Abstract
Uveal melanoma is the most common malignant tumor of the adult eye. Fifty percent of tumors will eventually metastasize, and there are no effective treatments for them. Recent studies of uveal melanoma have identified activating mutations in GNAQ and GNA11, loss-of-function mutations in the tumor suppressor gene BAP1, and recurrent mutations in codon 625 of SF3B1. Previous studies have reported the existence of a higher frequency of GNA11 than GNAQ mutations, frequent BAP1 loss, and rare SF3B1 mutations in metastatic uveal melanoma. We analyzed a cohort of 30 uveal melanoma metastases for the occurrence of GNAQ, GNA11, and SF3B1 mutations, as well as BAP1 loss, and correlated these parameters with clinical and histopathologic features. Most (92%) tumors were composed of cells with an epithelioid or mixed (<100% spindle cells) morphology. Tumor samples composed exclusively of spindle cells were rare (n=2, 8%). Most tumors showed a moderate to marked degree of nuclear pleomorphism (n=24, 96%), and contained hyperchromatic, vesicular nuclei with variably conspicuous nucleoli. GNA11 mutations were considerably more frequent than GNAQ mutations (GNA11, GNAQ, and wild-type in 18 (60%), 6 (20%), and 6 (20%) cases, respectively). SF3B1 mutation was found in 1 of 26 tumors (4%), whereas loss of BAP1 expression was present in 13 of 16 tumors (81%). Patients with GNA11-mutant tumors had poorer disease-specific survival (60.0 vs 121.4 months, P=0.03) and overall survival (50.6 vs 121.4 months, P=0.03) than those with tumors lacking GNA11 mutations. The survival data, combined with the predominance of GNA11 mutations in metastases, raises the possibility that GNA11-mutant tumors may be associated with a higher risk of metastasis and poorer prognosis than GNAQ-mutant tumors. Further studies of uveal melanoma are required to investigate the functional and prognostic relevance of oncogenic mutations in GNA11 and GNAQ.
Similar content being viewed by others
Main
Approximately 5% of all melanomas occur in the eye, of which most arise in the uveal tract (iris, ciliary body, or choroid).1 Uveal melanoma is the most common malignant tumor of the eye, with an incidence of about four cases per million per annum.2 Diagnosis often occurs late in the course of disease, and prognosis is generally poor with a 10-year mortality rate of around 40%.3 The most common site of metastasis is the liver.4, 5 Clinical parameters associated with prognosis include the site of tumor, tumor size, and extraocular invasion.6 Histopathologic prognostic factors include cell type (epithelioid or spindle), mitotic/Ki-67 index, extravascular matrix patterns, and vascular invasion.6, 7, 8
Advances in genetics have led to a better understanding of the events involved in the development of these tumors. An important finding was the identification of chromosome 3 loss correlating with metastasis and poor prognosis.9, 10 Recently, BAP1, a gene located in the area frequently lost on chromosome 3, was found to carry inactivating mutations in a large proportion (84%) of metastasizing uveal melanomas.11 BAP1 is an ubiquitin hydroxyl carboxy-terminal hydrolase. Mutations either affect the active domain of the enzyme or lead to a frameshift or nonsense mutation, resulting in translation of a non-functional protein. Although the functional role of BAP1 is still not fully understood, studies have implicated a role in chromatin remodeling12 as well as cell cycle control.13
Uveal melanomas lack activating mutations in BRAF or NRAS,14 which commonly occur in cutaneous melanoma (approximately 50% and 20%, respectively).15 Eighty to 90% of uveal melanomas harbor activating mutations in GNAQ or GNA119, 16 in a mutually exclusive pattern. The original report of GNA11 mutations showed a substantial difference in the distribution of mutations in primary and metastatic tumors. The ratio of GNA11 to GNAQ mutations was 0.7 (52:72) in primary tumors, and 2.6 (13:5) in metastases.16
Recently, mutations in codon 625 of SF3B1 on chromosome 2 were found in uveal melanomas. SF3B1 mutations were almost always mutually exclusive of BAP1 loss and were associated with a good prognosis. They were not detected in five metastases analyzed.17 Described as a splice factor, SF3B1 was assumed to have a role in excising introns from premessenger RNA. Harbour et al17 also proposed that the effect of SF3B1 mutation on tumorigenesis might be linked to an effect on chromatin remodeling. Future studies will be required to delineate the functional role of SF3B1 mutations.
