Abstract
Technological advances in DNA recovery and sequencing have drastically expanded the scope of genetic analyses of ancient specimens to the extent that full genomic investigations are now feasible and are quickly becoming standard1. This trend has important implications for infectious disease research because genomic data from ancient microbes may help to elucidate mechanisms of pathogen evolution and adaptation for emerging and re-emerging infections. Here we report a reconstructed ancient genome of Yersinia pestis at 30-fold average coverage from Black Death victims securely dated to episodes of pestilence-associated mortality in London, England, 1348–1350. Genetic architecture and phylogenetic analysis indicate that the ancient organism is ancestral to most extant strains and sits very close to the ancestral node of all Y. pestis commonly associated with human infection. Temporal estimates suggest that the Black Death of 1347–1351 was the main historical event responsible for the introduction and widespread dissemination of the ancestor to all currently circulating Y. pestis strains pathogenic to humans, and further indicates that contemporary Y. pestis epidemics have their origins in the medieval era. Comparisons against modern genomes reveal no unique derived positions in the medieval organism, indicating that the perceived increased virulence of the disease during the Black Death may not have been due to bacterial phenotype. These findings support the notion that factors other than microbial genetics, such as environment, vector dynamics and host susceptibility, should be at the forefront of epidemiological discussions regarding emerging Y. pestis infections.
Similar content being viewed by others
Main
The Black Death of 1347–1351, caused by the bacterium Yersinia pestis2,3, provides one of the best historical examples of an emerging infection with rapid dissemination and high mortality, claiming an estimated 30–50% of the European population in only a five-year period4. Discrepancies in epidemiological trends between the medieval disease and modern Y. pestis infections have ignited controversy over the pandemic’s aetiologic agent5,6. Although ancient DNA investigations have strongly implicated Y. pestis2,3 in the ancient pandemic, genetic changes in the bacterium may be partially responsible for differences in disease manifestation and severity. To understand the organism’s evolution it is necessary to characterize the genetic changes involved in its transformation from a sylvatic pathogen to one capable of pandemic human infection on the scale of the Black Death, and to determine its relationship with currently circulating strains. Here we begin this discussion by presenting the first draft genome sequence of the ancient pathogen.
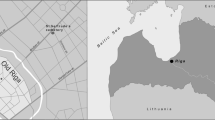
Y. pestis is a recently evolved descendent of the soil-dwelling bacillus Yersinia pseudotuberculosis7, which in the course of its evolution acquired two additional plasmids (pMT1 and pPCP1) that provide it with specialized mechanisms for infiltrating mammalian hosts. To investigate potential evolutionary changes in one of these plasmids, we reported on the screening of 46 teeth and 53 bones from the East Smithfield collection of London, England for presence of the Y. pestis-specific pPCP1 (ref. 3). Historical data indicate that the East Smithfield burial ground was established in late 1348 or early 1349 specifically for interment of Black Death victims8 (Supplementary Figs 1 and 2), making the collection well-suited for genetic investigations of ancient Y. pestis. DNA sequence data for five teeth obtained via molecular capture of the full Y. pestis-specific pPCP1 revealed a C to T damage pattern characteristic of authentic endogenous ancient DNA9, and assembly of the pooled Illumina reads permitted the reconstruction of 98.68% of the 9.6-kilobase plasmid at a minimum of twofold coverage3.
To evaluate the suitability of capture-based methods for reconstructing the complete ancient genome, multiple DNA extracts from both roots and crowns stemming from four of the five teeth which yielded the highest pPCP1 coverage3 were used for array-based enrichment (Agilent) and subsequent high-throughput sequencing on the Illumina GAII platform10. Removal of duplicate molecules and subsequent filtering produced a total of 2,366,647 high quality chromosomal reads (Supplementary Table 1a, b) with an average fragment length of 55.53 base pairs (Supplementary Fig. 4), which is typical for ancient DNA. Coverage estimates yielded an average of 28.2 reads per site for the chromosome, and 35.2 and 31.2 for the pCD1 and pMT1 plasmids, respectively (Fig. 1a, c, d and Supplementary Table 1b, c). Coverage was predictably low for pPCP1 (Fig. 1e) because probes specific to this plasmid were not included on the arrays. Coverage correlated with GC content (Supplementary Fig. 6), a trend previously observed for high-throughput sequence data11. The coverage on each half of the chromosome was uneven due to differences in sequencing depth between the two arrays, with 36.46 and 22.41 average reads per site for array 1 and array 2, respectively. Although greater depth contributed to more average reads per site, it did not increase overall coverage, with both arrays covering 93.48% of the targeted regions at a minimum of onefold coverage (Supplementary Table 1b). This indicates that our capture procedure successfully retrieved template molecules from all genomic regions accessible via this method, and that deeper sequencing would not result in additional data for CO92 template regions not covered in our data set.
