strains existed even well after the divergence of the MTBC lineage and perhaps still exists. We also found intermediate situations where the SNP distribution clearly suggested recombination events that were more ancient and likely followed by accumulation of a few mutations in the recipient or the donor strains (Supplementary Fig. 7a). These data provide new solid evidence for the discussion on potential interstrain gene flux in M. tuberculosis40,41. Our findings also raise puzzling questions about the (micro)environments and mechanisms that favor or have favored such extensive DNA exchanges. The high number of apparently recent recombination episodes, as suggested by numerous perfect large sequence matches detected among sequences from different STB lineages together with the almost exclusive isolation of STB strains from affected individuals around the Horn of Africa strongly suggests a common local source. Aquatic environments rich in mycobacteria, potentially residing in protozoan hosts24,42, are one possible opportunity for genetic exchange to occur, as suggested by a recent report on the detection of MTBC DNA in rural water sources in Ethiopia (E. Wellington, personal communication). The presence of a 55-kb genomic segment corresponding to a putative complete phage-encoding region inserted into the lysT gene (Lys tRNA) of the STB-I strain (Fig. 2) suggests a possible mediation by phages, although alternative mechanisms, such as DNA transfer by conjugation, reported for Mycobacterium smegmatis under biofilm conditions43, could also be involved.
STB strains show reduced persistence
To determine whether these genetic differences between the STB and MTBC strains affect host-pathogen interactions, we first measured the growth of these strains in in vitro cultures. Most STB strains grew 2 to 3 times faster than M. tuberculosis strains, both in liquid media (Supplementary Fig. 8) and on solid media (data not shown) at 30 °C and 37 °C, in line with previous observations10,11. Following infection of BALB/c mice (Fig. 4) and C57BL/6 mice (Supplementary Fig. 9) by aerosol, the STB strains effectively multiplied in the lungs and disseminated to the spleens during the acute infection phase but consistently persisted less well during the chronic infection phase compared to M. tuberculosis. Whereas the latter was able to persist in the lungs for up to 30 weeks at levels close to those seen in the acute phase (peaking at 3 weeks with around 1 × 107 to 1 × 108 colony-forming units (CFUs)), the infection levels of all STB strains dropped by at least 1 log at all (and by 2 to 3 logs at most) later time points in these organs (P = 0.05 by Mann-Whitney test, except for day 130 for STB-D, STB-K and STB-L). The strongest difference with M. tuberculosis was observed for STB-K, the strain phylogenomically most distant from MTBC strains, for which bacterial counts were undetectable after 30 weeks in BALB/c mice (Fig. 4c,d). Similar trends were observed in spleens, with STB-K also almost completely cleared at day 210. In parallel, histopathological analyses showed less intense lung lesions and inflammation 128 d after infection with the STB strains compared to M. tuberculosis infection, with STB-K showing the least damage (Fig. 4 and Supplementary Table 7). Furthermore, C57BL/6 mice intravenously infected with high doses of STB survived in contrast to controls infected with M. tuberculosis strains of different lineages (data not shown), confirming the decreased virulence of STB strains.
(a,b) CFUs recovered from the lungs (a) and spleens (b) of BALB/c mice 0–130 d after intranasal infection with 1,000 CFUs. (c,d) In an independent experiment, CFUs recovered from the lungs (c) and spleens (d) of BALB/c mice 0–210 d after intranasal infection with 1,000 CFUs. Data are shown as the median and range of CFUs from four mice. (e) Histopathological sections of the lungs of BALB/c infected mice 128 d after intranasal infection with 1,000 CFUs. Scale bar, 5 mm. (f) Magnified images (20×) of boxed regions in e. Blue circles show bronchi. A, alveoli; V, blood vessels.