In this study, we evaluated clinical, pathologic, and genetic features of a cohort of uveal melanoma metastases and analyzed their associations with survival.
Materials and methods
Sample Selection
The databases of Melanoma Institute Australia and the Department of Dermatology, Essen were searched for metastatic ocular melanoma of posterior uveal origin (arising either from the choroid or ciliary body). The study was carried out in accordance with Ethics Committee guidelines at each of the participating institutions.
Histopathology and Immunohistochemistry
Tissue blocks containing tumor were sectioned at thicknesses of 5 μm (for hematoxylin–eosin staining and immunohistochemistry, IHC) and 10 μm (for microdissection and DNA isolation). BAP1 IHC was performed according to methods that have been described previously.18 All histologic and IHC sections were reviewed by at least two histopathologists (KGG, JvdN and RM).
DNA Isolation
Ten 10-μm-thick sections of paraffin-embedded tissue were deparaffinized according to the following protocol: two steps of 10 min xylene, 5 min 100% ethanol, 5 min 95% ethanol, and 5 min 70% ethanol, and then rinsing in water. After drying, tumor tissue was manually macrodissected from the sections. Genomic DNA was isolated using the QIAamp DNA Mini Kit (Qiagen, Hilden, Germany) according to the manufacturer’s instructions. DNA from frozen tissue was directly applied to the Qiagen kit for purification.
Sanger Sequencing
The following primers were used: for GNAQ, exon 5—5′-TGATCATCGTCATTCAAGAGAA-3′ (F), 5′-AGAAACATGATAGAGGTGACATTTT-3′ (R), exon 4—5′-TGGTGTGATGGTGTCACTGACATTCTCAT-3′ (F), 5′-AGCTGGGAAATAGGTTTCATGGACTCAGT-3′ (R); and for GNA11, exon 5—5′-CGCTGTGTCCTTTCAGGATG-3′ (F), 5′-CCACCTCGTTGTCCGACT-3′ (R), exon 4—5′-GTGCTGTGTCCCTGTCCTG-3′ (F), 5′-GGCAAATGAGCCTCTCAGTG-3′ (R). Primers for the first 120 bp of exon 14 of SF3B1 were as follows: 5′-TGTTTACATTTTAGGCTGCTGGT-5′ (F), 5′-GCCAGGACTTCTTGCTTTTG-3′ (R). The PCR reaction conditions were 10 mM dNTPmix, 1 U Thermoprime Plus DNA Polymerase (Abgene), 1 x Ready Mix PCR Buffer IV with 15 mM MgCl2 (Thermo Scientific), and 10 pM of each primer. PCR consisted of 35 cycles of 95 °C (45 s), 57 °C (45 s), and 72 °C (45 s) after initial denaturation at 95 °C for 5 min. PCR reaction products were purified with the QIAquick PCR Purification kit (Qiagen) and then used as templates for sequencing in both directions.
Associations of GNAQ, GNA11, and BAP1 Status with Clinical and Pathologic Parameters
We investigated associations of mutation status with available clinical and pathologic parameters using χ2 tests and Fisher’s exact tests as appropriate. Univariate Cox regression models were used to analyze associations of mutation status with overall survival from (a) the time of diagnosis of metastasis of uveal melanoma, and (b) the time of diagnosis of primary uveal melanoma to the time of death or last follow-up. Cases in which the specified end points were missing were censored. All statistical analyses were performed using IBM SPSS Statistics software (version 20.0; International Business Machines, Armonk, NY, USA). A P-value of ⩽0.05 was considered statistically significant.
Results
Thirty-seven metastases from primary uveal melanoma were identified from the institutional databases. Tumor DNA of sufficient quality for genetic analysis was isolated from 30 tumors, which were included in the study (Table 1). The metastases were located in the skin (10, 33%), liver (9, 30%), lung (1, 3%), brain (1, 3%), spleen (1, 3%), lymph node (1, 3%), mediastinum (1, 3%), and unknown sites (6, 20%). Most tumor samples (n=21) were paraffin-embedded, and a few (n=9) were frozen tissues. In cases with minimal biopsy material (n=4), no sectioning for H&E slides or IHC was possible, and only DNA analysis was performed.
