Abstract
Analysis of the intestinal B-cell system and properties of immunoglobulin A, the main antibody isotype produced in the gut, has dominated the rise of mucosal immunology as a discipline. Seminal work established concepts describing the induction, transport, and function of mucosal antibodies. Still, open questions remain and we lack a comprehensive view of how the various sites and pathways of immunoglobulin A induction are integrated to respond to gut antigens. Next-generation sequencing (NGS) offers a novel approach to study B-cell responses, which might substantially enhance our tool box to answer key questions in the field and to take the next steps toward therapeutic exploitation of the mucosal B-cell system. In this review we discuss the potential, challenges, and emerging solutions for gut B-cell repertoire analysis by NGS.
Similar content being viewed by others
Setting the Stage: The Primary and Secondary B Cell Pools in the Gut
A large part of the intestinal immune system is devoted to the induction and maturation of B-cell responses and antibody production. B cells are the most abundant cell population in gut-associated lymphoid tissue such as Peyer’s patches, small intestinal isolated lymphoid follicles, the appendix, and colonic follicles.1 Characteristics of gut-associated lymphoid tissue are large germinal centers, indicating that within these sites B-cell responses are constitutively induced and refined. B cells activated at these sites express gut tropic homing cues, enter the intestinal lamina propria, and differentiate into antibody-secreting plasma cells.1 These processes establish the largest plasma cell population in the body, which in human gut has been estimated to comprise ∼7 × 1010 plasma cells positioned in the gut lamina propria.2 This plasma cell population locally produces antibodies, which by the action of the polymeric immunoglobulin (Ig) receptor are shuttled across the gut epithelium and secreted into the gut lumen. Secretory Igs neutralize toxins, provide protection against enteropathogens, and regulate the intestinal microbiota. The functions of secretory antibodies have been reviewed in ref. 3.
The configuration of the B-cell receptor (BCR), which is a key determinant of antigen specificity and unique for each B-cell clone, is assessable via sequencing-based approaches. Naive B cells express a surface BCR, which allows B cells to recognize antigens and to initiate B-cell activation and expansion, which eventually results in the generation of memory B cells and antibody-secreting plasma cells. The BCRs (such as the secreted BCRs, which are the antibodies/Igs) in mice and humans are all composed of heavy and light chains. Heavy and light chains combine constant and variable Ig domains. The amino-terminal variable Ig domains, referred to as VH and VL for the heavy and light chain, respectively, confer antigen specificity. In contrast, the constant domain of the heavy chain (CH) couples the BCR/antibody to cellular receptors and other components of the immune system and thereby confers effector functions. Some isotypes including secretory IgA assemble to higher molecular weight complexes. In the gut, IgA is produced mostly as a dimer comprising two identical sets of paired heavy and light chains. The structure of IgA has been reviewed in ref. 4.
Variable heavy and light chain domains are generated during B-cell development through somatic recombination. Gene segments selected from an array of alternative V (variable), D (diversity), and J (joining) gene segments are recombined to form the variable antigen binding VH domain of the heavy chain, whereas the VL domain is assembled through recombination of V and J segments without a D gene segment.5 Beyond the combination of gene segments, additional diversity of the BCRs is generated through imprecise joining of the gene segments, including nucleotide P and N addition and deletion, which further increases the number of unique variations of the BCRs by several orders of magnitude.6 Combinatorial and junctional diversity along with the pairing of alternative heavy and light chains can generate >1013 alternative BCRs in humans. This theoretical diversity by far exceeds the number of B cells present at a given time in an organism (typically 1011 in a human and 108 in a mouse). Consequently, the VH- and VL-encoding sequences are characteristic for a given B-cell clone and it is highly unlikely for the same sequence to appear twice in the same individual.7, 8 The highest variability is encoded in the V(D)J recombination site generally referred to as complementarity determining region 3 (CDR3). The CDR3 has a key role in determining antigen specificity, and sequence analysis of CDR3 regions can be used to identify clonally related B cells (see below).
B-cell activation results in clonal expansion. On activation, B cells can modify their BCR through class switch recombination (CSR) and somatic hypermutation (SHM). SHM mutates the variable domains and allows B cells to increase the affinity of their BCR to a given antigen. In contrast, CSR does not alter VH and VL domains but results in changes in the constant region, thereby altering effector functions but not antigen specificity.
The antigenic load inducing Ig responses in the intestine seems vast. The intestine, in particular its distal parts, is colonized by a dense population of microbiota, including bacteria, viruses, fungi, and helminths. In addition, pathogens ingested with the diet confront the intestinal immune system and food in itself provides a rich source of foreign antigens. Considering the complexity of the intestinal antigen load, it does not seem surprising that several alternative mechanisms of Ig induction and maturation have been identified. However, the contribution and relevance of some proposed mechanisms are heatedly discussed and some findings reported in mice contrast with observations made in humans.9, 10, 11 This uncertainty impedes the development of mucosal vaccines inducing effective secretory Ig responses.9, 12 Moreover, therapeutic manipulation of the microbiota might require a better understanding of how microbiota induce Ig responses.13 Establishing a comprehensive concept of intestinal Ig responses will require the combination of a broad range of technical approaches. We anticipate that next-generation sequencing (NGS) will be a particularly promising approach to supplement other advancements in the field, such as a more complete understanding of gut plasma cell phenotypes and composition. This notwithstanding, NGS is no magic bullet. At present, it is impossible to reliably predict antibody specificity/antigens recognized based on sequence information alone. This gap between Ig sequence information and IgA reactivity in the gut eventually requires the production of functional antibodies and characterization of their binding profiles (see “The gap between sequence and antigen specificity”). Moreover, the pathways of intestinal IgA induction are complex. For example, in the gut germinal center formation and SHM can occur in the absence of cognate BCR engagement;14 however, there are controversial findings on the ability of B cells to undergo local switch recombination in the gut lamina propria15, 16 and lymphotoxin produced by innate lymphoid cells contributes to intestinal IgA production.17 Therefore, the generation of NGS-based data sets and their interpretation require exact sophisticated phenotypic description and isolation of the starting material/cells, and need to go hand in hand with other non-NGS-based approaches. In this review we will focus on the potential and the pitfalls of NGS in the context of the efforts to better understand intestinal B-cell responses. We will not discuss in detail other exciting new developments in the field of mucosal B-cell responses.
