Abstract
Previously, we have identified a putative novel rapidly growing Mycobacterium species, referred to as TNTM28, recovered from the sputum of an apparently immunocompetent young man with an underlying pulmonary disease. Here we provide a thorough characterization of TNTM28 genome sequence, which consists of one chromosome of 5,526,191 bp with a 67.3% G + C content, and a total of 5193 predicted coding sequences. Phylogenomic analyses revealed a deep-rooting relationship to the Mycobacterium fortuitum complex, thus suggesting a new taxonomic entity. TNTM28 was predicted to be a human pathogen with a probability of 0.804, reflecting the identification of several virulence factors, including export systems (Sec, Tat, and ESX), a nearly complete set of Mce proteins, toxin-antitoxins systems, and an extended range of other genes involved in intramacrophage replication and persistence (hspX, ahpC, sodA, sodC, katG, mgtC, ClpR, virS, etc.), some of which had likely been acquired through horizontal gene transfer. Such an arsenal of potential virulence factors, along with an almost intact ESX-1 locus, might have significantly contributed to TNTM28 pathogenicity, as witnessed by its ability to replicate efficiently in macrophages. Overall, the identification of this new species as a potential human pathogen will help to broaden our understanding of mycobacterial pathogenesis.
Similar content being viewed by others
Introduction
Non tuberculous mycobacteria (NTM) are environmental germs that can invade a host and cause lung, skin or lymphatic infections in immunocompetent patients or cause disseminated infections, particularly in immunocompromised individuals1,2,3. Pulmonary infections due to NTM are increasingly recognized worldwide through improved culture and identification techniques4,5. NTM species are generally subdivided on the basis of growth into rapid and slow growing mycobacteria (RGM and SGM, respectively)6. Like the pathogenic Mycobacterium tuberculosis complex (MTBC) members, pathogenicity in NTM is predominantly correlated with slow growth7, yet some RGM species have been associated with true microbiological diseases8.
Among RGM, Mycobacterium abscessus, Mycobacterium fortuitum, and Mycobacterium chelonae, account amongst the most clinically relevant species8. M. abscessus is the most frequently isolated from clinical respiratory specimens, while M. fortuitum is the most common from non-respiratory specimens. The latter species, mainly found in soil, dust, water, and animal sources, encompasses a large group of emerging opportunistic pathogens, generally subdivided into three biovars, referred to as the M. fortuitum complex9,10,11. Reports on novel M. fortuitum-related species have significantly increased over the past decades12,13. Currently, the M. fortuitum complex consists of a monophyletic group of several species including, but not limited to, M. fortuitum stricto sensus (strain CT6), Mycobacterium mageritense, Mycobacterium conceptionense, Mycobacterium septicum, Mycobacterium peregrinum, Mycobacterium porcinum, and Mycobacterium senegalense14.
Infection with M. fortuitum complex species can cause a variety of clinical diseases, being most frequently associated with skin and soft tissues in both immunocompetent and immunocompromised patients15,16. The presence of M. fortuitum species in the respiratory tract has been mainly reported following simple colonization or ephemeral infection. However, a true lung infection due to M. fortuitum remains infrequent, and generally occurs in patients with gastroesophageal disease or in elderly patients with chronic cough17,18,19. Isolation of new M. fortuitum complex species, particularly those associated with pulmonary diseases, is thus worthy of consideration since it could provide new clues to better understand the evolution and pathogenesis of this mycobacterial group.
Previously, we have described a putative new RGM species, referred to as TNTM28, isolated from the sputum of an apparently immunocompetent young man presenting with an underlying pulmonary disease20. Based on sequence polymoprphisms in 16 rRNA, hsp65, and rpoB gene sequences, this non photochromogenic RGM was found to be related to the M. fortuitum complex. Here we provide a genome-based description of TNTM28, which is confirming its phylogenetic link to the M. fortuitum complex, but also shows enough differences to justify the status of a new species. TNTM28 was found to display several features reminiscent of a pathogenic species.
Results
Phenotypic characterization and further genetic analysis of TNTM28
In Löwenstein-Jensen (LJ) medium, TNTM28 appeared as small, nonpigmented, hemishperic colonies (approx. 1 mm in diameter) with a rough morphotype, mostly grouped in rosettes (Fig. 1a). Microscopic analysis of TNTM28 bacilli showed a Gram-positive type, and also displayed red color after Ziehl–Neelsen straining, where the bacteria tended to form large aggregates (Fig. 1b). TNTM28 colonies grew on LJ agar within 2 to 4 days at 37 °C (optimum), in the presence or absence of 5% NaCl. Growth did also occur at 30 °C, albeit less efficiently than at temperatures between 33 and 37 °C. No growth occurred at 42 °C.
Biochemical tests showed that TNTM28 was niacin-negative, but proved positive when tested for arylsulfatase production (after 3 and 14 days), alkaline phosphatase, nitrate reductase, and thermostable catalase activities. TNTM28 was capable to hydrolyse Tween-80 and urea, and was found competent for iron uptake.
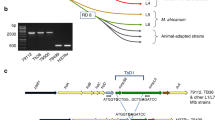
As mentioned above, previous analyses based on 16S rRNA and rpoB gene sequence polymorphism revealed that TNTM28 was phylogenetically related to members of the M. fortuitum complex. Here, we refined such a phylogenetic analysis by including a larger dataset of the M. fortuitum complex group. As shown in the rpoB-based phylogenetic tree depicted in Fig. 1c, TNTM28 was related to the M. fortuitum complex, but proved quite divergent from other species, being deeply rooted to Mycobacterium septicum strain ATCC 700731T (AY147167), Mycobacterium alvei CIP103464 (HM807430), and Mycobacterium peregrinum CIP 105382T (AY147166) with 94.49%, 94.23% and 94.06% rpoB gene sequence similarity, respectively.