Histologic Analysis
Histologic evaluation was performed in 25 tumors for which slides were available or could be prepared. Twenty-three (92%) tumors were composed of cells with an epithelioid or mixed (<100% spindle cells) morphology. Tumor samples composed exclusively of spindle cells were rare (n=2, 8%). Most tumors showed a moderate to marked degree of nuclear pleomorphism (n=24, 96%), and contained hyperchromatic, vesicular nuclei with variably conspicuous nucleoli. The cytoplasm in most tumors was moderate in amount and amphophilic. The following parameters varied widely: degree of pigmentation, number of admixed macrophages, extent of tumor necrosis (Table 1), and tumor mitotic rate (from 0 to 20 per mm2). Lymphovascular and perineural invasion (LVI and PNI, respectively) were rare (Table 2).
GNAQ/GNA11 Mutation Status
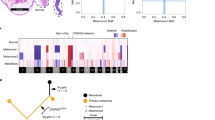
Sequencing of GNAQ and GNA11 was successful in all 30 tumors (Tables 1 and 2). Exon 5 of both genes, which contains the hotspot mutation leading to the most frequent alteration of the Q209 amino acid, was amplified and sequenced in all cases (Figure 1). In tumors lacking mutations in exon 5 of both GNAQ and GNA11, exon 4 of both genes, which harbors the less frequent R183 mutation, was sequenced (Figure 1).
Mutations in GNA11 were identified in 18 (60%) tumors (1 brain, 7 skin, 1 lung, 5 liver, 1 mediastinum, 2 unknown site, and 1 sequenced from the primary). There were mutations in exon 5 in 15 tumors, all of which were c.626A>T leading to a Q209L amino-acid change. In addition, three c.547C>T mutations in exon 4, resulting in R183C substitutions, were detected. Mutations in GNAQ were identified in six (20%) tumors (3 liver, 1 lymph node, 1 skin metastasis, and 1 sequenced from the primary). All mutations, four c.626A>C and two c.626A>T resulting in Q209P and Q209L amino-acid substitutions, respectively, were found in exon 5.
SF3B1 Mutation Status
In 29 tumors, we sequenced the first 120 base pairs of exon 14 of SF3B1, which contains the published hotspot in codon 625. High-quality results allowing confident mutation detection were available in 26 cases. Only one tumor (1/26, 4%) harbored a c.1873C>T mutation leading to an R625C amino-acid change.
BAP1 Expression
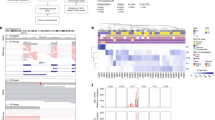
BAP1 status was analyzed by immunohistochemistry. This allows the detection of loss of protein expression due to either inactivating mutations or epigenetic silencing mechanisms. BAP1 loss was found in 13/16 (81%) tumors analyzed (Figure 2).
Morphology and BAP1 immunohistochemistry in metastatic uveal melanoma. (a and b) Mitotically active tumor composed of epithelioid cells, showing loss of expression of BAP1 ((a) hematoxylin–eosin, × 400; (b) BAP1 IHC, × 400). (c and d) Tumor composed of nests of epithelioid cells, with expression of BAP1 ((c) hematoxylin–eosin, × 200; (d) BAP1 IHC, × 200). (e and f) Tumor composed of a predominance of spindle-shaped cells, showing loss of expression of BAP1 ((e) hematoxylin–eosin, × 200; (f) BAP1 IHC, × 200). Note: the presence of BAP1 expression is indicated by nuclear expression of red chromogen.
Associations of Mutation Status with Clinical and Pathologic Parameters
There were no statistically significant associations of GNAQ or GNA11 mutation status with: age at either primary diagnosis or metastasis, sex, status (alive or deceased), site of metastasis, cell pleomorphism, pigmentation, melanophages, LVI, PNI, or loss of BAP1 by IHC (Table 3). Cell type showed a significant association (P=0.01), with the only two tumors composed exclusively of spindle cells having neither a GNAQ nor GNA11 mutation. Tumor-infiltrating lymphocytes (TILs) were present in 7/16 (44%) GNA11-mutant tumors and 3/4 (75%) GNAQ-mutant tumors, but were absent in all 5 tumors wild type for GNAQ and GNA11 (P=0.08).
Median follow-up durations were 68.2 (range 9.8–346.6) months from the diagnosis of primary tumor, and 8.6 (range 0.0–42.3) months from the diagnosis of first distant metastasis. The interval between primary tumor and first distant metastasis ranged between 5.1 and 307.2 months, with a median of 50.8 months.