What to Expect from NGS Sequencing
The application of Ig repertoire analysis is multi-fold, ranging from novel approaches of antibody discovery and potential use of Ig genes as biomarkers to fundamentally new approaches, to studying the formation of B-cell repertoires in health, aging, and disease,18, 19 (reviewed in ref. 20). NGS enables us to compare snapshots of the B-cell repertoire from different tissues, ages, disease stages, and points in time, and to observe how repertoire diversity, clonotypes, and characteristics of affinity maturation vary with these parameters. Seminal work established how B cells generate diverse BCRs during their differentiation.5 Yet, as opposed to T cells, the B-cell repertoire is further modified on B-cell activation by SHM and CSR. Lineage tree analysis allows describing B-cell maturation that occurs after cell activation. Such approaches have long been used to observe the relationships between B cells sampled under different conditions. Yet, the predictive power of B-cell repertoire analysis has taken a great leap forward with the increase in numbers of sequences obtained by NGS compared with that obtained by Sanger sequencing. Along with this quantitative difference there is a qualitative difference when considering the biological questions that can be addressed. Experimental models frequently concentrate on responses to a given model antigen or pathogen. The use of NGS holds the promise to broaden the perspective and to track global changes in the B-cell system during normal immune system development, in response to infection or vaccination, as well as in various disease states.
One important question in the field concerns the contribution of alternative IgA inductive sites to mucosal Ig production. There is a broad consensus that IgA induction takes place in classical secondary lymphoid tissues such as Peyer’s patches and flexible lymphoid follicles such as isolated lymphoid follicles. However, maturation of B-cell responses in the gut draining mesenteric lymph nodes and in situ CSR, i.e., IgA induction, in the intestinal lamina propria have also been described.11, 21, 22 Moreover, a recent report by Lycke and colleagues23 suggested that IgA responses appear synchronized throughout Peyer’s patches, which adds additional complexity to the spatio-temporal organization of intestinal B-cell responses. NGS allows comparing of the Ig repertoire in different compartments and at various time points. This type of information allows tracking mucosal B-cell responses and will contribute to better understanding of how Ig responses are integrated across various compartments and to target antigens to defined compartments for therapeutic purposes. Related to these aspects, in humans, who in contrast to mice have two IgA isotypes, there is an ongoing discussion on the role of local CSR in the gut lamina propria.15, 16 NGS alone might not finally settle this question but the construction and analysis of clonal trees (see below, “Analyzing B Cells Repertoires: Somatic Mutations and B-cell phylogeny”) comprising IgA1, IgA2, and other Ig isotype encoding sequences might shed new light on the interrelation of these isotypes. Other relevant questions concern the modes of IgA induction. IgA induction in the intestine can occur in T cell-dependent as well as T cell-independent processes, and is thought to involve various unique B-cell populations, such as B1 cells24 and transitional B cells,25 besides conventional B2 cells. Moreover, early B-cell development can occur in the gut lamina propria.26 In depth, analysis of Ig repertoire information will help to understand the contribution of alternative IgA-inducing pathways and B-cell subpopulations, B-cell differentiation, and diversification. Finally, lineage tree analysis can be used to track intestinal B cells at far higher resolution compared with classical Sanger sequencing-based studies. These insights will cast a high-resolution picture of antibody maturation in the gut immune system and might help the design of effective vaccination studies in the future.
At present, several NGS sequencing platforms are available, all of which easily yield several million reads at an affordable price. The advantages and disadvantages of NGS platforms have been reviewed in ref. 27). Yet, a particular challenge in using NGS to characterize intestinal B-cell responses arises from data analysis and interpretation. In particular, SHM poses a problem to the interpretation of Ig repertoire data, which is to distinguish bona-fide mutations introduced by SHM from sequencing errors introduced during sample preparation and NGS, as discussed below. Exploiting NGS requires the combination of classical immunological expertise with bioinformatics/systems immunology, a step which is not always easily accessible to the individual disciplines.
Besides technical aspects related to data analysis, special consideration needs to be given to the origin of starting material. Human blood samples are easily accessible. Yet, the B-cell repertoires observed in blood samples do not necessarily represent the overall repertoire in a human or mouse. Considering that 1 ml of human blood will contain more than a million B cells, blood-based Ig repertoire analysis seems suitable to provide a reasonable estimate of the Ig repertoire in circulating B cells. However, the circulating B-cell repertoire is a sort of averaged representation of different B-cell subsets and responses going on in various tissues. Thus, the Ig repertoire in blood does not necessarily reflect the Ig repertoire in the intestine, which is dominated by antigen-experienced plasma cells. Indeed, we observed that the IgA repertoire obtained from intestinal biopsies (not containing intestinal follicles) showed very low similarity to the Ig repertoire observed in the blood taken from the same individual (Thomsen and Pabst, unpublished data).