Finally, we determined whether these variations could be correlated to differences in the innate or adaptive immune responses elicited by infection. The STB and M. tuberculosis strains were similarly able to induce maturation of innate immunity cells in vitro, such as dendritic cells derived from wild-type, Tlr2-null, Tlr4-null or double-knockout C57BL/6 mice (data not shown), suggesting shared major pathogen-associated molecular patterns (PAMPs)44. Consistently, substantial recruitment of activated innate immune cells, for example, CD11b+BST-2+ and CD11c+MHC-IIhi cells, was observed in vivo in the lung parenchyma of severe combined immunodeficient (SCID) mice after 3 weeks of infection by STB strains but to a lesser extent than with M. tuberculosis infection (data not shown). With regard to adaptive responses, massive recruitment of activated CD4+ or CD8+ T cells, showing CD44 modulation and CD45RB, CD27 and CD62L downregulation, was detected in the lungs of C57BL/6 mice after 13 weeks of infection by strains with smooth colony morphology. Again, the responses were overall quantitatively lower for STB strains compared to M. tuberculosis strains, especially for STB-K (Supplementary Fig. 10), in line with the lower virulence and persistence of the STB strains.
Discussion
With the larger pan-genome reflecting the ancestral, wider gene pool of tubercle bacilli, their lower virulence and faster growth, especially at temperatures below 37 °C, plausibly reflecting broader environmental adaptability, STB strains might thus come nearer to the as-yet-unknown missing link between the obligate pathogen M. tuberculosis and environmental mycobacteria. We propose that M. tuberculosis has evolved its widespread, pathogenic lifestyle starting from a pool of STB-like mycobacteria by gaining additional virulence and persistence mechanisms through a potential combination of (i) loss of gene function, (ii) acquisition of new genes via horizontal gene transfer, (iii) interstrain recombination of gene clusters and (iv) fixation of SNPs. The data presented here suggest further experiments to examine which of these genetic events were involved. Primary candidates are MTBC-specific genes (Supplementary Table 4), including the prophage-like phiRv1- and phiRv2-encoding regions reported to be important for late infection45, the genes encoding PE_PGRS33 or other MTBC-acquired PE or PPE proteins known to enhance cellular toxicity46, the region encoding the polyketide synthases Pks8 and Pks17, the large prophage region in STB-I and/or CRISPR-Cas systems. The insights gained through our analysis bring new perspectives to the identification of potential targets to combat tuberculosis infection and disease.
URLs.
Magnifying Genome (MaGe) server, https://www.genoscope.cns.fr/agc/microscope/about/collabprojects.php?P_id=44ancreLogin; MycoBrowser database, http://mycobrowser.epfl.ch/; The Mycobacteriophage Database, http://phagesdb.org/.
Methods
Bacterial strains and MLST.
The 55 STB and 10 reference MTBC isolates are described in Supplementary Table 1. Twelve housekeeping genes were selected for MLST47 (Supplementary Table 2). Phylogenetic groupings were identified by split decomposition analysis48 on the concatenated target sequences.
Genome sequencing.
Genomic DNA was extracted from cultured single bacterial colonies as described12. For genome sequencing of STB-D, STB-J, STB-K and STB-L, Sanger reads from 10-kb fragment shotgun libraries at 4- to 4.9-fold coverage were assembled with contigs obtained from Newbler assemblies of 454/Roche reads at 13- to 18.1-fold coverage, using Arachne49. Scaffolds were validated using the Mekano interface (Genoscope). Primer walking, PCR and in vitro transposition were used for finishing. The assembled consensus sequences were validated using Illumina reads at 45- to 105-fold coverage and consed functionalities and by mapping of termini sequences from BAC libraries50. High-quality contiguous genome sequences of 4,420 kb (STB-L, 9 contigs), 4,432 kb (STB-D, 12 contigs), 4,524 kb (STB-J, 11 contigs) and 4,525 kb (STB-K, 9 contigs) were generated. Remaining gaps estimated not to exceed 2 kb correspond to GC-rich and repetitive regions coding for PE_PGRS proteins and/or the pks5 region (STB-J). For STB-A, a fully finished contiguous sequence of 4,482,059 bp was obtained using ∼80,000 shotgun Sanger reads, Illumina-generated reads and finishing12,28. Whole-genome shotgun data from strains STB-E, STB-G, STB-H and STB-I were generated using Illumina HiSeq technology and single lanes. Resulting reads that covered up to 900× of the genomes of these STB strains were assembled using Velvet software51, and contigs were ordered using M. canettii CIPT 140010059 (STB-A) and M. tuberculosis H37Rv as reference genomes.
Annotation and comparative genomics.