No statistically significant associations were identified between various clinical, pathologic, or genetic factors and BAP1 expression status (Table 4) or survival following the diagnosis of metastasis (Table 5). However, disease-specific survival and overall survival (measured from the date of diagnosis of the primary tumor) were significantly poorer in (a) patients with GNA11-mutant tumors than in those with tumors lacking GNA11 mutations; and (b) in patients with metastases exhibiting evidence of LVI (Table 5 and Figure 3). Multivariable survival analyses were not performed because of the small number of tumors in subgroups within the study cohort.
Discussion
Traditionally, prediction of prognosis in uveal melanoma patients has relied on evaluation of clinical and pathologic criteria such as tumor size and cell type. More recently, several genetic tests have been shown to aid prognostic prediction. Chromosome 3 loss,19 BAP1 loss,11 and RNA expression profiling20, 21 have high predictive values in terms of forecasting the course of disease and the risk of metastasis. SF3B1 codon 625 mutations also appear to be a genetic marker of good prognosis.17
Our finding that the majority of uveal melanoma metastases showed loss of BAP1 expression (13/16, 81%) is consistent with prior findings,11 and confirming that BAP1 loss is a marker of bad prognosis. Only one SF3B1 codon 625 mutation was identified in 26 tumors tested (3.8%), which is much lower than the percentage detected in primary tumors (18.6%) and consistent with the report that found SF3B1 mutations to be a good prognostic marker.17 The SF3B1-mutant tumor was one of the three tumors that retained expression of BAP1 protein. This suggests that metastasis may still occur in patients whose tumors harbor SF3B1 codon 625 mutations and retain BAP1 expression. Identifying additional markers that detect those SF3B1-mutant tumors that are prone to metastasize would be of great clinical relevance. The presence of SF3B1 mutation in only one tumor precluded meaningful analyses of clinicopathologic features and survival associated with SF3B1 mutation.
The majority of metastases were composed of an epithelioid or mixed-cell population, which is in keeping with the known adverse prognostic profile associated with these cell types relative to pure spindle cell tumors.22, 23 Of potential interest is the finding that metastases composed predominantly of spindle cells and those lacking TILs were wild type for both GNAQ and GNA11. However, the number of tumors in these groups is very small, and analysis of larger numbers of tumors will be required to validate these findings.
The presence of LVI was associated with significantly poorer survival (as measured from the date of diagnosis of the primary tumor). This finding may indicate a greater propensity for tumor dissemination, and is consistent with the findings of a recent study in which the presence of intravascular tumor was associated with poorer prognosis.24
Although currently not assumed to be of prognostic relevance, the type of driving oncogene involved may have a significant supplemental effect on long-term survival and outcome. Mutations in GNAQ and GNA11 have been shown to be mutually exclusive,16 with tumors harboring only a single mutation in either gene. We found a strong bias toward GNA11 mutations in our cohort. Eighteen tumors harbored GNA11 mutations compared to six with GNAQ mutations and six tumors lacking either mutation. The ratio of GNA11 to GNAQ mutations was 3.0. Our finding of the predominance of GNA11 mutations in uveal melanoma metastases fits well with the original report of GNA11 mutations in uveal melanoma.16 Of the 163 primary tumors analyzed, 52 (32%) had activating mutations in GNA11 and 72 (45%) had the corresponding mutation in GNAQ. On the other hand, in 23 metastases, 13 (57%) had GNA11 and only 5 (22%) harbored GNAQ mutations. Our data confirm the over-representation of GNA11 mutations in metastases, and raises the possibility of prognostic significance associated with the type of oncogenic mutation. Validation and further exploration of these findings in additional, larger cohorts is warranted.