In addition, studies performed on human material are frequently limited to small samples. Thus, Ig repertoires observed in these samples might not always represent the complexity of the entire Ig repertoire even in the tissues of interest. Besides blood, gut biopsies can be easily obtained. Based on histological examination the number of plasma cells present in a regular-sized biopsy has been estimated to be ∼75,000 IgA-secreting plasma cells.28 Our own unpublished observations hint at a considerably lower number. Still, we may safely assume that the human gut is densely populated by IgA-secreting plasma cells, but by much lower numbers of B cells expressing other isotypes. Thus, gut biopsies represent particularly valuable material to study the repertoire of gut IgA secreting plasma cells.
In conclusion, along with computational questions related to the processing of NGS data sets, repertoire analysis needs to consider, first, the immunological questions related to the nature of the sampled material and, second, limitations in the amount of starting material that might have an impact on the relationship between the sampled repertoire to the full original repertoire.
Assessing the B-Cell Repertoire: Maintaining Quantitative Information
Several alternative approaches have been reported to obtain B-cell repertoire sequence information. Genomic DNA or RNA are used as starting material. In both cases the quantities of material required for NGS necessitate prior PCR amplification of the genes of interest. Thus, care needs to be taken to ensure representative amplification of all potential gene segments. This demand is met most easily by using a mixture of primers for amplification that anneal to the various alternative gene segments. If RNA is used as starting material, 5′ race offers an alternative to the use of primer pools and ensures unbiased amplification of the various VH gene families.29 However, the number of RNA molecules per cell varies greatly between individual cells even in the same pool, e.g., expression of Ig-encoding transcripts is much higher in plasma cells than in naive B cells. Thus, if the starting material contains various B-cell types, the number of sequences observed in RNA-based Ig repertoires does not reliably reflect cell numbers. Genomic DNA as starting material has the advantage of yielding sequence numbers that are more representative of cell numbers. In addition, DNA-based repertoire analysis produces sequences of the non-productively rearranged alleles, which can serve as a non-selected internal control. On the other hand, only RNA-based analysis can retain part of the constant region and thus retain information on the isotype expressed by each cell. Thus, the particular scientific questions determine whether DNA- or RNA-based repertoire analysis is more suitable.
Additional difficulties in obtaining quantitative information of cell numbers/sequence frequencies are inherent to the PCR technique. Variability in PCR amplification can result in dramatic skewing of sequence representation. In a PCR reaction of 20 cycles, each sequence may be amplified between 0 to 20 times, so that in a worst-case scenario the number of molecules obtained from a single molecule can vary between one and over a million. Early repertoire studies dealt with this problem by discarding all but one representative of each sequence/B-cell clone—this retains the V(D)J segment information but loses much information about repertoire diversity and sizes of clonally related B cell pools. An alternative is to use information on the number of unique sequences obtained per clone,30, 31, 32 which allows a rough assessment of clone size and diversity, as discussed below.
An elegant way to retain the quantitative information—and to also correct for PCR and sequencing errors—is to add random oligomers or “unique molecular identifiers” (UMIs) to the primers in the first reaction.33 UMIs are random sequences of sufficient complexity such that every primer contains a different UMI sequence. Thus, the “progeny” of every original sequence can be identified via its UMI, and counting UMIs allows a more reliable estimate of the true number of cells carrying BCRs with certain sequences. In addition, UMI can be used to obtain corrected sequences through a “majority vote” for every nucleotide position (eliminating errors that occurred in later replication cycles or in sequencing, and thus are seen only in a few copies of the sequence), and identical sequences coming from different cells (if using genomic DNA) can be enumerated—all via sophisticated computational analysis and error-correction algorithms.34 Figure 1 summarizes the main steps in analysing Ig-NGS data. The numbers of sequence reads in the raw data may vary from a few thousands per sample (when the source is DNA extracted from preserved biopsies) to millions (from fresh DNA or RNA samples). The reduction in the number of sequences during data cleaning, collapsing and error correction depends on the thresholds and criteria used; one may lose between 50–80% of the reads. The remaining sequences may represent highly diverse repertoires, out of the estimated 108 clones in a human—e.g., in blood samples—or be dominated by one or more large clones, as in samples from B-cell malignancies.
A flow chart showing the main choices and analysis steps required in processing next-generation sequencing (NGS) data of immunoglobulin genes. UMI, unique molecule identifiers. Cleanup, removal of all sequence that do not conform to basic criteria of quality, length, assignment to specific samples, primer quality, etc. Collapsing, if UMI are present, then artifact duplicate sequences can be removed while keeping the true duplicates (that have different UMIs). If UMIs are not present, then it is best to remove all duplicates and base the analysis on unique sequences. Segment assignment, identification of the most likely V(D)J segments comprising each sequence. Clone assignment, grouping sequences that likely originated from the same B-cell clone. Repertoire analysis, quantifying segment and segment combination usage and sharing. This is the basis for measuring repertoire diversity and comparing between repertoires (bottom of right path). For detailed mutation analysis (left path), one must also remove or correct as many errors and artifact insertions/deletions as possible. Lineage trees may then be created on the basis of the remaining (presumed real) mutations and tree shape may be analyzed. Further mutation analyses are most accurate when based on lineage tree shapes.
Analysing B-Cell Repertoires: Gene Segment Usage and Identification of Clonotypes
The analysis of NGS data requires the automated processing of sequence reads (Figure 2). Typically, sequences are processed through quality control filters. In a second step, sequences are assigned to different samples according to their MID (molecular identification) tags, which allow combining multiple samples in a single NGS sequence run. All of these steps can be performed by well-documented and easy-to-use resources.31 The third step is to extract the quantitative (and possibly pairing) information. At this step, sequence errors may be corrected, e.g., by the use of UMIs, as described in the previous section.