Annotation and genome comparisons were performed with the Microscope platform52, Artemis and Artemis comparison tool (ACT)53. When applicable, annotations were transferred from those of M. tuberculosis orthologs in the TubercuList/Mycobrowser database, using BLAST matches of > 90% protein sequence identity, an alignable region of >80% of the shortest protein length in pairwise comparisons and visual inspection of the gene synteny. Pairwise average nucleotide identities were calculated using JSpecies54. The core and accessory genomes of STB and M. tuberculosis were determined as described16.
SNP and indel analysis.
The SNiPer pipeline (Genoscope), based on the SSAHA2 package55, was used to map Illumina reads and detect SNPs and indels of STB strains against a corrected version56 of the M. tuberculosis H37Rv reference sequence (NC_000962)12. After the exclusion of ambiguous sequences mapping to repeat regions, an average of 4.7 million split paired-end reads of 36 bp (STB-A, STB-D, STB-J and STB-L) or trimmed at 50 bp (STB-K) were mapped at a resulting genome coverage of >40×. SNPs with base coverage of <10×, base quality of <25 or heterozygosity of >0.2 were removed. ACT53 comparison files were created using MUMmer and NUCmer softwares57 to visualize the SNP distribution in local genome regions.
Calculation of dN/dS ratios.
dN/dS ratios were calculated on orthologs conserved in all STB strains and M. tuberculosis H37Rv, as identified by bidirectional best hits, an alignable region of >80% and sequence identity of ≥30%. Pairwise, concatenated codon alignments between M. tuberculosis H37Rv and each STB strain were generated using PAL2NAL58, after respective protein alignments obtained with MUSCLE59. Synonymous and nonsynonymous substitutions were defined using Nei-Gojobori method-based SNAP35 or maximum likelihood–based PAML36. STB T-cell antigen, essential and nonessential gene categories, as well as T-cell epitope codon concatenates, were constructed as described30.
Recombination.
The genomes of M. tuberculosis H37Rv, M. bovis AF2122/97 and the five STB strains were aligned using progressiveMauve60. Given a pair of aligned genomes i and j, the number of SNPs xij observed between i and j within a region of length l follows a binomial distribution B(l, pij), where pij is the expected proportion of recombination-free nucleotide differences between taxa i and j. Regions containing at least one pair of sequences i and j with an unexpectedly high or low number xij of SNPs, that is, min(P(X ≥ xij), P(X ≤ xij)) < 0.05, where X ∼ B(l, pij), were identified using a 200-character-long sliding window along the conserved (core) portions of the multiple-genome alignment. The value pij inside each window was estimated as the proportion of SNPs between i and j within the 10,000 aligned characters flanking the sliding window on both sides. To obtain a reference phylogeny, all regions of ≥500 characters in length (excluding gaps) that did not contain an unexpected number of SNPs were concatenated. The derived supermatrix was used to infer a phylogenetic tree with the Neighbor-Joining algorithm61 on the pairwise nucleotide p distances. All regions of ≥500 characters in length with a significantly high or low number of SNPs were inspected visually for detection of the concentration of homoplasic characters using ACT53, leading to the identification of similarities between strains incongruent with the phylogenetic tree. The proportion of protein-coding sequences within the core genome probably affected by interstrain recombination was assessed with the pairwise homoplasy index62, the maximum X2 test63 and the neighbor similarity score64.
Bacterial growth assays.
The growth rates of STB and reference MTBC strains in liquid media were measured using a BACTEC 460 system (Beckton-Dickinson) as recommended by the manufacturer.
Mouse infection experiments and histopathological and cell analyses.
Mice were maintained according to the Institut Pasteur de Lille and Paris guidelines for laboratory animal husbandry. Animal experiments were approved by the Nord-Pas-De-Calais ethical committee (CEEA 15/2009) and the Institut Pasteur Hygiene Committee (authorization number 75-1469), in accordance with European and French guidelines (Directive 86/609/CEE and Decree 87–848). Eight-week-old female BALB/c mice were infected by the intranasal route with 1,000 CFUs of either STB or M. tuberculosis H37Rv strains. At the indicated times, four mice per group were sacrificed, and colony counting was performed from homogenized individual lungs and spleens as described65. For histopathological evaluation, whole lungs were harvested from three BALB/c mice for each group 128 d after infection, fixed in 4% formalin and embedded in paraffin. Four-mm-thick sections were stained with hematoxylin and eosin. Virulence and cell analysis–based immunological assays using C57BL/6 and/or SCID mice were performed as described66,67. Adaptive immune cells from infected mice were prepared, incubated with conjugated monoclonal antibodies (Beckton-Dickinson), fixed and analyzed using a CyAn system and Summit (Beckman Coulter) and FlowJo (Treestar) software.