a, c–e, Coverage plots for the chromosome (a) and the plasmids pMT1 (c), pCD1 (d) and pPCP (e). Coverage in blue, GC content in green. Scale lines indicate 10-, 20-, 30-, 40- and 50-fold coverage and 10%, 20%, 30%, 40% and 50% GC content. For plasmids, red corresponds to coding regions, yellow to mobile elements. Chromosome shows median coverage per gene. Plasmids show each site plotted. Coverage distributions for the plasmids are shown in Supplementary Fig. 5. b, Distributions show chromosomal coverage of array 1 (blue) and array 2 (red), indicating that deeper sequencing increases the number of reads per site, but does not substantially influence overall coverage.
Genome architecture is known to vary widely among extant Y. pestis strains12. To extrapolate gene order in our ancient genome, we analysed reads mapping to the CO92 reference for all extracts stemming from a single individual who yielded the highest coverage (individual 8291). Despite the short read length of our ancient sequences and the highly repetitive nature of the Y. pestis genome, 2,221 contigs matching CO92 were extracted, comprising a total of 4,367,867 bp. To identify potential regions of the ancient genome that are architecturally distinct from CO92, all reads not mapping to the CO92 reference were in turn considered for contig construction. After filtering for a minimum length of 500 bp, 2,134 contigs remained comprising 4,013,009 bp, of which 30,959 stemmed from unmapped reads. Conventional BLAST search queried against the CO92 genome identified matches for 2,105 contigs. Evidence of altered architecture was identified in 10 contigs (Supplementary Table 2). An example of such a structural variant is shown in Fig. 2, where reference-guided assembly incorporating unmapped reads to span the breakpoint validates its reconstruction. This specific genetic orientation is found only in Y. pseudotuberculosis and Y. pestis strains Mictrotus 91001, Angola, Pestoides F and B42003004, which are ancestral to all Y. pestis commonly associated with human infections (branch 1 and branch 2 strains13,14). Furthermore, discrepancies in the arrangement of this region in branch 1 and branch 2 modern Y. pestis strains indicate that rearrangements occurred as separate events on different lineages.
Reads mapped at positions A (blue) and B (green) are 231 kb apart in the linearized CO92 genome. Adjacent sequence is high coverage although only 18× and 20× is shown due to space constraints (black) for A and B, respectively. The structural variant was assembled using reads that did not map to CO92 (red). Its position is shown on the linearized Microtus 91001 chromosome where the 9,096 bp contig maps with 100% identity.
Single-nucleotide differences between our ancient genome and the CO92 reference surprisingly consisted of only 97 chromosomal positions, and 2 and 4 positions in the pCD1 and pMT1 plasmids, respectively (Supplementary Table 3), indicating tight genetic conservation in this organism over the last 660 years. Twenty-seven of these positions were unreported in a previous analysis of extant Y. pestis diversity14 (Supplementary Tables 3 and 4). Comparison of our ancient genome to its ancestor Y. pseudotuberculosis revealed that the medieval sequence contained the ancestral nucleotide for all 97 positions, indicating that it does not possess any derived positions absent in other Y. pestis strains. Two previously reported chromosomal differences3 were not present in our genomic sequence data, suggesting that they probably derived from deaminated cytosines that would have been removed in the current investigation via uracil-DNA-glycosylase treatment before array capture.