The oncogenic mutation data suggest that GNA11-mutant tumors may have a higher tendency to metastasize than GNAQ-mutant tumors. If this is the case, one might expect that the survival of patients with GNA11-mutant tumors should be significantly lower than that for patients with GNAQ-mutant tumors. Indeed, we found that survival from the diagnosis of the primary tumor was significantly poorer in patients with GNA11-mutant tumors compared with patients lacking GNA11 mutations in their tumors (Table 5 and Figure 3). Since GNA11 and GNAQ mutations are early events in the evolution of uveal melanoma,16 it is reasonable to presume that there is fidelity between the presence of these mutations in metastases and the primary tumors from which they arose. However, the survival data in a previous study16 showed that GNA11-mutant tumors had a better prognosis than GNAQ-mutant tumors. One of the major limitations of the survival analysis in that study is the duration of available follow-up. Initially, the GNA11- and GNAQ-mutant tumor groups contained 26 and 43 patients, respectively, but the majority of patients were censored early in the follow-up period; there were only 5 and 3 tumors, respectively, at 5 years after diagnosis, and only 2 patients per group remained at 10 years. Given that well-documented long-term follow-up studies have shown that uveal melanoma patients can die of disease 10 or more years after diagnosis,3, 4, 25, 26 it is clear that more complete, longer follow-up will be required to establish whether GNA11-mutant tumors do indeed have a significantly worse overall survival than GNAQ-mutant tumors.
Our results should be interpreted with some caution. The study cohort is small, and is highly selective, namely uveal melanoma metastases, and therefore reflects patients with high-risk disease. There is also some sample bias; the percentage of cutaneous metastases in the cohort is higher than that expected in the general uveal melanoma population, in which hepatic metastases would predominate. This probable selection bias likely reflects the ease of resection and low morbidity associated with biopsying metastases involving superficial cutaneous sites compared with hepatic or other visceral lesions. Nevertheless, the results generate an interesting hypothesis that is worthy of further study: that GNA11 mutations are associated with aggressive disease and poor prognosis in uveal melanoma. It is conceivable that the poor prognostic significance of GNA11 mutations applies particularly to patients with high-risk disease.
In summary, we found that in metastatic uveal melanoma, GNA11 mutations were more frequent, and were associated with poorer survival. In addition, loss of BAP1 expression was frequent, and mutation of SF3B1 at codon 625 was rare. Larger cohorts, optimally with comparable sets of primary tumors and long-term follow-up, will be needed to validate our findings, which imply more aggressive behavior of tumors harboring GNA11 mutations. If a prognostic significance of GNA11 is validated, it may influence management of affected patients. For example, the intensity of follow-up could be stratified according to the oncogene mutation harbored, and potentially treatment strategies should also be adjusted based on the oncogene mutation present.
References
Shildkrot Y, Wilson MW . Update on posterior uveal melanoma: treatment of the eye and emerging strategies in the prognosis and treatment of metastatic disease. Curr Opin Ophthalmol 2009;20:504–510.
Chang AE, Karnell LH, Menck HR . The National Cancer Data Base report on cutaneous and noncutaneous melanoma: a summary of 84,836 cases from the past decade. The American College of Surgeons Commission on Cancer and the American Cancer Society. Cancer 1998;83:1664–1678.
Kujala E, Tuomaala S, Eskelin S et al. Mortality after uveal and conjunctival melanoma: which tumour is more deadly? Acta Ophthalmol 2009;87:149–153.
Kujala E, Makitie T, Kivela T . Very long-term prognosis of patients with malignant uveal melanoma. Invest Ophthalmol Vis Sci 2003;44:4651–4659.
Singh AD, Borden EC . Metastatic uveal melanoma. Ophthalmol Clin N Am 2005;18:143–150 ix.
Griewank KG, Murali R . Pathology and genetics of uveal melanoma. Pathology 2013;45:18–27.
McLean IW, Foster WD, Zimmerman LE . Uveal melanoma: location, size, cell type, and enucleation as risk factors in metastasis. Hum Pathol 1982;13:123–132.
Mudhar HS, Parsons MA, Sisley K et al. A critical appraisal of the prognostic and predictive factors for uveal malignant melanoma. Histopathology 2004;45:1–12.
Van Raamsdonk CD, Bezrookove V, Green G et al. Frequent somatic mutations of GNAQ in uveal melanoma and blue naevi. Nature 2009;457:599–602.
Abdel-Rahman MH, Cebulla CM, Verma V et al. Monosomy 3 status of uveal melanoma metastases is associated with rapidly progressive tumors and short survival. Exp Eye Res 2012;100:26–31.
Harbour JW, Onken MD, Roberson ED et al. Frequent mutation of BAP1 in metastasizing uveal melanomas. Science 2010;330:1410–1413.
Scheuermann JC, de Ayala Alonso AG, Oktaba K et al. Histone H2A deubiquitinase activity of the Polycomb repressive complex PR-DUB. Nature 2010;465:243–247.