Accession codes.
The complete genome sequence for the STB-A strain (CIPT 140010059) was deposited under accession HE572590. Genome sequences of strains STB-D (CIPT 140060008), STB-J (CIPT 140070017), STB-K (CIPT 140070010) and STB-L (CIPT 140070008) were deposited in the European Molecular Biology Laboratory (EMBL) database under project/accession codes PRJEB94/FO203507, PRJEB93/FO203508, PRJEB92/FO203509 and PRJEB95/FO203510, respectively. Illumina-derived whole-genome shotgun sequences for strains STB-E (CIPT 140070002), STB-G (CIPT 140070005), STB-H (CIPT 140070013) and STB-I (CIPT 140070007) were deposited in the EMBL whole-genome sequence repository under project/accession codes PRJEB584/CAOL00000000, PRJEB585/CAOM00000000, PRJEB586/CAON00000000 and PRJEB587/CAOO00000000, respectively.
Accession codes
Primary accessions
EMBL/GenBank/DDBJ
European Nucleotide Archive
NCBI Reference Sequence
Referenced accessions
NCBI Reference Sequence
References
Dye, C. & Williams, B.G. The population dynamics and control of tuberculosis. Science 328, 856–861 (2010).
Sreevatsan, S. et al. Restricted structural gene polymorphism in the Mycobacterium tuberculosis complex indicates evolutionarily recent global dissemination. Proc. Natl. Acad. Sci. USA 94, 9869–9874 (1997).
Wirth, T. et al. Origin, spread and demography of the Mycobacterium tuberculosis complex. PLoS Pathog. 4, e1000160 (2008).
Brosch, R. et al. A new evolutionary scenario for the Mycobacterium tuberculosis complex. Proc. Natl. Acad. Sci. USA 99, 3684–3689 (2002).
Supply, P. et al. Linkage disequilibrium between minisatellite loci supports clonal evolution of Mycobacterium tuberculosis in a high tuberculosis incidence area. Mol. Microbiol. 47, 529–538 (2003).
Hirsh, A.E., Tsolaki, A.G., DeRiemer, K., Feldman, M.W. & Small, P.M. Stable association between strains of Mycobacterium tuberculosis and their human host populations. Proc. Natl. Acad. Sci. USA 101, 4871–4876 (2004).
van Soolingen, D. et al. A novel pathogenic taxon of the Mycobacterium tuberculosis complex, Canetti: characterization of an exceptional isolate from Africa. Int. J. Syst. Bacteriol. 47, 1236–1245 (1997).
Fabre, M. et al. High genetic diversity revealed by variable-number tandem repeat genotyping and analysis of hsp65 gene polymorphism in a large collection of “Mycobacterium canettii” strains indicates that the M. tuberculosis complex is a recently emerged clone of “M. canettii”. J. Clin. Microbiol. 42, 3248–3255 (2004).
Gutierrez, M.C. et al. Ancient origin and gene mosaicism of the progenitor of Mycobacterium tuberculosis. PLoS Pathog. 1, e5 (2005).
Fabre, M. et al. Molecular characteristics of “Mycobacterium canettii” the smooth Mycobacterium tuberculosis bacilli. Infect. Genet. Evol. 10, 1165–1173 (2010).
Koeck, J.L. et al. Clinical characteristics of the smooth tubercle bacilli ′'Mycobacterium canettii' infection suggest the existence of an environmental reservoir. Clin. Microbiol. Infect. 17, 1013–1019 (2011).
Cole, S.T. et al. Deciphering the biology of Mycobacterium tuberculosis from the complete genome sequence. Nature 393, 537–544 (1998).
Garnier, T. et al. The complete genome sequence of Mycobacterium bovis. Proc. Natl. Acad. Sci. USA 100, 7877–7882 (2003).
Springer, B., Stockman, L., Teschner, K., Roberts, G.D. & Bottger, E.C. Two-laboratory collaborative study on identification of mycobacteria: molecular versus phenotypic methods. J. Clin. Microbiol. 34, 296–303 (1996).