To place our ancient genome in a phylogenetic context, we characterized all 1,694 previously identified phylogenetically informative positions14 (Supplementary Table 4), and compared those from our ancient organism against aggregate base call data for 17 publicly available Y. pestis genomes and the ancestral Y. pseudotuberculosis. When considered separately, sequences from three of the four victims fall only two substitutions from the root of all extant human pathogenic Y. pestis strains (Fig. 3a), and they show a closer relationship to branch 1 Y. pestis than to branch 2; however, one of the four victims (individual 6330) was infected with a strain that contained three additional derived positions seen in all other branch 1 genomes14. This suggests either the presence of multiple strains in the London 1348–1350 pandemic or microevolutionary changes accruing in one strain, which is known to occur in disease outbreaks15. Additional support for Y. pestis microevolution is indicated by the presence of several variant positions for which sequence data from one individual shows two different nucleotides at comparable frequencies (Supplementary Table 5). Position 2896636, for example, is a known polymorphic position in extant Y. pestis populations14, and this position shows the fixed derived state in one individual (6330) and the polymorphic state in another (individual 8291) at minimum fivefold coverage (Supplementary Fig. 7). This provides a remarkable example of microevolution captured during an historical pandemic. The remaining variance positions are unchanged in the 18 extant Yersinia genomes, thus they may be unique to the ancient organism and are, therefore, of further interest. Additional sampling of ancient genomes will assist in determining the frequency of these mutations in co-circulating Y. pestis strains, and will clarify the emergence of branch 2 strains that are as yet unreported in ancient samples.
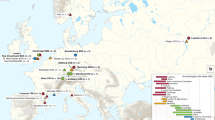
a, Median network of ancient and modern Y. pestis based on 1,694 variant positions in modern genomes14. Coloured circles represent different clades as defined in ref. 13. Gray circles represent hypothetical nodes. b, Phylogenetic tree using 1,694 variable positions. Divergence time intervals are shown in calendar years, with neighbour-joining bootstrap support (blue italic) and Bayesian posterior probability (blue). Grey box indicates known human pathogenic strains. A, NZ ACNQ01000; Nepal516, NC 008149; KIM10, NC 004088; B, NZ AAYT01000; C, NZ ABAT01000; D, NZ ACNS01000; E, NZ AAYS01000; F, NZ AAOS02000; CO92, NC 003143; G, NZ ABCD01000; H, NZ AAYV01000; I, NC 014029; J, NZ AAYR01000; Antiqua, NC 008150. c, Geographical origin of genome sequences used in a and b. d, Geographical spread of the Black Death from infection routes reported in ref. 4.
Consistent tree topologies were produced using several construction methods and all major nodes were supported by posterior probability (pp) values of >0.96 and bootstrap values >90 (Fig. 3b and Supplementary Figs 8 and 9). The trees place the East Smithfield sequence close to the ancestral node of all extant human pathogenic Y. pestis strains (only two differences in 1,694 positions) and at the base of branch 1 (Fig. 3b). A secure date for the East Smithfield site of 1348–1350 allowed us to assign a tip calibration to the ancient sequence and thus date the divergence time of the modern genomes and the East Smithfield genome using a Bayesian approach. Temporal estimates indicate that all Y. pestis commonly associated with human infection shared a common ancestor sometime between 668 and 729 years ago (ad 1282–1343, 95% highest probability density, HPD), encompassing a much smaller time interval than recently published estimates14 and further indicating that all currently circulating branch 1 and branch 2 isolates emerged during the thirteenth century at the earliest (Fig. 3b), potentially stemming from an Eastern Asian source as has been previously suggested14. This implies that the medieval plague was the main historical event that introduced human populations to the ancestor of all known pathogenic strains of Y. pestis. This further questions the aetiology of the sixth to eighth century Plague of Justinian, popularly assumed to have resulted from the same pathogen: our temporal estimates imply that the pandemic was either caused by a Y. pestis variant that is distinct from all currently circulating strains commonly associated with human infections, or it was another disease altogether.
Although our approach of using an extant Y. pestis reference template for bait design precluded our ability to identify genomic regions that may have been present in the ancient organism and were subsequently lost in CO92, genomic comparisons of our ancient sequence against its closest outgroups may yield valuable insights into Y. pestis evolution. The Microtus 91001 strain is the closest branch 1 and branch 2 relative confirmed to be non-pathogenic to humans16, hence genetic changes may represent contributions to the pathogen’s adaptation to a human host. Comparisons against this outgroup revealed 113 changes (Supplementary Table 6a, b), many of which are found in genes affecting virulence-associated functions like biofilm formation (hmsT), iron-acquisition (iucD) or adaptation to the intracellular environment (phoP). Similarly, although its virulence potential in humans has yet to be confirmed to our knowledge, Y. pestis B42003004 isolated from a Chinese marmot population17 has been identified as the strain closest to the ancestral node of all Y. pestis commonly associated with human plague, and thus may provide key information regarding the organism’s evolution. Full genome comparison against the East Smithfield sequence revealed only eight single-nucleotide differences (Supplementary Table 6c), six of which result in non-synonymous changes (Supplementary Table 6d). Although these differences probably do not affect virulence, the influence of gene loss, gene gain or genetic rearrangements, all of which are well documented in Y. pestis12,18, is as yet undetermined. In more recent evolutionary terms, single-nucleotide differences in several known pathogenicity-associated genes were found between our ancient genome and the CO92 reference sequence (Supplementary Table 3), which may represent further adaptations to human hosts.