General features of TNTM28 genome
After assembling and filtering, based on median coverage ≥ 100 X, the pre-processed 5,469,922 paired-end reads resulted in 50 scaffolds with a total length of 5,526,191 bp. Using the genome of M. fortuitum strain CT6 (the nearest genome sequence) as reference, 28 scaffolds (5,493,022 bp) were mapped, while the remaining 22 (35,969 bp) proved unplaced. The maximum length of scaffolds was estimated to be 851,167 bp. The assembly had N50 and N90 values of 338,742 pb and 103,464 pb, respectively. The genome of TNTM28 displayed a high G + C content of 67.3%, and contained no plasmid replicons. A total of 5193 protein-coding genes (CDS) was identifed, along with 52 tRNA, 1 tmRNA, a single 16S-5S-23S ribosomal RNA operon, and 3 putative CRISPRs loci (Table 1). A circular representation of TNTM28 genome is shown in Fig. 2a. The genome sequence of TNTM28 showed no major genomic rearrangements, being mostly syntenic to that of M. fortuitum CT6 strain (Fig. 2b). Five incomplete prophage regions (A to E) have been identified. They mainly consisted of hypothetical and phage-like protein sequences (Supplementary Figure S1).
Graphical map of TNTM28 genome. (a) Circular map of the 5.49 Mb TNTM28 chromosome performed with GCview Server. The circles represent, from outside to inside: rings 1 and 4 show protein-coding genes oriented in the forward and reverse orientations, respectively. Rings 2 and 3 display genes on forward and reverse strand, respectively. Ring 5 shows G + C% content plot (black). Ring 6 shows GC skews, with positive and negative values being indicated with green and purple colors, respectively. Positions of the prophage regions (P1 to P5) are indicated in the innermost circle. (b) Genome alignment performed using Mauve software between TNTM28 with its closest species, M. foruitum, strain CT6. Boxes with identical colors represent locally collinear blocks (LCBs), indicating homologous DNA regions shared between the two genomes without sequence rearrangement. Lines collate aligned segments between genomes. The vertical bars denote the conservation level, and upward and downward orientations relative to the genome line indicates collinear and inverted regions, respectively. Sequences outside colored blocks do not have homologs in the other genome. Red lines indicate contig boundaries within the assembly.
Functional annotation
The COGs content of the genome sequence of TNTM28 was analyzed and compared to that of five M. fortuitum complex species (M. fortuitum, M. peregrinum, M. porcinum, M. senegalense, and M. conceptionense). As shown in Fig. 3, the total number of genes in the six species varied from 4716 to 5570. Curiously, TNTM28 displayed the lowest number of genes, which reflects its particular phylogenetic status. Overall, 3763 shared COGs have been identified among the six species (Fig. 3). We found 64 genes unique to TNTM28, which could have contributed to specific phenotypic traits. These TNTM28-specific genes grouped into 18 gene ontology (GO) categories, particularly energy production and conversion (Supplementary Fig. S2).
The TNTM28 genome sequence was functionally annotated using the orthoMCL database, according to which 32% of the genes have been assigned to either ill-defined functional categories (“R” and “S” categories) or had no homologue (Table 2). The most represented functional categories were “lipid transport and metabolism” (category I; ~ 8.7%), “transcription” (category K; ~ 8.0%), “secondary metabolites biosynthesis, transport and catabolism” (category Q; ~ 7.9%), energy production and conversion (category C, ~ 6.8%), and “amino acid transport and metabolism (category E, ~ 6.0%).
Phylogenomic analysis
The phylogenetic placement of TNTM28 within the Mycobacterium genus was carried out based on MAUVE alignment of the genome sequence of 53 mycobacterial species. As shown in Fig. 4a, TNTM28 branched off from the common ancestor of the M. fortuitum complex, occupying a separate phylogenetic branch intermediate between this complex and M. smegmatis. Orthologous average nucleotide identity (orthoANI) scores rangin from 84.46 (M. fortuitum CT6) to 85.24% (M. septicum) were obtained (Fig. 4b).
Genes potentially acquired through horizontal transfer
The identification of five prophage regions in the genome of TNTM28 is a witness of the occurrence of putative lateral gene transfer (LGT) events during its evolution. Therefore, we sought for horizontally transferred genomic islands (GIs). These acquired regions may encode several beneficial factors endowing the bacillus with a new virulence and drug resistance patterns, as well as higher adaptability to changing environmental conditions. Using the IslandViewer 4 platform and its improved GI prediction tool, the IslandPath-DIMOB21, eight GIs varying in length from 8510 to 37,995 bp have been identified (Supplementary Fig. S3, Supplementary Table S1). Overall, these horizontally transferred regions accounted for 2.28% of TNTM28 genome size and entailed 150 genes (2.88% of the coding capacity), 66% of which encoded hypothetical proteins.
Among the transferred genes with a known function, a large part was devoted to cell wall/membrane biogenesis and/or integrity (n = 7), transcription, as there were 6 helix-turn-helix (HTH) transcriptional regulators of various families (TetR/AcrR, GntR, and AraC), and detoxification (n = 4). We identified reductases (n = 2) and dehydrogenases (n = 2), which are likely to be involved in energy production and conversion, antitoxins (RelB and HipB), genes involved in lipid metabolism (hsaD and fgD), DNA repair (mutT2 and xseA), and carbohydrate metabolism. In addition, a copy of a hemagglutinin gene (hbhA) had been gained through a HGT event.