Goris, J. et al. DNA-DNA hybridization values and their relationship to whole-genome sequence similarities. Int. J. Syst. Evol. Microbiol. 57, 81–91 (2007).
Stinear, T.P. et al. Insights from the complete genome sequence of Mycobacterium marinum on the evolution of Mycobacterium tuberculosis. Genome Res. 18, 729–741 (2008).
Veyrier, F.J., Dufort, A. & Behr, M.A. The rise and fall of the Mycobacterium tuberculosis genome. Trends Microbiol. 19, 156–161 (2011).
Sassetti, C.M., Boyd, D.H. & Rubin, E.J. Genes required for mycobacterial growth defined by high density mutagenesis. Mol. Microbiol. 48, 77–84 (2003).
Sassetti, C.M. & Rubin, E.J. Genetic requirements for mycobacterial survival during infection. Proc. Natl. Acad. Sci. USA 100, 12989–12994 (2003).
Griffin, J.E. et al. High-resolution phenotypic profiling defines genes essential for mycobacterial growth and cholesterol catabolism. PLoS Pathog. 7, e1002251 (2011).
Becq, J. et al. Contribution of horizontally acquired genomic islands to the evolution of the tubercle bacilli. Mol. Biol. Evol. 24, 1861–1871 (2007).
Makarova, K.S. et al. Evolution and classification of the CRISPR-Cas systems. Nat. Rev. Microbiol. 9, 467–477 (2011).
Hendrix, R.W., Smith, M.C., Burns, R.N., Ford, M.E. & Hatfull, G.F. Evolutionary relationships among diverse bacteriophages and prophages: all the world's a phage. Proc. Natl. Acad. Sci. USA 96, 2192–2197 (1999).
Gordon, S.V., Bottai, D., Simeone, R., Stinear, T.P. & Brosch, R. Pathogenicity in the tubercle bacillus: molecular and evolutionary determinants. Bioessays 31, 378–388 (2009).
Deshayes, C. et al. Detecting the molecular scars of evolution in the Mycobacterium tuberculosis complex by analyzing interrupted coding sequences. BMC Evol. Biol. 8, 78 (2008).
Smith, N.H., Hewinson, R.G., Kremer, K., Brosch, R. & Gordon, S.V. Myths and misconceptions: the origin and evolution of Mycobacterium tuberculosis. Nat. Rev. Microbiol. 7, 537–544 (2009).
Reed, M.B. et al. A glycolipid of hypervirulent tuberculosis strains that inhibits the innate immune response. Nature 431, 84–87 (2004).
Bentley, S.D. et al. The genome of Mycobacterium africanum West African 2 reveals a lineage-specific locus and genome erosion common to the M. tuberculosis complex. PLoS Negl. Trop. Dis. 6, e1552 (2012).
Brosch, R. et al. Genome plasticity of BCG and impact on vaccine efficacy. Proc. Natl. Acad. Sci. USA 104, 5596–5601 (2007).
Comas, I. et al. Human T cell epitopes of Mycobacterium tuberculosis are evolutionarily hyperconserved. Nat. Genet. 42, 498–503 (2010).
Hershberg, R. et al. High functional diversity in M. tuberculosis driven by genetic drift and human demography. PLoS Biol. 6, e311 (2008).
Rocha, E.P. et al. Comparisons of dN/dS are time dependent for closely related bacterial genomes. J. Theor. Biol. 239, 226–235 (2006).
Castillo-Ramírez, S. et al. The impact of recombination on dN/dS within recently emerged bacterial clones. PLoS Pathog. 7, e1002129 (2011).
Croucher, N.J. et al. Rapid pneumococcal evolution in response to clinical interventions. Science 331, 430–434 (2011).
Korber, B. HIV signature and sequence variation analysis. in Computational Analysis of HIV Molecular Sequences (eds. Rodrigo, A.G. & Learn, G.H.) Ch. 4, 55–72 (Kluwer Academic Publishers, Dordrecht, The Netherlands, 2000).
Goldman, N. & Yang, Z. A codon-based model of nucleotide substitution for protein-coding DNA sequences. Mol. Biol. Evol. 11, 725–736 (1994).
Simeone, R. et al. Phagosomal rupture by Mycobacterium tuberculosis results in toxicity and host cell death. PLoS Pathog. 8, e1002507 (2012).