Through enrichment by DNA capture coupled with targeted high throughput DNA sequencing, we have reconstructed a draft genome for what is arguably the most devastating human pathogen in history, and revealed that the medieval plague of the fourteenth century was probably responsible for its introduction and widespread distribution in human populations. This indicates that the pathogen implicated in the Black Death has close relatives in the twenty-first century that are both endemic and emerging19. Introductions of new pathogens to populations are often associated with increased incidence and severity of disease20 and although the mechanisms governing this phenomenon are complex21, genetic data from ancient infectious diseases will provide invaluable contributions towards our understanding of host–pathogen coevolution. The Black Death is a seminal example of an emerging infection, travelling across Europe and claiming the lives of an estimated 30 million people in only 5 years, which is much faster than contemporary rates of bubonic or pneumonic plague infection22 and dissemination7,8. Regardless, although no extant Y. pestis strain possesses the same genetic profile as our ancient organism, our data suggest that few changes in known virulence-associated genes have accrued in the organism’s 660 years of evolution as a human pathogen, further suggesting that its perceived increased virulence in history23 may not be due to novel fixed point mutations detectable via the analytical approach described here. At our current resolution, we posit that molecular changes in pathogens are but one component of a constellation of factors contributing to changing infectious disease prevalence and severity, where genetics of the host population24, climate25, vector dynamics26, social conditions27 and synergistic interactions with concurrent diseases28 should be foremost in discussions of population susceptibility to infectious disease and host–pathogen relationships with reference to Y. pestis infections.
Methods Summary
DNA from dental pulp was extracted and converted into sequencing libraries as previously described3. Potential sequencing artefacts resulting from deaminated nucleotides were eliminated by treatment of the DNA extracts with uracil-DNA-glycosylase and endonuclease VIII. DNA extracts were subsequently converted into sequencing libraries and amplified to incorporate unique sequence tags on both ends of the molecule. Two Agilent DNA capture arrays were designed for capture of the full Y. pestis chromosome (4.6 megabases), and the pCD1 (70 kb) and pMT1 (100 kb) plasmids using the modern Y. pestis strain CO92 (accession numbers NC_003143, NC_003131, NC_003134) for bait design with 3 bp tiling density. Serial array capture was performed over two copies of each array using the enriched fraction from the first round of capture as a template for a second round. The resulting products were amplified and pooled in equimolar amounts. All templates were sequenced for 76 cycles from both ends on the Illumina GAII platform, and reads merged into single fragments were included in subsequent analyses only if forward and reverse sequences overlapped by a minimum of 11 bp. Reads were mapped against the CO92 genome using the software BWA, and molecules with the same start and end coordinates were removed with the rmdup program in the samtools suite. Reference-guided sequence assembly was performed using Velvet version 1.1.03, with mapped and unmapped reads supplied in separate channels. Single-nucleotide differences were determined at a minimum of fivefold coverage and base frequency of at least 95% for both a pooled data set for all individuals and one in which all individuals were treated separately. A median network was constructed on these base calls using SplitsTree4. Phylogenetic trees were constructed using parsimony, neighbour-joining (MEGA 4.1) and Bayesian methods, and coalescence dates were determined in BEAST using both a strict and a relaxed molecular clock (Supplementary Fig. 9).
References
Stoneking, M. & Krause, J. Learning about human population history from ancient and modern genomes. Nature Rev. Genet. 12, 603–614 (2011)
Haensch, S. et al. Distinct clones of Yersinia pestis caused the Black Death. PLoS Pathog. 6, e1001134 (2010)
Schuenemann, V. J. et al. Targeted enrichment of ancient pathogens yielding the pPCP1 plasmid of Yersinia pestis from victims of the Black Death. Proc. Natl Acad. Sci. USA 10.1073/pnas.1105107108 (29 August, 2011).