Putative pathogenic features encoded in the TNTM28 genome
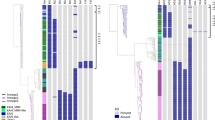
The genome of TNTM28 was found to contain a range of genes which in other mycobacteria are involved in mycobacterial pathogenicity, as defined within the virulence factor database22, endowing this new species with a relatively high probability (0.804) of being a pathogen for humans, as predicted by PathogenFinder23. A detailed distribution of 231 putative virulence genes among related SGM and pathogenic mycobacteria is provided in Supplementary Table S2. In addition, a heatmap was generated with dendograms showing the clustering of 20 mycobacterial species based on the presence or absence of each one of the 231 virulence genes, which further confirmed the relatedness of TNTM28 and the M. fortuitum group (Fig. 5).
Of particular interest, the TNTM28 genome contained an almost complete ESX-1 locus. As shown in Supplementary Fig. 4Sa, this type VII secretion locus, which in many SGM species is involved in virulence, lacked only espJ and espK, with all genes critical for secretion, namely EccA to EccE and MycP, being present. In pathogenic mycobacteria, including Mycobacterium marinum and Mycobacteirum leprae, ESX-1 secretion and function have been shown to be dependent upon the distal espACD operon, which has not been found in RGM genomes analysed so far24. Strikingly, scrutiny of the TNTM28 genome uncovered a distal operon comprising three genes (GEECPEIM_00946, GEECPEIM_00945, and GEECPEIM_00944) whose organization resembled that of the espACD operon (Supplementary Fig. 4Sb), with one gene, GEECPEIM_00945, showing significant orthology with the EspC product, as predicted by reciprocal best hits analysis (Supplementary data 1 to 3). The other two genes, GEECPEIM_00946 and GEECPEIM_00944, though positioned in a way similar to that of espA and espD, respectively, displayed no siginificant orthology with the latters. A similar situation was found in M. fortuitum (Supplementary Fig. 4Sb), but neither in M. abscessus nor in M. smegmatis (data not shown).
Like other NTMs, the TNTM28 genome harbored the ESX-3 locus. In addition, it contained an ESX-4 locus, representing the most ancestral ESX system known for mycobacteria25,26. However, the ESX-4 locus of TNTM28 contained no orthologue of eccE4, which in M. abscessus was deemed crucial for ESX-4 functions, notably with regard to the ability to block phagosomal acidification27.
Furthermore, the genome of TNTM28 encoded a large number (n = 40) of mce proteins, a gene family shown to be critical for invasion and persistence of mycobacteria in host macrophages and non-phagocytic mammalian cells, with mce4 being implicated in cholesterol catabolism28,29. Homologs to M. tuberculosis mce1A, mce1B, mce3E, mce3F, which are lacking in M. abscessus, have been found in the TNTM28 genome. Importantly, the latter genome contained the full set of mce4 genes (mce4A to mce4F). Additional mce genes (mce5 to mce9) specified by other NTM species and Actinomycetales species have been found30,31.
Like the majority of RGM species, we identified a few members of the PE/PPE multigene families, mainly those associated with the three Esx clusters (Esx-1, Esx3, and Esx-4)31.
We also identified seven members of the Sec export system (secA, secD, secE, secF, secG, secY, and yajC), which have been shown to be critical for M. tuberculosis virulence as they ensure the transport to the cytoplasmic membrane, and beyond, of several virulence factors. In particular, the presence of secA and secY, the motor protein and the major component of the translocon, respectively, may have endowed TNTM28 with the ability to ensure many functions important for its survival in the host32. Besides, we identified at least three sec-independent protein secretion pathway components.
Survival of mycobacteria within the host is greatly dependent upon their ability to produce cell wall-associated lipids, siderophores and other biologically active molecules33,34. In this respect, the genome of TNTM28 was found to be well equipped since it contained 40 genes involved in polyketide biosynthesis and 17 others involved in non-ribosomal peptide synthesis (NRPS). The existence of a pks15 gene copy along with ppsA-E genes is noteworthy. In slow growing mycobacteria this locus was shown to be involved in the synthesis of phenolic glycolipids (PGL), representing major virulence factors of pathogenic mycobacteria35, whose presence in TNTM28 can now be experimentally verified. Furthermore, TNTM28 genome contained a gene copy of mmpL8, which encodes a product required for the synthesis of sulfolipid-1 (SL-1), a compound that is able to prevent the fusion of phagosome with lysosome to form the phagolysosome in macrophages. It also blocks oxidative phosphorylation and inhibits the production of reactive oxygen36. Yet, unlike M. abscessus, the TNTM28 genome lacked genes encoding phospholipase C-type enzymes, whose function in certain SGM has previously been proposed as virulence factor37, a hypothesis that has recently been dismissed38.
Assessment of TNTM28 intramacrophagic growth
The particular phenotypic and genomic features of TNTM28, which all converged towards its pathogenicity have promted us to assess its ability to replicate in macrophages. As shown in Fig. 6, TNTM28 replicated efficiently in PMA-differentiated THP-1 macrophages, with titers comparable to smooth and rough variants of M. abscessus.
Discussion
In the present study we described the genome sequence of TNTM28, a new rapidly growing NTM species. Several features might indicate that this species could be endowed with a virulence potential. Firstly, it has been recovered from a patient presenting with a typical pulmonary disease. Review of the patient's clinical record by taking into account the American Thoracic Society (ATS)/Infectious Disease Society of America (IDSA) guidelines, argued for a true NTM pulmonary disease39. Secondly, TNTM28 displayed two main phenotypic features reminiscent of pathogenic mycobacteria. Indeed, TNTM28 colonies showed a rough morphotype, and Ziehl–Neelsen-stained cells tended to form large aggregates. Both characteristics have been associated with persistence inside phagocytes, which is most likely linked to the bacillus’s ability to escape from phagocytosis5,39. Another argument for an enhanced level of pathogenicity in TNTM28 was its ability to replicate in macrophages equally well as M. abscessus, one of the most pathogenic RGM species40,41.