Structure of antibody chains and immunoglobulin (Ig) amplicons. (a) Combination of heavy and light chain V(D)J gene segments and junctional diversity—addition of N nucleotides yields more than 1012 different antibody encoding sequences. (b) Placement of forward and reverse primers to amplify Ig genes. Forward primers may be placed within the variable region or anneal to upstream sequences added by race PCR to the 5′-end. Reverse primers can be placed in conserved 3′- regions (e.g., the alternative J gene segments) or, in case of amplification from cDNA, in the isotype-specific constant regions. (c) Beside Ig gene-specific regions, forward and reverse primers comprise additional sequences allowing for next-generation sequencing (NGS) and data analysis. UMI, unique molecular identifier, were used to quantify NGS repertoire information and determine clonal sizes. MID, molecular identification tags, identify samples and allow combining multiple samples in one sequence run. (We prefer not to use the term “barcode”, as it has been alternately used for both MID tags and UMIs.) Adapter sequences are needed for technical reasons for the NGS sequencing platforms.
Subsequently, V, D and J segments (or V and J in case of light-chain repertoires) have to be identified. At present, the main database of Ig (and T-cell receptor) gene sequences is part of the IMmunoGeneTics set of databases and tools (http://www.imgt.org/). Tools for segment identification and junction analysis include IMmunoGeneTics’s own V-Quest,35, 36, 37, 38 NIH’s Ig-BLAST, JOINSOLVER,39 SoDA,40, 41 or iHMMune-align.42 The first four tools rely on IMmunoGeneTics, except for iHMMune-align; iHMMune-align relies on its own database, from which errors in IMmunoGeneTics were eliminated and some observed polymorphisms were added.43
Analysis of gene segment usage has been performed in various species by conventional Sanger sequencing and more recently by NGS. In all species, preferential gene segment usage was observed. A comprehensive analysis in this respect has been done in zebrafish.44, 45 The total number of B cells in an individual zebrafish is only about 300,000. Thus, NGS sequencing of the entire B-cell repertoire is feasible without the sampling problem inherent to human and, to some extent, also mouse studies. Full sequencing of the zebrafish B-cell repertoire revealed an unexpected preferential usage of only a few V(D)J combinations, in particular in young fish.44 Such stereotypic usage of gene segments became less obvious in mature zebrafish, but was still above the stochastically expected similarities in gene segment usage.45
Similarly, in humans and mice, waves of B-cell development seem to differ in their respective collection of gene segments preferentially used.46 Thus, we may conclude that the primary B-cell repertoire is less diverse than one might expect from the potential diversity that could be generated for the respective repertoire sizes.
The next step in sequence analysis is to identify the sequences belonging to clonally related groups (clonotypes, that is, sequences originating from one progenitor B cell). In T-cell receptors, clonal identity is established based on the V and J segments, and identical CDR3 junction regions. In BCRs, this process is more complex due to somatic mutations introduced by SHM. Hence, the information contained in the CDR3 region is usually not sufficient for clonal identification. Moreover, as identification of the original segments is also comfounded by SHM. No method for BCR clonal identification is 100% certain, although the probability of correct identification obviously increases with the number of unique sequences obtained from the clone.30 Clustering-based methods are at present the best for clonal identification, as they do not impose any artificial cutoffs;42 the cluster cutoff is dictated by the data.
When assessing gene segment usage and segment combinations, it is important to keep in mind which question is being addressed. If the focus is on B-cell development and BCR rearrangement, the relevant information is the number of clones using each gene segment combination, and hence clone sizes do not matter. On the other hand, when studying the peripheral repertoire and immune responses, clone sizes matter, as the larger clones may have been selected to expand. Furthermore, when assessing whether certain segment combinations or clonotypes are preferentially expanded, it is not sufficient to show that their numbers are higher than those of other clonotypes, because these numbers may be affected by rearrangement efficiency and PCR primer biases. Thus, to evaluate over- or underrepresentation of clonotypes, the frequency of certain segment combinations should be compared with the expected frequency if all gene segments were expressed independently.47 Under the latter assumption, the expected probability of observing a given combination VxJy, P(VxJy), is the product of the probabilities of observing each of these segments, i.e., P(Vx)*P(Jy); the latter probabilities are the observed frequencies of each segment in the database (see appendix in ref. 48). We have recently used this method to identify the overrepresented segment combinations in gastritis and gastric lymphomas; many of those we found in gastric lymphomas were already known to occur in other lymphomas, but new ones were also identified.47
Analysing B-Cell Repertoires: Diversity of the Intestinal B-Cell Repertoire
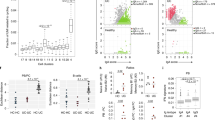
Further information obtained by NGS relates to Ig repertoire diversity. Ig repertoire diversity differs between anatomical sites and time points, and changes with aging and during immune responses.49, 50, 51, 52, 53 Thus, Ig repertoire diversity is characteristic of an individual’s B-cell compartment. Several techniques to address B-cell repertoire diversity have been established in the pre-NGS era (reviewed in ref. 54). A comprehensive view of global diversity can be obtained by measuring the length distribution of the CDR3 region by a method called spectratyping. For a highly diverse B-cell population, spectratyping will show a Gaussian distribution of CDR3 length, although the precise features of the distribution vary with age, tissue of origin, and disease.49, 53, 55 Notably, intestinal IgA plasma cells were reported to show a non-Gaussian distribution.56 This observation correlated with other studies that observed clonally related IgA encoding sequences by conventional Sanger sequencing even when only comparably few sequences had been analysed.57, 58 These observations were either interpreted to indicate the local proliferation of plasmablasts within the lamina propria28 or to suggest the dissemination of a previously expanded population of B cells throughout the intestine.57 The use of NGS has allowed us to revisit this issue. NGS sequencing of the intestinal IgA repertoire revealed that, indeed, a comparably low number of B-cell clones is highly expanded in the intestinal B-cell pool, although besides this expanded population a high number of non-expanded B cells is also present.59
To systematically assess Ig repertoire diversity, several groups have used diversity measures originally developed by ecologists to quantify habitat biodiversity (reviewed in ref. 48). As reports published so far used diversity measures in a rather sporadic manner, we performed a systematic evaluation of diversity measures, methods for estimating the diversity of the original repertoire a sample was taken from, and measures of similarity (or distance) between repertoires. We found that the best measure to use in every case depends on sample size and the resolution in which the repertoire is presented. For example, when the repertoire is examined only at the level of gene-segment combinations, then even in samples smaller than 104 sequences the estimated diversity is close to the full repertoire diversity, although at the level of amino-acid-sequence-only estimates based on sample sizes above 2 × 105 sequences reach the full repertoire diversity (Pickman and Mehr, unpublished data).