Benedictow, O. J. The Black Death 1346–1353: The Complete History (Boydell Press, 2004)
Scott, S. & Duncan, C. J. Biology of Plagues (Cambridge Univ. Press, 2001)
Cohn, S. K. The Black Death Transformed: Disease and Culture in Early Renaissance Europe (Arnold Publication, 2003)
Achtman, M. et al. Yersinia pestis, the cause of the plague, is a recently emerged clone of Yersinia pseudotuberculosis. Proc. Natl Acad. Sci. USA 96, 14043–14048 (1999)
Cowal, L., Grainger, I., Hawkins, D. & Mikulski, R. 2008. The Black Death Cemetery, East Smithfield, London (Museum of London Archaeology Service, 2008)
Briggs, A. W. et al. Patterns of damage in genomic DNA sequences from a Neandertal. Proc. Natl Acad. Sci. USA 104, 14616–14621 (2007)
Hodges, E. et al. Hybrid selection of discrete genomic intervals on custom-designed microarrays for massively parallel sequencing. Nature Protocols 4, 960–974 (2009)
Green, R. E. et al. A complete Neandertal mitochondrial genome sequence determined by high-throughput sequencing. Cell 134, 416–426 (2008)
Chain, P. S. G. et al. Insights into the evolution of Yersinia pestis through whole-genome comparison with Yersinia pseudotuberculosis. Proc. Natl Acad. Sci. USA 101, 13826–13831 (2004)
Achtman, M. et al. Microevolution and history of the plague bacillus, Yersinia pestis. Proc. Natl Acad. Sci. USA 101, 17837–17842 (2004)
Morelli, G. et al. Yersinia pestis genome sequencing identifies patterns of global phylogenetic diversity. Nature Genet. 42, 1140–1143 (2010)
Harris, S. R. et al. Evolution of MRSA during hospital transmission and intercontinental spread. Science 327, 469–474 (2010)
Song, Y. et al. Complete genome sequence of Yersinia pestis strain 91001, an isolate avirulent to humans. DNA Res. 11, 179–197 (2004)
Eppinger, M. et al. Draft genome sequences of Yersinia pestis isolates from natural foci of endemic plague in China. J. Bacteriol. 191, 7628–7629 (2009)
Pouillot, F., Fayolle, C. & Carniel, E. Characterization of chromosomal regions conserved in Yersinia pseudotuberculosis and lost by Yersinia pestis. Infect. Immun. 76, 4592–4599 (2008)
Stenseth, N. C. et al. Plague: past, present, and future. PLoS Med. 5, e3 (2008)
Baum, J. & Bar-Gal, G. K. in Emerging Pathogens Archaeology, Ecology & Evolution of Infectious Diseases (eds Greenblat, C. L. & Spigleman, M. ) 67–78 (Oxford Univ. Press, 2003)
Brown, N. F. et al. Crossing the line: selection and evolution of virulence traits. PLoS Pathog. 2, e42 (2006)
WHO . Interregional meeting on prevention and control of plague. 〈http://www.who.int/csr/resources/publications/WHO_HSE_EPR_2008_3w.pdf〉 (2008)
Wood, J. W., Ferrell, R. J. & DeWitte-Aviña, S. N. The temporal dynamics of the fourteenth-century Black Death: new evidence from ecclesiastical records. Hum. Biol. 75, 427–448 (2003)
Joosten, M. H. A. J., Cosijnsen, T. J. & De Wit, P. J. G. Host resistance to a fungal tomato pathogen lost by a single base pair change in an avirulence gene. Nature 367, 384–386 (1994)
Xu, L. et al. Nonlinear effect of climate on plague during the third pandemic in China. Proc. Natl Acad. Sci. USA (2011)
Keeling, M. K. & Gilligan, C. A. Metapopulation dynamics of bubonic plague. Nature 407, 903–906 (2000)
Barrett, R., Kuzawa, C. W., McDade, T. & Armelagos, G. J. Emerging and re-emerging infectious diseases: the third epidemiologic transition. Annu. Rev. Anthropol. 27, 247–271 (1998)
Singer, M. & Clair, S. Syndemics and public health: reconceptualizing disease in bio-social context. Med. Anthropol. Q. 17, 423–441 (2003)
Acknowledgements
We thank W. White (deceased), J. Bekvalac and R. Redfern from the Museum of London Centre for Human Bioarchaeology for access to samples, M. Kircher and S. Forrest for assistance with computational analysis, G. Wright for support throughout the project, past and present members of the McMaster Ancient DNA Centre for support throughout the project, and D. Poinar for constructive comments on earlier versions of the manuscript. We also thank S. Pääbo and the Max Planck Institute of Evolutionary Anthropology for use of their clean room facilities and molecular biology lab. Funding was provided by the Carl Zeiss Foundation (J.K.), the Human Genetics department of the Medical faculty in Tübingen (J.K.), the Canada Research Chairs program (H.N.P., G.B.G.), the Canadian Institute for Health Research (H.N.P.), the Social Science and Humanities Research Council of Canada (H.N.P.), the Michael G. DeGroote Institute for Infectious Disease Research (H.N.P., B.K.C., D.J.D.E.), an Early Research award from the Ontario Ministry of Research and Education (H.N.P.), the Natural Sciences and Engineering Research Council of Canada (D.J.D.E.), the James S. McDonnell Foundation (D.J.D.E.), and the University at Albany Research Foundation and Center for Social and Demographic Analysis and the Wenner-Gren Foundation (S.N.D.).