Phylogenetic and phylogenomic analyses linked TNTM28 to the M. fortuitum complex. This finding came as no surprise given the ubiquitous presence of M. fortuitum-related species in the environment. Furthermore, M. fortuitum encompasses a large group of emerging opportunistic pathogens whose members have frequently been associated with pathogenic conditions in both immunocompetent and immunocompromised individuals11. Of particular interest, this new species, which has derived immediately from the common ancestor of all members of the M. fortuitum complex, displayed a smaller genome size compared to other pathogenic RGM (~ 1 Mb difference), such as M. fortuitum and M. abscessus complexes. Such a genomic reduction could have tightened the parasitic lifestyle of TNTM28, thereby enhancing its pathogenicity42.
As detailed in the Results section, the genome of TNTM28 contained a range of potential virulence genes, which might have promoted its ability for intracellular replication and persistence. Indeed, compared to other RGMs, TNTM28 specified numerous genes whose homologues exist essentially in pathogenic mycobacteria and are thus worthy of consideration. Among these was the mgtC gene, a key player in intramacrophage survival, being important for virulence in diverse intracellular pathogens43,44. This gene is usually absent from the genomes of several RGM, such as M. fortuitum, M. smegmatis, M. gilvum, etc., but like TNTM28, this gene does exist in the genome of M. abscessus. This finding is consistent with the fact that the latter two species were found to replicate equally well in macrophages. Furthermore, the genome of TNTM28 harbored a copy of the clgR gene, which has been shown in M. tuberculosis to be involved in the modulation of phagosome maturation45, and which was lacking in the majority of RGM, with the exception of M. smegmatis. In M. tuberculosis, ClgR activates the transcription of genes encoding a larger network of protein homeostatic and regulatory systems. ClgR-regulated transcriptional activation of these systems is essential for M. tuberculosis to replicate in macrophages by enabling the bacillus to control the phagosome pH45. By contrast, TNTM28 lacked the sapM gene, which encodes a secretory acid phosphatase, and whose disruption in M. tuberculosis translated into the inability of the mutant to arrest the phagosomal maturation with a severe growth defect in THP-1 macrophages46. Among other genes of importance was trpD, which encodes an anthranilate phosphoribosyltransferase involved in tryptophan biosynthesis. In SGM, trpD has been shown to play important roles during infection47. Aside from TNTM28, only M. abscessus was found to contain a gene copy of trpD. Interestingly, this gene proved essential for lung colonization by M. tuberculosis in mice48. Another distinctive feature of TNTM28, yet shared with M. fortuitum only, was the existence of a treS gene, which encodes a trehalose synthase enzyme, converting trehalose into maltose and vice versa. Deletion of the treS gene was shown to significantly prolonge the time to death in a chronic infection model in mice49.
Additional genes allowing TNTM28 to cope with the harsh intramacrophagic environment appear to have been brought by HGT. At least seven genes (murB, lprQ, fhaB, yidC, yidD, fgD2, and ispH1) involved in cell wall biogenesis might have been transferred to TNTM28, as such. These genes are of particular interest given the critical role played by the mycobacterial cell wall, whose extraordinarily complex nature significantly contribute to the ability of pathogenic mycobacteria to manipulate and evade human immune system50,51. Furthermore, and considering the vital role of cholesterol for optimal growth and persistence within the host52,53,54, the transfer of hsaD, a gene encoding a hydrolase involved in cholesterol catabolism, is likely to be of significance for TNTM28 virulence. Indeed, HsaD proved essential for survival of M. tuberculosis inside macrophages55. Moreover, and with regard to the cholesterol metabolism, it is worth mentioning that TNTM28 genome contained nearly the full set of mce genes, including the mce4 complex, which has been shown to be essential for cholesterol import56,57.
Among other notable putatively transferred genes that might have increased TNTM28 resistance to macrophage defensive arsenals was mutT2, given its pivotal role in protecting the bacillus against reactive oxygen species58.
Compared to the majority of RGM, TNTM28 harbored a near-complete ESX-1 locus, which could have endowed it with a relatively enhanced virulence, although the role of ESX-1 in RGM species such as M. smegmatis is rather linked to horizontal gene transfer than to virulence59. The ESX-1 region, along with ESX-3 and ESX-4 are ancestral regions and were thus found in the genomes of most mycobacteria60. The role of ESX-1 in survival in the macrophage and the overall bacillus’s pathogenicity has been largely demonstrated for MTBC members and slow growing mycobacteria, whose genome encode a complete ESX-1 system, as well as an associated espACD operon, which was not found in RGM so far61,62,63. It is worth noting that we have identified in TNTM28, but neither in M. abscessus nor in M. smegmatis, a locus structurally similar to espACD, and which, most intriguingly, was found to encode for an ortholog of EspC, the main modulator of ESX-1 function24. Therefore, it remains to be seen whether such an operon, does serve the same function(s) as the espACD operon of pathogenic mycobacteria. Should it be the case, this may also compensate for the lack in TNTM28 of the gene encoding Eis N-acetyl transferase protein, whose deletion mutant in M. abscessus proved strongly attenuated in macrophages64. Moreover, TNTM28 contained an ESX-4 copy, which in the absence of ESX-1, was shown to play a prominent role in M. abscessus growth in vivo27.
In summary, we identified a new RGM species displaying several phenotypic and genomic hallmarks that argue for its pathogenicity. Acquisition of TNTM28 virulence traits seemed to have benefited of a highly permissive environment for gene exchange, thereby favoring transition to pahogenicity. Because TNTM28 deep rooting in the phylogenetic tree, compared to the Mycobacterium fortuitum complex, being the unique representative of a newly derived branch, we propose a new taxonomic entity with the provisional name “Mycobacterium fortunisiensis sp. Nov”, which also refers to Tunis, the origin of isolation.