Analysing B-Cells Repertoires: Somatic Mutations and B-Cell Phylogeny
Somatic mutations pose a major problem to Ig repertoire analysis, which is to distinguish bona fide somatic mutations from PCR- and sequencing-derived mistakes. Typically, individual sequences are aligned to a library of template sequences. Somatic mutations complicate sequence alignments and sequence analysis has to be performed without a definitive template. Usually, the germline gene segments (V, J and sometimes D, if it can be identified with certainty) closest to the observed sequence are assumed to represent the gene segments recombined in the clone’s founder B cell. As there is no way to know the junction sequences in the original founder cell, the consensus CDR3 sequence is assumed to be that of the founder B cell, which minimizes the number of artifact mutations introduced by this choice.30 Mutations are considered valid if they pass defined sequencing quality criteria. In the case of 454 sequencing, which tends to create insertions and deletions (so called indels) near homopolymer tracts, these indels have to be evaluated; we have created a program that evaluates indels and mutations, and discards sequences carrying indels that are likely to be sequencing artifacts.31 This evaluation is based on whether the indel in question has or has not appeared in other (unique) sequences in the same clone. Where UMIs are used, legitimate mutations and indels can be distinguished with higher certainty from PCR and sequencing errors by consensus analysis, as explained above.
Finally, for every clonotype, somatic mutations can be ordered according to the most likely succession of mutations by creating lineage trees. The use of lineage trees to elucidate clonal relationships between B cells from different sources has preceded NGS.60, 61 For example, lineage tree analysis in ulcerative colitis patients established clonal relationships between B cells in the inflamed colon segments, in the non-inflamed margins of those segments, and in the nearby lymph nodes.62 Studies in mice investigating the response to oral vaccination revealed clonally related B cells in different Peyer’s patches and gut segments.23 Thus, drawing the lineage trees in itself is very useful to elucidate response dynamics.61, 63, 64, 65 Measurement of lineage tree properties has been shown to reveal the features of the humoral response, SHM, and selection. In addition, lineage tree analysis can be used to identify and enumerate somatic mutations more precisely than direct comparison of germline and observed sequence. The tree enables us to identify the most likely ancestor sequence to every mutation, count every mutation only once if it has only appeared once in the tree (even if it appears in many sequences), and identify reversion mutations. Figure 3 depicts an example of such analysis. We obtained small intestinal gut biopsies at 5, 9, 13, and 17 weeks of age from the same animal. The IgA repertoire was determined by NGS and used for lineage tree analysis (C. Linder, L. Hazanov, I. Iosselevitch, O. Pabst and R. Mehr, unpublished data). The depicted tree represents one out of many trees depicting the phylogeny of sets of clonally related B cells over time. Thus, lineage tree analysis offers a comprehensive way to track mucosal antibody responses in time and throughout compartments.
A sample lineage tree containing sequences from serial mouse gut biopsy samples. Biopsies have been obtained at 5, 9, 13, and 17 weeks of age from the same animal. The germline sequence represents for the original unmutated sequence, composed of the germline V(D)J segments and the clonal consensus junctional nucleotides. Open (non-colored) white nodes represent deduced, unobserved intermediate sequences. The different node colors indicate the mouse age (in weeks) at which each sequence was found (see legend at top left). More than one unique sequence may be assigned to a node, if the sequences differ only at the edges, or come from different samples (i.e., different mouse ages). In this tree, two such node are depicted as “tandem nodes”. Numbers next to edges indicate the number of mutations represented by each edge; edges without numbers represent one mutation.
The Gap between Sequence and Antigen Specificity
Notwithstanding the usefulness of repertoire and lineage tree analysis, the BCR/antibody encoding sequence in itself cannot be used to predict the nature of the antigen and specific epitope of the antibody. Moreover, the functionality of a given antibody needs to be analysed in a wider context than its binding affinity to a given antigen.
Results obtained by Ig sequencing (including seminal work performed with Sanger sequencing) seem to contradict a classical concept in IgA biology: the concept of natural gut IgA antibodies. Natural IgA antibodies are thought to bind a broad range of antigens, e.g., commensal bacteria, with only moderate affinity.10 Importantly, in analogy to natural IgM, natural IgA should be produced by germline sequences and lack somatic mutations. However, IgA encoding sequences obtained from human or murine gut plasma cells carry frequent somatic mutations.58, 59 Thus, IgA sequence information does not support the idea that natural IgA, defined as non-mutated IgA, makes a major contribution to the overall IgA pool.
To link BCR sequence information to the nature of the antigen, functional antibodies need to be produced based on the acquired sequences. This step is demanding because, first, today antibody expression and characterization are not yet compatible with high-throughput methods and, second, pairing information on heavy and light chain is required.