Kalscheuer, R., Weinrick, B., Veeraraghavan, U., Besra, G.S. & Jacobs, W.R. Jr. Trehalose-recycling ABC transporter LpqY-SugA-SugB-SugC is essential for virulence of Mycobacterium tuberculosis. Proc. Natl. Acad. Sci. USA 107, 21761–21766 (2010).
Felsenstein, J. The evolutionary advantage of recombination. Genetics 78, 737–756 (1974).
Achtman, M. Insights from genomic comparisons of genetically monomorphic bacterial pathogens. Phil. Trans. R. Soc. Lond. B 367, 860–867 (2012).
Namouchi, A., Didelot, X., Schock, U., Gicquel, B. & Rocha, E.P. After the bottleneck: genome-wide diversification of the Mycobacterium tuberculosis complex by mutation, recombination, and natural selection. Genome Res. 22, 721–734 (2012).
Mba Medie, F., Ben Salah, I., Henrissat, B., Raoult, D. & Drancourt, M. Mycobacterium tuberculosis complex mycobacteria as amoeba-resistant organisms. PLoS ONE 6, e20499 (2011).
Nguyen, K.T., Piastro, K., Gray, T.A. & Derbyshire, K.M. Mycobacterial biofilms facilitate horizontal DNA transfer between strains of Mycobacterium smegmatis. J. Bacteriol. 192, 5134–5142 (2010).
Medzhitov, R. & Janeway, C.A. Innate immunity recognition and control of adaptive immune responses. Semin. Immunol. 10, 351–353 (1998).
Aagaard, C. et al. A multistage tuberculosis vaccine that confers efficient protection before and after exposure. Nat. Med. 17, 189–194 (2011).
Cadieux, N. et al. Induction of cell death after localization to the host cell mitochondria by the Mycobacterium tuberculosis PE_PGRS33 protein. Microbiology 157, 793–804 (2011).
Maiden, M.C. et al. Multilocus sequence typing: a portable approach to the identification of clones within populations of pathogenic microorganisms. Proc. Natl. Acad. Sci. USA 95, 3140–3145 (1998).
Huson, D.H. & Bryant, D. Application of phylogenetic networks in evolutionary studies. Mol. Biol. Evol. 23, 254–267 (2006).
Batzoglou, S. et al. ARACHNE: a whole-genome shotgun assembler. Genome Res. 12, 177–189 (2002).
Brosch, R. et al. Use of a Mycobacterium tuberculosis H37Rv bacterial artificial chromosome library for genome mapping, sequencing, and comparative genomics. Infect. Immun. 66, 2221–2229 (1998).
Zerbino, D.R. & Birney, E. Velvet: algorithms for de novo short read assembly using de Bruijn graphs. Genome Res. 18, 821–829 (2008).
Vallenet, D. et al. MicroScope: a platform for microbial genome annotation and comparative genomics. Database (Oxford) 2009, bap021 (2009).
Carver, T. et al. Artemis and ACT: viewing, annotating and comparing sequences stored in a relational database. Bioinformatics 24, 2672–2676 (2008).
Richter, M. & Rossello-Mora, R. Shifting the genomic gold standard for the prokaryotic species definition. Proc. Natl. Acad. Sci. USA 106, 19126–19131 (2009).
Ning, Z., Cox, A.J. & Mullikin, J.C. SSAHA: a fast search method for large DNA databases. Genome Res. 11, 1725–1729 (2001).
Niemann, S. et al. Genomic diversity among drug sensitive and multidrug resistant isolates of Mycobacterium tuberculosis with identical DNA fingerprints. PLoS ONE 4, e7407 (2009).
Delcher, A.L., Phillippy, A., Carlton, J. & Salzberg, S.L. Fast algorithms for large-scale genome alignment and comparison. Nucleic Acids Res. 30, 2478–2483 (2002).
Suyama, M., Torrents, D. & Bork, P. PAL2NAL: robust conversion of protein sequence alignments into the corresponding codon alignments. Nucleic Acids Res. 34, W609–W612 (2006).
Edgar, R.C. MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res. 32, 1792–1797 (2004).
Darling, A.E., Mau, B. & Perna, N.T. progressiveMauve: multiple genome alignment with gene gain, loss and rearrangement. PLoS ONE 5, e11147 (2010).