Methods
De novo sequencing and assembly
Genomic DNA (gDNA) was extracted using standard phenol-choloroform method and was sequenced on the HiSeq2500 Technology (Illumina Inc., San Diego, CA, USA) with paired-end application. The gDNA was quantified using Quant-iT™ PicoGreen® ds DNA reagent, (Invitrogen, CA, USA). Paired-end library was constructed using "NEXTflex PCRFree Kit" according to Nextflex Illumina protocol. Automated cluster generation and paired end sequencing with dual index reads were performed in a single run in 2 × 107-pb. The 5.469.922 paired-end reads were firstly processed with FastQC and Trimmomatic sotwares65 before de novo genome assembling.
Paired-end reads were assembled using SPAdes genome assembler v.3.10.166. Illumina reads were re-mapped into the scaffolds using the paired-end mode of Bwa mem v0.7.467 with default parameters. After converting output SAM files to BAM files by SAMtools68, coverage mapping was computed by BEDtools v2.17.069.
Filtered Scaffolds were ordered and oriented using CONTIGuator 2.7.470 using M. fortuitum CT6, complete genome (Genbank CP011269) as reference in order to distinguish chromosome scaffolds and unplaced scaffolds. We used progressiveMauve for multiple genome alignment71.
Genome annotation and phylogenetic analyses
Functionnal annotation was performed using the Prokaryotic Genome Annotation System (Prokka) v1.12 pipeline72. CRISPR loci were searched for using the CRISPRfinder program online73 (last update, 2017-05-09). The genomic circular representation of TNTM28 was generated using CGView server (http://stothard.afns.ualberta.ca/cgview_server/) and putative prophages were found using PHASTER (PhAge Search Tool) software74 based on the actinobacteriophage Database at phageDB.org and the online plasmid search tool http://plasmid.med.havard.edu/PLASMID/home.xhtml.
The prediction of tRNA was processed using ARAGORN program75, whereas ribosomal RNAs were predicted using RNAmmer76.
We performed functional annotation of genes using the Clusters of Orthologous Groups (COGs) database (http://www.ncbi.nlm.nih.gov/COG) using BLASTP (E value < 1e−5 and > 50% coverage).
The Phylogenomic tree was constructed using an identity matrix based on Mauve software (http://gel.ahabs.wisc.edu/mauve) genome alignment71. OrthoANI values were calculated between TNTM28 and 9 sequenced Mycobacterium species genomes using Orthologous Average Nucleotide Identity Tool77. Putative virulence genes were found using Pathogen Finder tool23 against the VFDB database (http://www.mgc.ac.cn/VFs/).
Horizontally transferred Genomic Islands (GIs) were identified using IslandPahth-DIMOB78.
Hierarchical clustering of closely related sets of virulence genes was generated after z-score normalization of the data using Euclidean distance.
Pan genome analysis
The pan genome analysis, the core accessory, and unique genes was performed using the Bacterial Pan Genome Analysis Tool (BPGA)79.
ANI and DDH values were calculated using the GGDC version 2.0 online tool80.
In vitro growth assessment of TNTM28 in THP-1 derived macrophages
THP-1 human monocyte-like cells (TIB-202D) were purchased from the American type culture collection (ATCC), directly amplified and stored in liquid nitrogen. Only low passage cells (number of passages < 11) were used in the experiments. The purchased THP-1 cell line was authenticated and tested against microbial contaminants, including mycoplasmas, by ATCC.
THP-1 cells were grown in RPMI 1640, GlutaMAX (Life Technologies) containing 10% heat-inactivated fetal bovine serum (Life Technologies), seeded at a density of 7.5 × 104 cells per well in 96-well plates and differentiated into macrophages by incubation with 50 mM phorbol-myristate-acetate (PMA) for 3 days.
For infection, bacteria cultured in Sauton medium without agitation were sonicated, added to macrophages at a multiplicity of infection (MOI) of 0.05 (~ 1 bacteria per 20 THP cells), and incubated for 2 h. Sauton medium was used because it allows the production of more complex polar lipids81. After phagocytosis, 0.1 mg/ml of amikacin was added for one hour to eliminate extracellular bacteria and cells were incubated for up to 6 days at 37 °C and 5% CO2. At various times, the macrophages were lysed with 0.1% Triton-X100 in PBS and the lysates were plated in serial dilutions on 7H11 + OADC plates to determine the intracellular survival of the bacteria in c.f.u. The experiments were performed at least four biological replicas, each in triplicate (technical replicas).
Data availability
The rpoB, hsp65, 16 S rRNA and sodA gene sequences of strain TNTM28 was deposited in GenBank under accession number MK762879, MK751438, MK630280 and MK778075, respectively. Illumina reads for TNTM28 have been deposited at GenBank under accession number VOMB00000000. All data generated and analysed during this study are included in this manuscript and its supplementary information files.
References
Chan, E. D. & Iseman, M. D. Underlying host risk factors for nontuberculous mycobacterial lung disease. Semin. Respir. Crit. Care Med 34, 110–123 (2013).
Jenkins, P. A., Campbell, I. A. & Research Committee of The British Thoracic Society. Pulmonary disease caused by Mycobacterium xenopi in HIV-negative patients: Five year follow-up of patients receiving standardised treatment. Respir. Med. 97, 439–444 (2003).
Cook, J. L. Nontuberculous mycobacteria: Opportunistic environmental pathogens for predisposed hosts. Br. Med. Bull. 96, 45–59 (2010).
