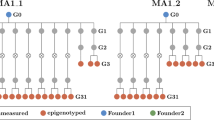
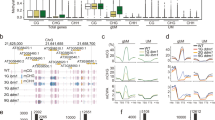
Abstract
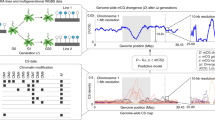
Heritable epigenetic polymorphisms, such as differential cytosine methylation, can underlie phenotypic variation1,2. Moreover, wild strains of the plant Arabidopsis thaliana differ in many epialleles3,4, and these can influence the expression of nearby genes1,2. However, to understand their role in evolution5, it is imperative to ascertain the emergence rate and stability of epialleles, including those that are not due to structural variation. We have compared genome-wide DNA methylation among 10 A. thaliana lines, derived 30 generations ago from a common ancestor6. Epimutations at individual positions were easily detected, and close to 30,000 cytosines in each strain were differentially methylated. In contrast, larger regions of contiguous methylation were much more stable, and the frequency of changes was in the same low range as that of DNA mutations7. Like individual positions, the same regions were often affected by differential methylation in independent lines, with evidence for recurrent cycles of forward and reverse mutations. Transposable elements and short interfering RNAs have been causally linked to DNA methylation8. In agreement, differentially methylated sites were farther from transposable elements and showed less association with short interfering RNA expression than invariant positions. The biased distribution and frequent reversion of epimutations have important implications for the potential contribution of sequence-independent epialleles to plant evolution.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Accession codes
Data deposits
DNA sequencing data are deposited in the European Nucleotide Archive (http://www.ebi.ac.uk/ena/data/view/ERP000902) under accession number ERP000902.
References
Richards, E. J. Population epigenetics. Curr. Opin. Genet. Dev. 18, 221–226 (2008)
Paszkowski, J. & Grossniklaus, U. Selected aspects of transgenerational epigenetic inheritance and resetting in plants. Curr. Opin. Plant Biol. 14, 195–203 (2011)
Vaughn, M. W. et al. Epigenetic natural variation in Arabidopsis thaliana. PLoS Biol. 5, e174 (2007)
Rangwala, S. H. et al. Meiotically stable natural epialleles of Sadhu, a novel Arabidopsis retroposon. PLoS Genet. 2, e36 (2006)
Slatkin, M. Epigenetic inheritance and the missing heritability problem. Genetics 182, 845–850 (2009)
Shaw, R. G., Byers, D. L. & Darmo, E. Spontaneous mutational effects on reproductive traits of Arabidopsis thaliana. Genetics 155, 369–378 (2000)
Ossowski, S. et al. The rate and molecular spectrum of spontaneous mutations in Arabidopsis thaliana. Science 327, 92–94 (2010)
Law, J. A. & Jacobsen, S. E. Establishing, maintaining and modifying DNA methylation patterns in plants and animals. Nature Rev. Genet. 11, 204–220 (2010)
Teixeira, F. K. et al. A role for RNAi in the selective correction of DNA methylation defects. Science 323, 1600–1604 (2009)
Reinders, J. et al. Compromised stability of DNA methylation and transposon immobilization in mosaic Arabidopsis epigenomes. Genes Dev. 23, 939–950 (2009)
Widman, N., Jacobsen, S. E. & Pellegrini, M. Determining the conservation of DNA methylation in Arabidopsis. Epigenetics 4, 119–124 (2009)
Lynch, M. Evolution of the mutation rate. Trends Genet. 26, 345–352 (2010)
Cokus, S. J. et al. Shotgun bisulphite sequencing of the Arabidopsis genome reveals DNA methylation patterning. Nature 452, 215–219 (2008)
Lister, R. et al. Highly integrated single-base resolution maps of the epigenome in Arabidopsis. Cell 133, 523–536 (2008)
Zhang, Y. et al. QDMR: a quantitative method for identification of differentially methylated regions by entropy. Nucleic Acids Res. 39, e58 (2011)
Zilberman, D., Gehring, M., Tran, R. K., Ballinger, T. & Henikoff, S. Genome-wide analysis of Arabidopsis thaliana DNA methylation uncovers an interdependence between methylation and transcription. Nature Genet. 39, 61–69 (2007)
Zhang, X. et al. Genome-wide high-resolution mapping and functional analysis of DNA methylation in Arabidopsis. Cell 126, 1189–1201 (2006)
Fahlgren, N. et al. MicroRNA gene evolution in Arabidopsis lyrata and Arabidopsis thaliana. Plant Cell 22, 1074–1089 (2010)
Kasschau, K. D. et al. Genome-wide profiling and analysis of Arabidopsis siRNAs. PLoS Biol. 5, e57 (2007)