Paired VH/VL information can be obtained by single-cell PCR. In the most comprehensive of such studies on IgA antibodies, Wardemann and colleagues67 expressed >200 functional IgA antibodies in vitro. This antibody panel was tested for binding to a set of “typical” gut antigens which led the authors to conclude that most IgA antibodies are highly specific for their respective antigen.66 Recently, the Wardemann and colleagues67 further refined the single cell-based approach using barcoded primers to amplify VH and VL regions. Subsequently, amplicon pools representing a large number of single B cells were characterized by cost-saving NGS to combine the benefits of NGS with a straightforward technique to retain pairing VH/VL information and to express functional antibodies. Following standard bulk amplification of Ig-encoding genes from tissues or cell pools, information on VH/VL pairing is lost. At best, correlation of heavy and light chain repertoires can predict likely pairings for highly frequent sequences, an approach which was successfully taken to express functional antibodies from highly polarized Ig repertoires observed in immunized mice.68 However, compared with heavy chains, light chains are less reliable to identify clonotypes7 and B-cell development in the gut biases light chains in mice.26 Thus, direct assessment of VH/VL pairing is required to systematically extend Ig-repertoire information to antibody expression and antigen specificity. A sophisticated approach to preserve pairing information of the T-cell receptor α and βchains during PCR was based on cell emulsion PCR.69 Similarly, deposition of single B cells in microwell plates was successfully used to obtain matched information on VH/VL pairing in human blood B cells.70
Determining antigen specificity through antibody expression and screening can be accompanied by computational methods for structure determination. These approaches rely on the fact that the general antibody structure is known, and in many cases a similar antibody can be found; the structures of most CDR loops are rather preserved, although CDR3 presents a bigger challenge.71 Yet, even assuming the antibody structure is known, predicting the specificity in silico (in order to narrow the range of antigens that need to be screened) is a much more daunting challenge than predicting T-cell receptor epitopes. Unlike peptides presented by major histocompatibility complex molecules, BCR antigens (a) include all possible molecules, not only proteins; (b) appear in their native unprocessed form, rather than as linear peptides; and (c) may include conformational protein epitopes consisting of structurally adjacent amino acids that come from different parts of the sequence. Finally, antibody specificity might not be determined by the BCR-encoding sequence alone. In fact, IgA and the secretory component associated with the secretory Ig complex are highly glycosylated and glycans have been shown to contribute to antigen binding.72 Thus, Ig sequencing will certainly move the field forward, but has to go hand in hand with further technical and computational inventions.
Outlook
The last few years have seen enormous advances not only in the experimental techniques for Ig repertoire sequencing, but also in the computational methods and tools available to analyse sequencing data. As the technical issues reviewed here are being addressed, we are getting closer to a point where wet-lab and computational approaches can be standardized in a way that allows results from different experiments to be directly compared, and data may be joined for large meta-studies. We expect that along with a more complete understanding of gut plasma cell subset composition in health and disease, these advances will make it possible to clarify many remaining open questions regarding the generation and shaping of mucosal antibody repertoires in health, aging, and disease.
References
Brandtzaeg, P. Mucosal immunity: induction, dissemination, and effector functions. Scand. J. Immunol. 70, 505–515 (2009).
Brandtzaeg, P. & Baklien, K. Immunohistochemical studies of the formation and epithelial transport of immunoglobulins in normal and diseased human intestinal mucosa. Scand. J. Gastroenterol. Suppl. 36, 1–45 (1976).
Mantis, N.J., Rol, N. & Corthesy, B. Secretory IgA’s complex roles in immunity and mucosal homeostasis in the gut. Mucosal Immunol. 4, 603–611 (2011).
Woof, J.M. & Russell, M.W. Structure and function relationships in IgA. Mucosal Immunol. 4, 590–597 (2011).
Tonegawa, S. Somatic generation of antibody diversity. Nature 302, 575–581 (1983).
Perlot, T. & Alt, F.W. Cis-regulatory elements and epigenetic changes control genomic rearrangements of the IgH locus. Adv. Immunol. 99, 1–32 (2008).
Saada, R., Weinberger, M., Shahaf, G. & Mehr, R. Models for antigen receptor gene rearrangement: CDR3 length. Immunol. Cell Biol. 85, 323–332 (2007).
Jackson, K.J., Kidd, M.J., Wang, Y. & Collins, A.M. The shape of the lymphocyte receptor repertoire: lessons from the B cell receptor. Front. Immunol. 4, 263 (2013).
Spencer, J., Klavinskis, L.S. & Fraser, L.D. The human intestinal IgA response; burning questions. Front. Immunol. 3, 108 (2012).
Slack, E., Balmer, M.L., Fritz, J.H. & Hapfelmeier, S. Functional flexibility of intestinal IgA - broadening the fine line. Front. Immunol. 3, 100 (2012).
Pabst, O. New concepts in the generation and functions of IgA. Nat. Rev. Immunol. 12, 821–832 (2012).
Lycke, N. Y. IgA B cell responses to gut mucosal antigens: do we know it all? Front. Immunol. 4, 368 (2013).
Lozupone, C.A., Stombaugh, J.I., Gordon, J.I., Jansson, J.K. & Knight, R. Diversity, stability and resilience of the human gut microbiota. Nature 489, 220–230 (2012).
Casola, S. et al. B cell receptor signal strength determines B cell fate. Nat. Immunol. 5, 317–327 (2004).
He, B. et al. Intestinal bacteria trigger T cell-independent immunoglobulin A(2) class switching by inducing epithelial-cell secretion of the cytokine APRIL. Immunity 26, 812–826 (2007).
Lin, M., Du, L., Brandtzaeg, P. & Pan-Hammarstrom, Q. IgA subclass switch recombination in human mucosal and systemic immune compartments. Mucosal Immunol. 7, 511–520 (2014).
Kruglov, A.A. & Nedospasov, S.A. Microbiota, intestinal immunity, and mouse bustle. Acta Naturae 6, 6–8 (2014).