Johansen, M. D., Herrmann, J. L. & Kremer, L. Non-tuberculous mycobacteria and the rise of Mycobacterium abscessus. Nat. Rev. Microbiol. 18, 392–407 (2020).
Johnson, M. M. & Odell, J. A. Nontuberculous mycobacterial pulmonary infections. J. Thorac. Dis. 6, 210–220 (2014).
Jarzembowski, J. A. & Young, M. B. Nontuberculous mycobacterial infections. Arch. Pathol. Lab. Med. 132, 1333–1341 (2008).
Leclerc, M. C., Thomas, F. & Guégan, J. F. Evidence for phylogenetic inheritance in pathogenicity of Mycobacterium. Antonie Van Leeuwenhoek 83, 265–274 (2003).
De Groote, M. A. & Huitt, G. Infections due to rapidly growing mycobacteria. Clin. Infect. Dis. 42, 1756–1763 (2006).
Falkinham, J. O. Nontuberculous mycobacteria in the environment. Clin. Chest Med. 23, 529–551 (2002).
Brown-Elliott, B. A. & Wallace, R. J. Jr. Clinical and taxonomic status of pathogenic nonpigmented or late-pigmenting rapidly growing mycobacteria. Clin. Microbiol. Rev. 15, 716–746 (2002).
Wallace, R. J. Jr., Brown, B. A. & Griffith, D. E. Nosocomial outbreaks/pseudo-outbreaks caused by nontuberculous mycobacteria. Annu. Rev. Microbiol. 52, 453–490 (1998).
Schinsky, M. F. et al. Mycobacterium septicum sp. nov., a new rapidly growing species associated with catheter-related bacteraemia. Int. J. Syst. Evol. Microbiol. 50, 575–581 (2000).
Schinsky, M. F. et al. Taxonomic variation in the Mycobacterium fortuitum third biovariant complex: Description of Mycobacterium boenickei sp. nov., Mycobacterium houstonense sp. nov., Mycobacterium neworleansense sp. nov. and Mycobacterium brisbanense sp. nov. and recognition of Mycobacterium porcinum from human clinical isolates. Int. J. Syst. Evol. Microbiol. 54, 1653–1667 (2004).
Gupta, R. S., Lo, B. & Son, J. Phylogenomics and comparative genomic studies robustly support division of the genus Mycobacterium into an emended genus Mycobacterium and four novel genera. Front. Microbiol. 9, 67 (2018).
Erber, J. et al. Successful bedaquiline-containing antimycobacterial treatment in post-traumatic skin and soft-tissue infection by Mycobacterium fortuitum complex: A case report. BMC Infect. Dis. 20, 365 (2020).
Diaz, M., Huff, T. N. & Libertin, C. R. Nontuberculous mycobacterial infections of the lower extremities: A 15-year experience. J. Clin. Tuberc. Other Mycobact. Dis. 15, 100091 (2019).
Park, S. et al. Clinical significance of Mycobacterium fortuitum isolated from respiratory specimens. Respir. Med. 102, 437–442 (2008).
Okamori, S. et al. Natural history of Mycobacterium fortuitum pulmonary infection presenting with migratory infiltrates: A case report with microbiological analysis. BMC Infect. Dis. 18, 1 (2018).
Radzniwan, M. R., Tohid, H., Ahmad, S., Mohd, A. F. & Md Anshar, F. Isolation of Mycobacterium fortuitum in sputum specimens of a patient with chronic cough: Is it clinically significant?. Malays. Fam. Physician 9, 38–41 (2014).
Gharbi, R., Mhenni, B., Ben Fraj, S. & Mardassi, H. Nontuberculous mycobacteria isolated from specimens of pulmonary tuberculosis suspects, Northern Tunisia: 2002–2016. BMC Infect. Dis. 19, 819 (2019).
Bertelli, C. et al. IslandViewer 4: Expanded prediction of genomic islands for larger-scale datasets. Nucleic Acids Res. 45, W30–W35 (2017).
Chen, L. et al. VFDB: A reference database for bacterial virulence factors. Nucleic Acids Res. 33, D325–D328 (2005).
Cosentino, S., Voldby Larsen, M., Møller Aarestrup, F. & Lund, O. PathogenFinder–distinguishing friend from foe using bacterial whole genome sequence data. PLoS ONE 8, e77302 (2013).
Lou, Y., Rybniker, J., Sala, C. & Cole, S. T. EspC forms a filamentous structure in the cell envelope of Mycobacterium tuberculosis and impacts ESX-1 secretion. Mol. Microb. 103, 26–38 (2017).
Gey Van Pittius, N. C. et al. The ESAT-6 gene cluster of Mycobacterium tuberculosis and other high G+C Gram-positive bacteria. Genome Biol. 2, 1–18 (2001).
Dumas, E. et al. Mycobacterial pan-genome analysis suggests important role of plasmids in the radiation of type VII secretion systems. Genome Biol. Evol. 8, 387–402 (2016).
Laencina, L. et al. Identification of genes required for Mycobacterium abscessus growth in vivo with a prominent role of the ESX-4 locus. Proc. Natl. Acad. Sci. USA. 115, E1002–E1011 (2018).
Zhang, F. & Xie, J. P. Mammalian cell entry gene family of Mycobacterium tuberculosis. Mol. Cell. Biochem. 352, 1–10 (2011).
Griffin, J. E. et al. High-resolution phenotypic profiling defines genes essential for mycobacterial growth and cholesterol catabolism. PLoS Pathog. 7, e1002251 (2011).
Casali, N. & Riley, L. W. A phylogenomic analysis of the Actinomycetales mce operons. BMC Genomics 8, 60 (2007).
Fedrizzi, T. et al. Genomic characterization of nontuberculous mycobacteria. Sci. Rep. 7, 45258 (2017).