Slotkin, R. K. et al. Epigenetic reprogramming and small RNA silencing of transposable elements in pollen. Cell 136, 461–472 (2009)
Lisch, D. Epigenetic regulation of transposable elements in plants. Annu. Rev. Plant Biol. 60, 43–66 (2009)
Pagnussat, G. C. et al. Genetic and molecular identification of genes required for female gametophyte development and function in Arabidopsis. Development 132, 603–614 (2005)
Cao, J. et al. Whole-genome sequencing of multiple Arabidopsis thaliana populations. Nature Genet. 10.1038/ng.911 (28 August 2011)
Tran, R. K. et al. DNA methylation profiling identifies CG methylation clusters in Arabidopsis genes. Curr. Biol. 15, 154–159 (2005)
Mirouze, M. et al. Selective epigenetic control of retrotransposition in Arabidopsis. Nature 461, 427–430 (2009)
Chodavarapu, R. K. et al. Relationship between nucleosome positioning and DNA methylation. Nature 466, 388–392 (2010)
Laurent, L. et al. Dynamic changes in the human methylome during differentiation. Genome Res. 20, 320–331 (2010)
Hollister, J. D. et al. Transposable elements and small RNAs contribute to gene expression divergence between Arabidopsis thaliana and Arabidopsis lyrata. Proc. Natl Acad. Sci. USA 108, 2322–2327 (2011)
Ossowski, S. et al. Sequencing of natural strains of Arabidopsis thaliana with short reads. Genome Res. 18, 2024–2033 (2008)
Schneeberger, K. et al. Simultaneous alignment of short reads against multiple genomes. Genome Biol. 10, R98 (2009)
Storey, J. D. & Tibshirani, R. Statistical significance for genomewide studies. Proc. Natl Acad. Sci. USA 100, 9440–9445 (2003)
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 25, 1754–1760 (2009)
Robinson, M. D., McCarthy, D. J. & Smyth, G. K. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26, 139–140 (2010)
Benjamini, Y. & Yekutieli, D. The control of the false discovery rate in multiple testing under dependency. Ann. Stat. 29, 1165–1188 (2001)
Wickham, H. ggplot2: Elegant Graphics for Data Analysis (Springer, 2009)
Acknowledgements
We are grateful to R. Shaw for providing seeds of the mutation accumulation lines. We thank C. Lanz for help with Illumina sequencing, J. Fitz for help with the Gbrowse implementation, K. Schneeberger for help with DNA mutation analysis, D. Zilberman for providing bisulphite sequencing protocols, and R. Clark, M. Pellegrini, R. Neher and J. Paszkowski for discussion. This work was supported by an HFSP Long-Term Fellowship (D.K.), the Volkswagen Foundation (O.S.), FP7 Collaborative Project AENEAS (contract KBBE-2009-226477), a Gottfried Wilhelm Leibniz Award of the DFG, and the Max Planck Society (D.W.).
Author information
Authors and Affiliations
Contributions
C.B., J.H. and D.W. conceived the study; C.B. performed the experiments; C.B., J.H., J.M., D.K. and O.S. analysed the data; K.B. provided advice on statistical analysis; and C.B. and D.W. wrote the paper with contributions from all authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Figures
This file contains Supplementary Figures 1-13 with legends and legends for Supplementary Tables 1-4. (PDF 3395 kb)
Supplementary Tables
This zipped file contains Supplementary Tables 1-4 (see Supplementary Information file for legends). (ZIP 3052 kb)
Rights and permissions
About this article
Cite this article
Becker, C., Hagmann, J., Müller, J. et al. Spontaneous epigenetic variation in the Arabidopsis thaliana methylome. Nature 480, 245–249 (2011). https://doi.org/10.1038/nature10555
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature10555
This article is cited by
-
Identification of epigenetically regulated genes involved in plant-virus interaction and their role in virus-triggered induced resistance
BMC Plant Biology (2024)
-
Comparative genetic and epigenetic of the Sphagneticola trilobata (L.) Pruski from different regions in China
BMC Plant Biology (2023)
-
DDM1-mediated gene body DNA methylation is associated with inducible activation of defense-related genes in Arabidopsis
Genome Biology (2023)
-
Cisplatin exposure alters tRNA-derived small RNAs but does not affect epimutations in C. elegans
BMC Biology (2023)
-
Re-analysis of publicly available methylomes using signal detection yields new information
Scientific Reports (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.