Saitou, N. & Nei, M. The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol. Biol. Evol. 4, 406–425 (1987).
Bruen, T.C., Philippe, H. & Bryant, D. A simple and robust statistical test for detecting the presence of recombination. Genetics 172, 2665–2681 (2006).
Smith, J.M. Analyzing the mosaic structure of genes. J. Mol. Evol. 34, 126–129 (1992).
Jakobsen, I.B. & Easteal, S. A program for calculating and displaying compatibility matrices as an aid in determining reticulate evolution in molecular sequences. Comput. Appl. Biosci. 12, 291–295 (1996).
Pethe, K. et al. The heparin-binding haemagglutinin of M. tuberculosis is required for extrapulmonary dissemination. Nature 412, 190–194 (2001).
Majlessi, L. et al. Influence of ESAT-6 secretion system 1 (RD1) of Mycobacterium tuberculosis on the interaction between mycobacteria and the host immune system. J. Immunol. 174, 3570–3579 (2005).
Bottai, D. et al. ESAT-6 secretion–independent impact of ESX-1 genes espF and espG1 on virulence of Mycobacterium tuberculosis. J. Infect. Dis. 203, 1155–1164 (2011).
Acknowledgements
We are grateful to S. Cole for help in initiating the M. canettii CIPT 140010059 genome sequencing project and advice, to H. Dong, T. Garnier, N. Honoré, M. Huerre, A. Kapopoulou and M. Monot for help and fruitful discussions and to Z. Rouy for help with sequence depositing. The work was supported in part by the Institut Pasteur (PTR 314 and PTR 383), European Community's Framework Programme 7 grant 260872, Wellcome Trust grant 098051 and Genoscope collaborative grant 114.
Author information
Authors and Affiliations
Contributions
P.S., M.-C.G. and R.B. designed the study. P.S. and R.B. analyzed data and wrote the manuscript, with comments from all authors. M.M., D.W., J.P., C.M. and S.D.B. annotated the genomes. S.M., E.C.B., M.A.Q., L. Ma, C.B. and V.B. performed and/or verified the finishing and assembly of sequences. D.R. and S.C. performed SNP analyses and database management, with the support of G.S. C.R., A.P., W.F., R.S., A.-S.D., E.W., A. Cascioferro and M.-C.G. performed mouse infection experiments, in vivo data analyses and/or mycobacterial growth assays. L. Majlessi, F.S., C. Locht and C. Leclerc conducted and/or analyzed immune assays. J.T., A. Criscuolo and S.B. conducted MLST, recombination and/or phylogenetic analyses. L.F. conducted histopathological analyses. V.K., M.O. and C.P. created bioinformatics tools and analyzed data. M.F. isolated STB strains. T.S. and T.P.S. conducted core genome and NeighborNet analyses.
Corresponding authors
Ethics declarations
Competing interests
P.S. is a consultant for Genoscreen. All other authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–10 and Supplementary Tables 1, 2 and 4–7 (PDF 6796 kb)
Supplementary Table 3
Alignment of orthologues among MTBC and STB genomes (XLS 1349 kb)
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-Share Alike 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/3.0/
About this article
Cite this article
Supply, P., Marceau, M., Mangenot, S. et al. Genomic analysis of smooth tubercle bacilli provides insights into ancestry and pathoadaptation of Mycobacterium tuberculosis. Nat Genet 45, 172–179 (2013). https://doi.org/10.1038/ng.2517
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.2517
This article is cited by
-
A smooth tubercle bacillus from Ethiopia phylogenetically close to the Mycobacterium tuberculosis complex
Nature Communications (2023)
-
Types and functions of heterogeneity in mycobacteria
Nature Reviews Microbiology (2022)
-
Inhibition of the Niemann-Pick C1 protein is a conserved feature of multiple strains of pathogenic mycobacteria
Nature Communications (2022)
-
Spatial clustering and genetic diversity of Mycobacterium tuberculosis isolate among pulmonary tuberculosis suspected patients, Arsi Zone, Ethiopia
BMC Pulmonary Medicine (2021)
-
Cobalamin is present in cells of non-tuberculous mycobacteria, but not in Mycobacterium tuberculosis
Scientific Reports (2021)