Simeone, R., Bottai, D., Frigui, W., Majlessi, L. & Brosch, R. ESX/type VII secretion systems of mycobacteria: Insights into evolution, pathogenicity and protection. Tuberculosis 95, S150–S154 (2015).
Lee, V. T. & Schneewind, O. Protein secretion and the pathogenesis of bacterial infections. Genes Dev. 15, 1725–1752 (2001).
Cole, S. T. et al. Deciphering the biology of Mycobacterium tuberculosis from the complete genome sequence. Nature 393, 537–544 (1998).
Stinear, T. P. et al. Insights from the complete genome sequence of Mycobacterium marinum on the evolution of Mycobacterium tuberculosis. Genome Res. 18, 729–741 (2008).
Converse, S. E. et al. MmpL8 is required for sulfolipid-1 biosynthesis and Mycobacterium tuberculosis virulence. Proc. Natl. Acad. Sci. USA. 100, 6121–6126 (2003).
Raynaud, C. et al. Phospholipases C are involved in the virulence of Mycobacterium tuberculosis. Mol Microbiol. 45, 203–217 (2002).
Le Chevalier, F. et al. Revisiting the role of phospholipases C in virulence and the lifecycle of Mycobacterium tuberculosis. Sci. Rep. 5, 16918 (2015).
Griffith, D. E. et al. An official ATS/IDSA statement: Diagnosis, treatment, and prevention of nontuberculous mycobacterial diseases. Am. J. Respir. Crit. Care Med. 175, 367–416 (2007).
Jönsson, B., Ridell, M. & Wold, A. E. Phagocytosis and cytokine response to rough and smooth colony variants of Mycobacterium abscessus by human peripheral blood mononuclear cells. APMIS 121, 45–55 (2013).
Choo, S. W. et al. Genomic reconnaissance of clinical isolates of emerging human pathogen Mycobacterium abscessus reveals high evolutionary potential. Sci. Rep. 4, 4061 (2014).
Moran, N. A. Microbial minimalism: Genome reduction in bacterial pathogens. Cell 108, 583–586 (2002).
Buchmeier, N. et al. A parallel intraphagosomal survival strategy shared by Mycobacterium tuberculosis and Salmonella enterica. Mol. Microbiol. 35, 1375–1382 (2000).
Alix, E. & Blanc-Potard, A. B. MgtC: A key player in intramacrophage survival. Trends Microbiol. 15, 252–256 (2007).
Estorninho, M. et al. ClgR regulation of chaperone and protease systems is essential for Mycobacterium tuberculosis parasitism of the macrophage. Microbiology 156, 3445–3455 (2010).
Puri, R. V., Reddy, P. V. & Tyagi, A. K. Secreted acid phosphatase (SapM) of Mycobacterium tuberculosis is indispensable for arresting phagosomal maturation and growth of the pathogen in guinea pig tissues. PLoS ONE 8, e70514 (2013).
Zhang, Y. J. et al. Tryptophan biosynthesis protects mycobacteria from CD4 T-cell-mediated killing. Cell 155, 1296–1308 (2013).
Lee, C. E., Goodfellow, C., Javid-Majd, F., Baker, E. N. & Shaun Lott, J. The crystal structure of TrpD, a metabolic enzyme essential for lung colonization by Mycobacterium tuberculosis, in complex with its substrate phosphoribosylpyrophosphate. J. Mol. Biol. 355, 784–797 (2006).
Murphy, H. N. et al. The OtsAB pathway is essential for trehalose biosynthesis in Mycobacterium tuberculosis. J. Biol. Chem. 280, 14524–14529 (2005).
Guenin-Macé, L., Siméone, R. & Demangel, C. Lipids of pathogenic Mycobacteria: Contributions to virulence and host immune suppression. Transbound. Emerg. Dis. 56, 255–268 (2009).
Ouellet, H., Johnston, J. B. & de Montellano, P. R. Cholesterol catabolism as a therapeutic target in Mycobacterium tuberculosis. Trends Microbiol. 19, 530–539 (2011).
Jankute, M., Cox, J. A., Harrison, J. & Besra, G. S. Assembly of the mycobacterial cell wall. Annu. Rev. Microbiol. 69, 405–423 (2015).
Rohde, K. H., Veiga, D. F., Caldwell, S., Balázsi, G. & Russell, D. G. Linking the transcriptional profiles and the physiological states of Mycobacterium tuberculosis during an extended intracellular infection. PLoS Pathog. 8, e1002769 (2012).
Wilburn, K. M., Fieweger, R. A. & VanderVen, B. C. Cholesterol and fatty acids grease the wheels of Mycobacterium tuberculosis pathogenesis. Pathog. Dis. 76, fty021 (2018).
Ryan, A. et al. Mechanism-based inhibition of HsaD: A C-C bond hydrolase essential for survival of Mycobacterium tuberculosis in macrophage. FEMS Microbiol. Lett. 350, 42–47 (2014).
Pandey, A. K. & Sassetti, C. M. Mycobacterial persistence requires the utilization of host cholesterol. Proc. Natl. Acad. Sci. USA 105, 4376–4380 (2008).
Nazarova, E. V. et al. Rv3723/LucA coordinates fatty acid and cholesterol uptake in Mycobacterium tuberculosis. Elife 6, e26969 (2017).
Sang, P. B. & Varshney, U. Biochemical properties of MutT2 proteins from Mycobacterium tuberculosis and M. smegmatis and their contrasting antimutator roles in Escherichia coli. J. Bacterial. 195, 1552–1560 (2013).
Derbyshire, K. M. & Gray, T. A. Distributive conjugal transfer: New insights into horizontal gene transfer and genetic exchange in mycobacteria. Microbiol. Spectr. 2, 04 (2014).