Wu, X. et al. Focused evolution of HIV-1 neutralizing antibodies revealed by structures and deep sequencing. Science 333, 1593–1602 (2011).
Jiang, N. et al. Lineage structure of the human antibody repertoire in response to influenza vaccination. Sci. Transl. Med. 5, 171ra119 (2013).
Georgiou, G. et al. The promise and challenge of high-throughput sequencing of the antibody repertoire. Nat. Biotechnol. 32, 158–168 (2013).
Macpherson, A.J., Geuking, M.B. & McCoy, K.D. Homeland security: IgA immunity at the frontiers of the body. Trends Immunol. 33, 160–167 (2012).
Suzuki, K., Maruya, M., Kawamoto, S. & Fagarasan, S. Roles of B-1 and B-2 cells in innate and acquired IgA-mediated immunity. Immunol. Rev. 237, 180–190 (2010).
Bergqvist, P. et al. Re-utilization of germinal centers in multiple Peyer’s patches results in highly synchronized, oligoclonal, and affinity-matured gut IgA responses. Mucosal Immunol. 6, 122–135 (2012).
Roy, B. et al. An intrinsic propensity of murine peritoneal B1b cells to switch to IgA in presence of TGF-beta and retinoic acid. PLoS ONE 8, e82121 (2013).
Vossenkamper, A. et al. A role for gut-associated lymphoid tissue in shaping the human B cell repertoire. J. Exp. Med. 210, 1665–1674 (2013).
Wesemann, D.R. et al. Microbial colonization influences early B-lineage development in the gut lamina propria. Nature 501, 112–115 (2013).
Metzker, M.L. Sequencing technologies - the next generation. Nat. Rev. Genet. 11, 31–46 (2010).
Yuvaraj, S. et al. Evidence for local expansion of IgA plasma cell precursors in human ileum. J. Immunol. 183, 4871–4878 (2009).
Choi, N.M. et al. Deep sequencing of the murine IgH repertoire reveals complex regulation of nonrandom V gene rearrangement frequencies. J. Immunol. 191, 2393–2402 (2013).
Michaeli, M., Barak, M., Hazanov, L., Noga, H. & Mehr, R. Automated analysis of immunoglobulin genes from high-throughput sequencing: life without a template. J. Clin. Bioinform. 3, 15 (2013).
Michaeli, M., Noga, H., Tabibian-Keissar, H., Barshack, I. & Mehr, R. Automated cleaning and pre-processing of immunoglobulin gene sequences from high-throughput sequencing. Front. Immunol. 3, 386 (2012).
Tabibian-Keissar, H. et al. PCR amplification and high throughput sequencing of immunoglobulin heavy chain genes from formalin-fixed paraffin-embedded human biopsies. Exp. Mol. Pathol. 94, 182–187 (2013).
Kivioja, T. et al. Counting absolute numbers of molecules using unique molecular identifiers. Nat. Methods 9, 72–74 (2011).
Bolotin, D.A. et al. Next generation sequencing for TCR repertoire profiling: platform-specific features and correction algorithms. Eur. J. Immunol. 42, 3073–3083 (2012).
Brochet, X., Lefranc, M.P. & Giudicelli, V. IMGT/V-QUEST: the highly customized and integrated system for IG and TR standardized V-J and V-D-J sequence analysis. Nucleic Acids Res. 36, W503–W508 (2008).
Giudicelli, V., Brochet, X. & Lefranc, M.P. IMGT/V-QUEST: IMGT standardized analysis of the immunoglobulin (IG) and T cell receptor (TR) nucleotide sequences. Cold Spring Harb. Protoc. 2011, 695–715 (2011).
Giudicelli, V., Chaume, D. & Lefranc, M.P. IMGT/V-QUEST, an integrated software program for immunoglobulin and T cell receptor V-J and V-D-J rearrangement analysis. Nucleic Acids Res. 32, W435–W440 (2004).
Lefranc, M.P. et al. IMGT, the international ImMunoGeneTics information system. Nucleic Acids Res. 37, D1006–D1012 (2009).
Souto-Carneiro, M.M., Longo, N.S., Russ, D.E., Sun, H.W. & Lipsky, P.E. Characterization of the human Ig heavy chain antigen binding complementarity determining region 3 using a newly developed software algorithm, JOINSOLVER. J. Immunol. 172, 6790–6802 (2004).
Munshaw, S. & Kepler, T.B. SoDA2: a Hidden Markov Model approach for identification of immunoglobulin rearrangements. Bioinformatics 26, 867–872 (2010).
Volpe, J.M., Cowell, L.G. & Kepler, T.B. SoDA: implementation of a 3D alignment algorithm for inference of antigen receptor recombinations. Bioinformatics 22, 438–444 (2006).
Gaeta, B.A. et al. iHMMune-align: hidden Markov model-based alignment and identification of germline genes in rearranged immunoglobulin gene sequences. Bioinformatics 23, 1580–1587 (2007).
Lee, C.E.H. et al. Reconsidering the human immunoglobulin heavy-chain locus: 1. An evaluation of the expressed human IGHD gene repertoire. Immunogenetics 57, 917–925 (2006).
Jiang, N. et al. Determinism and stochasticity during maturation of the zebrafish antibody repertoire. Proc. Natl Acad. Sci. USA 108, 5348–5353 (2011).
Weinstein, J.A., Jiang, N., White, R.A. 3rd, Fisher, D.S. & Quake, S.R. High-throughput sequencing of the zebrafish antibody repertoire. Science 324, 807–810 (2009).
Rogosch, T. et al. IgA response in preterm neonates shows little evidence of antigen-driven selection. J. Immunol. 189, 5449–5456 (2012).
Michaeli, M. et al. Immunoglobulin gene repertoire diversification and selection in the stomach - from gastritis to gastric lymphomas. Front. Immunol. 5, 264 (2014).