Author information
Authors and Affiliations
Contributions
K.I.B., S.N.D., D.J.D.E., J.K. and H.N.P. conceived the project. K.I.B., S.N.D., S.S. and J.W. performed skeletal sampling. K.I.B., J.K. and V.J.S. carried out laboratory work. H.A.B., K.I.B., J.K., M.M. and H.N.P. designed experiments. K.I.B., G.B.G., J.K., H.N.P., V.J.S. and N.W. analysed the data. B.K.C., D.J.D.E., D.A.H. and J.B.M. provided valuable interpretations. P.B. provided technical support. K.I.B., J.K. and H.N.P. wrote the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
This file contains Supplementary Material and Methods, Supplementary Figures 1-9 with legends, Supplementary Tables 1a, 1c, 2a, 2b (see separate files for Supplementary Tables 1b, 3a, 3b, 4, 5, 6a, 6b, 6c and 6d) and Supplementary References. (PDF 3537 kb)
Supplementary Table 1b
This file contains the description of all mapped reads. (XLS 15 kb)
Supplementary Table 3a
This file shows the nucleotide difference between the ancient genome and CO92 (chromosome). The original version of this table contained some errors and it was replaced on 30 November 2011. (XLS 30 kb)
Supplementary Table 3b
This file shows the nucleotide differences between the ancient genome and CO92 (plamids). (XLS 10 kb)
Supplementary Table 4
This file contains the known polymorphic positions used for phylogenetic reconstruction. The original version of this table contained some errors and it was replaced on 30 November 2011. (XLS 720 kb)
Supplementary Table 5
This file shows the variant positions in the ancient genome. (XLS 21 kb)
Supplementary Table 6a
This file shows the differences between B42003004 and the 18 modern genomes. (XLS 61 kb)
Supplementary Table 6b
This file shows the differences between B42003004 and the ancient genome. (XLS 24 kb)
Supplementary Table 6c
This file shows the differences between Mictrotus 91001 and the 18 modern genomes. (XLS 12 kb)
Supplementary Table 6d
This file shows the differences between Microtus 91001 and the ancient genome. (XLS 10 kb)
Rights and permissions
This article is distributed under the terms of the Creative Commons Attribution-Non-Commercial-Share Alike licence (http://creativecommons.org/licenses/by-nc-sa/3.0/), which permits distribution, and reproduction in any medium, provided the original author and source are credited. This licence does not permit commercial exploitation, and derivative works must be licensed under the same or similar licence.
About this article
Cite this article
Bos, K., Schuenemann, V., Golding, G. et al. A draft genome of Yersinia pestis from victims of the Black Death. Nature 478, 506–510 (2011). https://doi.org/10.1038/nature10549
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature10549
This article is cited by
-
Yersinia entomophaga Tc toxin is released by T10SS-dependent lysis of specialized cell subpopulations
Nature Microbiology (2024)
-
Identification of microbial pathogens in Neolithic Scandinavian humans
Scientific Reports (2024)
-
Effect of temperature on fast transmission of COVID-19 in low per capita GDP Asian countries
Scientific Reports (2023)
-
Ancient Clostridium DNA and variants of tetanus neurotoxins associated with human archaeological remains
Nature Communications (2023)
-
Yersinia pestis genomes reveal plague in Britain 4000 years ago
Nature Communications (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.