Eletr ZM, Wilkinson KD . An emerging model for BAP1’s role in regulating cell cycle progression. Cell Biochem Biophys 2011;60:3–11.
Spendlove HE, Damato BE, Humphreys J et al. BRAF mutations are detectable in conjunctival but not uveal melanomas. Melanoma Res 2004;14:449–452.
Sekulic A, Haluska P Jr., Miller AJ et al. Malignant melanoma in the 21st century: the emerging molecular landscape. Mayo Clin Proc 2008;83:825–846.
Van Raamsdonk CD, Griewank KG, Crosby MB et al. Mutations in GNA11 in uveal melanoma. N Engl J Med 2010;363:2191–2199.
Harbour JW, Roberson ED, Anbunathan H et al. Recurrent mutations at codon 625 of the splicing factor SF3B1 in uveal melanoma. Nat Genet 2013;45:133–135.
Wiesner T, Murali R, Fried I et al. A distinct subset of atypical Spitz tumors is characterized by BRAF mutation and loss of BAP1 expression. Am J Surg Pathol 2012;36:818–830.
Prescher G, Bornfeld N, Hirche H et al. Prognostic implications of monosomy 3 in uveal melanoma. Lancet 1996;347:1222–1225.
Tschentscher F, Husing J, Holter T et al. Tumor classification based on gene expression profiling shows that uveal melanomas with and without monosomy 3 represent two distinct entities. Cancer Res 2003;63:2578–2584.
Onken MD, Worley LA, Ehlers JP et al. Gene expression profiling in uveal melanoma reveals two molecular classes and predicts metastatic death. Cancer Res 2004;64:7205–7209.
McLean IW, Foster WD, Zimmerman LE et al. Modifications of Callender’s classification of uveal melanoma at the Armed Forces Institute of Pathology. Am J Ophthalmol 1983;96:502–509.
Gamel JW, McLean IW, Foster WD et al. Uveal melanomas: correlation of cytologic features with prognosis. Cancer 1978;41:1897–1901.
Ly LV, Odish OF, Wolff-Rouendaal D et al. Intravascular presence of tumor cells as prognostic parameter in uveal melanoma: a 35-year survey. Invest Ophthalmol Vis Sci 2010;51:658–665.
Singh AD, Topham A . Survival rates with uveal melanoma in the United States: 1973–1997. Ophthalmology 2003;110:962–965.
Kivela T, Eskelin S, Kujala E . Metastatic uveal melanoma. Int Ophthalmol Clin 2006;46:133–149.
Acknowledgements
We thank Sabine Prass, Marion Schwamborn, and Nicola Bielefeld for their excellent technical support. Assistance from staff of Melanoma Institute Australia and Royal Prince Alfred Hospital and funding support from the National Health and Medical Research Council (of the Commonwealth Government of Australia) and the Cancer Institute New South Wales is also gratefully acknowledged.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
Dirk Schadendorf is on the advisory board or has received honararia from Roche, Genetech, Novartis, Amgen, GSK, BMS, Boehringer Ingelheim, and Merck. Lisa Zimmer has honoraria from Roche, Bristol-Meyers Squibb, and Amgen, and travel support from Merck Sharp & Dohme and Bristol-Meyers Squibb. All the other authors declare no conlictof interest.
Additional information
This study was presented in part at the 102nd Annual Meeting of the United States and Canadian Academy of Pathology, held between 2 and 8 March in Baltimore, MD, USA.
Rights and permissions
About this article
Cite this article
Griewank, K., van de Nes, J., Schilling, B. et al. Genetic and clinico-pathologic analysis of metastatic uveal melanoma. Mod Pathol 27, 175–183 (2014). https://doi.org/10.1038/modpathol.2013.138
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/modpathol.2013.138
Keywords
This article is cited by
-
Deep learning classification of uveal melanoma based on histopathological images and identification of a novel indicator for prognosis of patients
Biological Procedures Online (2023)
-
The immune cell landscape of metastatic uveal melanoma correlates with overall survival
Journal of Experimental & Clinical Cancer Research (2021)
-
Frequent and Yet Unreported GNAQ and GNA11 Mutations are Found in Uveal Melanomas
Pathology & Oncology Research (2019)
-
Okuläre Melanome
Der Pathologe (2017)
-
The biology of uveal melanoma
Cancer and Metastasis Reviews (2017)