Mehr, R., Sternberg-Simon, M., Michaeli, M. & Pickman, Y. Models and methods for analysis of lymphocyte repertoire generation, development, selection and evolution. Immunol. Lett. 148, 11–22 (2012).
Ademokun, A., Wu, Y.C. & Dunn-Walters, D. The ageing B cell population: composition and function. Biogerontology 11, 125–137 (2010).
Ademokun, A. et al. Vaccination-induced changes in human B cell repertoire and pneumococcal IgM and IgA antibody at different ages. Aging Cell 10, 922–930 (2011).
Dunn-Walters, D.K. & Ademokun, A.A. B cell repertoire and ageing. Curr. Opin. Immunol. 22, 514–520 (2010).
Wu, Y.C. et al. High-throughput immunoglobulin repertoire analysis distinguishes between human IgM memory and switched memory B-cell populations. Blood 116, 1070–1078 (2010).
Gibson, K.L. et al. B-cell diversity decreases in old age and is correlated with poor health status. Aging Cell 8, 18–25 (2009).
Six, A. et al. The past, present, and future of immune repertoire biology - the rise of next-generation repertoire analysis. Front. Immunol. 4, 413 (2013).
Pickman, Y., Dunn-Walters, D. & Mehr, R. BCR CDR3 length distributions differ between blood and spleen and between old and young patients, and TCR distributions can be used to detect myelodysplastic syndrome. Phys. Biol. 10, 056001 (2013).
Stoel, M. et al. Restricted IgA repertoire in both B-1 and B-2 cell-derived gut plasmablasts. J. Immunol. 174, 1046–1054 (2005).
Holtmeier, W., Hennemann, A. & Caspary, W.F. IgA and IgM V(H) repertoires in human colon: evidence for clonally expanded B cells that are widely disseminated. Gastroenterology 119, 1253–1266 (2000).
Dunn-Walters, D.K., Boursier, L. & Spencer, J. Hypermutation, diversity and dissemination of human intestinal lamina propria plasma cells. Eur. J. Immunol. 27, 2959–2964 (1997).
Lindner, C. et al. Age, microbiota, and T cells shape diverse individual IgA repertoires in the intestine. J. Exp. Med. 209, 365–377 (2012).
Jacob, J. & Kelsoe, G. In situ studies of the primary immune response to (4-hydroxy-3-nitrophenyl)acetyl. II. A common clonal origin for periarteriolar lymphoid sheath-associated foci and germinal centers. J. Exp. Med. 176, 679–687 (1992).
Zuckerman, N.S. et al. Ectopic GC in the thymus of myasthenia gravis patients show characteristics of normal GC. Eur. J. Immunol. 40, 1150–1161 (2010).
Tabibian-Keissar, H. et al. B-cell clonal diversification and gut-lymph node trafficking in ulcerative colitis revealed using lineage tree analysis. Eur. J. Immunol. 38, 2600–2609 (2008).
Shahaf, G. et al. Antigen-driven selection in germinal centers as reflected by the shape characteristics of immunoglobulin gene lineage trees: a large-scale simulation study. J. Theor. Biol. 255, 210–222 (2008).
Zuckerman, N.S. et al. Ig gene diversification and selection in follicular lymphoma, diffuse large B cell lymphoma and primary central nervous system lymphoma revealed by lineage tree and mutation analyses. Int. Immunol. 22, 875–887 (2010).
Uduman, M., Shlomchik, M.J., Vigneault, F., Church, G.M. & Kleinstein, S.H. Integrating B cell lineage information into statistical tests for detecting selection in Ig sequences. J. Immunol. 192, 867–874 (2013).
Benckert, J. et al. The majority of intestinal IgA+ and IgG+ plasmablasts in the human gut are antigen-specific. J. Clin. Invest. 121, 1946–1955 (2011).
Busse, C.E., Czogiel, I., Braun, P., Arndt, P.F. & Wardemann, H. Single-cell based high-throughput sequencing of full-length immunoglobulin heavy and light chain genes. Eur. J. Immunol. 44, 597–603 (2014).
Reddy, S.T. et al. Monoclonal antibodies isolated without screening by analyzing the variable-gene repertoire of plasma cells. Nat. Biotechnol. 28, 965–969 (2010).
Turchaninova, M.A. et al. Pairing of T-cell receptor chains via emulsion PCR. Eur. J. Immunol. 43, 2507–2515 (2013).
DeKosky, B.J. et al. High-throughput sequencing of the paired human immunoglobulin heavy and light chain repertoire. Nat. Biotechnol. 31, 166–169 (2013).
Teplyakov, A. et al. Antibody modeling assessment II: structures and models. Proteins 82, 1563–1582 (2014).
Mathias, A. & Corthesy, B. N-Glycans on secretory component: mediators of the interaction between secretory IgA and gram-positive commensals sustaining intestinal homeostasis. Gut Microbes 2, 287–293 (2011).
Acknowledgements
We are indebted to Miri Michaeli, Irina Iosselevitch, and Olga Schulz for help in figure and manuscript preparation. This work was supported by Deutsche Forschungsgemeinschaft grant PA921/4-1 to O.P. and Israel Science Foundation grant 270/09, a Human Frontiers Science Program Research Grant and a grant from the Bar Ilan University–Sheba medical Center joint research fund to R.M.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declared no conflict of interest.
Rights and permissions
About this article
Cite this article
Pabst, O., Hazanov, H. & Mehr, R. Old questions, new tools: does next-generation sequencing hold the key to unraveling intestinal B-cell responses?. Mucosal Immunol 8, 29–37 (2015). https://doi.org/10.1038/mi.2014.103
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/mi.2014.103