Gcebe, N., Michel, A., Gey van Pittius, N. C. & Rutten, V. Comparative genomics and proteomic analysis of four non-tuberculous Mycobacterium Species and Mycobacterium tuberculosis complex: Occurrence of shared immunogenic proteins. Front. Microbiol. 7, 795 (2016).
Abdallah, A. M. et al. Type VII secretion–mycobacteria show the way. Nat. Rev. Microbiol. 5, 883–891 (2007).
Houben, E. N., Korotkov, K. V. & Bitter, W. Take five—Type VII secretion systems of Mycobacteria. Biochim. Biophys. Acta 1843, 1707–1716 (2014).
Gröschel, M. I., Sayes, F., Simeone, R., Majlessi, L. & Brosch, R. ESX secretion systems: Mycobacterial evolution to counter host immunity. Nat. Rev. Microbiol. 14, 677–691 (2016).
Dubois, V. et al. Mycobacterium abscessus virulence traits unraveled by transcriptomic profiling in amoeba and macrophages. PLoS Pathog. 15, e1008069 (2019).
Bolger, A. M., Lohse, M. & Usadel, B. Trimmomatic: A flexible trimmer for Illumina sequence data. Bioinformatics 30, 2114–2120 (2014).
Bankevich, A. et al. SPAdes: A new genome assembly algorithm and its applications to single-cell sequencing. J. Comput. Biol. 19, 455–477 (2012).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 25, 589–595 (2009).
Li, H. et al. The Sequence alignment/map (SAM) format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Quinlan, A. R. & Hall, I. M. BEDTools: A flexible suite of utilities for comparing genomic features. Bioinformatics 26, 841–842 (2010).
Galardini, M., Biondi, E. G., Bazzicalupo, M. & Mengoni, A. CONTIGuator: A bacterial genomes finishing tool for structural insights on draft genomes. Source Code Biol. Med. 6, 11 (2011).
Darling, A. C., Mau, B., Blattner, F. R. & Perna, N. T. Mauve: Multiple alignment of conserved genomic sequence with rearrangements. Genome Res. 14, 1394–1403 (2004).
Seemann, T. Prokka: Rapid prokaryotic genome annotation. Bioinformatics 30, 2068–2069 (2014).
Grissa, I., Vergnaud, G. & Pourcel, C. CRISPRFinder: A web tool to identify clustered regularly interspaced short palindromic repeats. Nucleic Acids Res. 35, W52–W57 (2007).
Arndt, D. et al. PHASTER: A better, faster version of the PHAST phage search tool. Nucleic Acids Res. 44, W16–W21 (2016).
Laslett, D. & Canback, B. ARAGORN, a program to detect tRNA genes and tmRNA genes in nucleotide sequences. Nucleic Acids Res. 32, 11–16 (2004).
Lagesen, K. et al. RNAmmer: Consistent and rapid annotation of ribosomal RNA genes. Nucleic Acids Res. 35, 3100–3108 (2007).
Lee, I., Ouk Kim, Y., Park, S. C. & Chun, J. OrthoANI: An improved algorithm and software for calculating average nucleotide identity. Int. J. Syst. Evol. Microbiol. 66, 1100–1103 (2016).
Bertelli, C. & Brinkman, F. S. L. Improved genomic island predictions with IslandPath-DIMOB. Bioinformatics 34, 2161–2167 (2018).
Chaudhari, N. M., Gupta, V. K. & Dutta, C. BPGA- an ultra-fast pan-genome analysis pipeline. Sci. Rep. 6, 24373 (2016).
Meier-Kolthoff, J. P., Auch, A. F., Klenk, H. P. & Göker, M. Genome sequence-based species delimitation with confidence intervals and improved distance functions. BMC Bioinform. 14, 60 (2013).
Burguière, A. et al. LosA, a key glycosyltransferase involved in the biosynthesis of a novel family of glycosylated acyltrehalose lipooligosaccharides from Mycobacterium marinum. J. Biol. Chem. 280, 42124–42133 (2005).
Acknowledgements
We thank Christiane Bouchier and Laurence Ma, from the Biomics platform of the Institut Pasteur, Paris, for library construction and genome sequencing, and Sinda Zarouk from Institut Pasteur, Tunis, for sample logistics.
This study was funded by the Tunisian Ministry of Higher Education and Scientific Research (LR16IPT01).
Author information
Authors and Affiliations
Contributions
H.M. conceived and supervised the study. R.G. and H.M. designed the experiments. R.G., B.M., and H.M. were involved in initial isolation and identification of TNTM28, the described new mycobacterial species. W.F. and R.B. performed the intramacrophagic growth assays. R.G. and V.K. performed de novo sequencing and data curation. R.G., V.K., R.B., and H.M., analyzed the data. R.G. and H.M. wrote the manuscript. R.G., V.K., W.F., R.B., and H.M. reviewed and edited the manuscript. All authors have read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Gharbi, R., Khanna, V., Frigui, W. et al. Phenotypic and genomic hallmarks of a novel, potentially pathogenic rapidly growing Mycobacterium species related to the Mycobacterium fortuitum complex. Sci Rep 11, 13011 (2021). https://doi.org/10.1038/s41598-021-91737-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-021-91737-8
This article is cited by
-
A nontuberculous mycobacterium could solve the mystery of the lady from the Franciscan church in Basel, Switzerland
BMC Biology (2023)
-
Prosthetic joint infection caused by an imipenem-resistant Mycobacterium senegalense
Brazilian Journal of Microbiology (2023)
-
Antibiotic delivery evaluation against Mycobacterium fortuitum using nanofluids containing carbon nanotubes
BMC Microbiology (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.