Abstract
Symbiosis between microbes and their mammalian host is vital to maintaining homeostasis. Symbiotic microbes within the gastrointestinal tract provide an array of benefits to the host, including promotion of host immunity. A coordinated effort of the host and symbiotic microbes deters the colonization and survival of many invading pathogens. However, pathogens have devised strategies to overcome these mechanisms. Furthermore, some pathogens can hijack host hormones and bacterial autoinducers to induce virulence traits. Intra- and inter-species (bacteria/bacteria) and interkingdom (bacteria/host) communication orchestrates the complex relationship among symbiotic microbes, invading pathogens, and their mammalian host. Insight into this communication will provide a foundation for the development of targeted antimicrobial therapies.
Similar content being viewed by others
Introduction
Infectious diseases wreak havoc on mankind. The mammalian immune system has in place a line of defense specialized in recognizing and eradicating invading pathogens; however, sometimes the pathogen evades these mechanisms and establishes disease in its host. Therapies like antibiotics and vaccination abet the immune system in its fight against pathogenic microbes. Over time, resistance to antibiotics has developed because of the intense selective pressure the antibiotics place on bacteria. Furthermore, although a number of vaccines have been successful, far too many infectious diseases still do not have efficacious vaccines. A desperate call for new therapeutics exists.
Understanding the complex relationship among the host, symbiotic microbes, and invading pathogens will provide important insight for the rational design of therapeutics. Bacteria can communicate with one another through hormone-like signals to modulate their gene expression1 in a process termed quorum sensing (QS).2 Additionally, these bacterial signals can modify mammalian cell-signal transduction,3 and likewise host hormones can cross signal to regulate bacterial gene expression4 in a process termed interkingdom signaling. Interference with the cell-to-cell signaling pathways that control bacterial virulence offers a promising new strategy in the treatment of bacterial infections. This review will discuss both the mechanisms employed by the host and symbiotic bacteria to impair pathogen virulence, as well as the conserved cell-to-cell signaling pathways implemented by pathogens that allow for exploitation of their host environment.
Antimicrobial Strategies Enlisted by the Host and Symbionts
The human gut hosts an estimated 500–1,000 species of bacteria.5, 6 A mutually beneficial relationship exists between the human intestine and many of its symbionts: the human intestine provides nutrients to the resident bacteria, whereas bacteria aid in the digestion of food and absorption of nutrients, produce vitamins such as biotin and vitamin K, regulate immune system function, and hinder the colonization of pathogenic microorganisms.7 Two major bacterial phyla, Firmicutes and Bacteroidetes, and five minor bacterial phyla, Proteobacteria, Actinobacteria, Fusobacteria, Cyanobacteria, and Verrucomicrobia, compose the intestinal gut flora in adult humans.8 Variations in gut microbiota, however, can result from genetic and environmental factors such as diet, living conditions, and birthplace.6
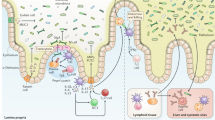
In a concerted effort, the host and symbiotic bacteria employ the use of physical, chemical, and cell-mediated antimicrobial strategies to prevent or impair pathogen survival and virulence. Physical barriers provide the first line of resistance against pathogens. The intestinal mucosal barrier, composed of a thick mucus layer, a layer of epithelial cells, and an underlying layer of cells composed predominantly by immune cells, provides both a physical and chemical barrier through the secretion of mucins, immunoglobulins, antimicrobial peptides, and lectins.9 Especially important in barrier function, immunoglobulins secreted by B-lymphocytes aid in the opsonization of microbes and prevent microbial penetration of the mucosal layer.9 Resident bacteria provide another crucial line of defense against the colonization of pathogens by competing for nutrition and attachment sites to the colonic epithelium, a mechanism known as the “barrier effect.”10
Intestinal epithelial cells produce two major classes of antimicrobial peptides: defensins and cathelicidins.11, 12 Defensins function by embedding into the microbial membrane to form pores that allow for the efflux of essential ions and nutrients.13 In addition to their antimicrobial activity, defensins and cathelicidins modulate the host immune response,14 in large part by forming local chemotactic gradients that promote the mobilization of leukocytes.12 Angiogenins represent another important class of antimicrobial proteins. Whereas mouse Ang1 and human ANG exhibit bactericidal and fungicidal activity against systemic pathogens, Ang4 acts selectively in the gut against enteric pathogens. Secreted by Paneth cells into the gut lumen, Ang4 expression is induced by the intestinal bacteria Bacteroides thetaiotamicron.15 Also influenced by the intestinal flora, the mouse C-type lectin RegIIIγ and its human counterpart HIP/PAP exert their bactericidal activity against Gram-positive bacteria via an interaction with peptidoglycan16 and are vital to antimicrobial protection in the mammalian gut.17
Symbiotic microbes provide another source of antimicrobial molecules within the mucosal barrier. Through the production of butyrate, a short-chain fatty acid that is the only known stimulus for cathelicidin expression,18 the enteric microflora aids in the stimulation of host innate immunity. The short-chain fatty acid has also been shown to decrease Shiga toxin expression in enterohemorrhagic Escherichia coli (EHEC).19 Additionally, symbiotic microbes can interfere with pathogen survival and growth through the production of potent toxins called bacteriocins. For example, Lactobacillus salivarius UCC118 produces the bacteriocin ABP-118 that is active against food-borne pathogens, including Bacillus, Listeria, Enterococcus, and Staphylococcus species.20, 21
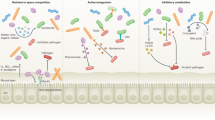
Although both the host and symbiotic microbes have evolved mechanisms to prevent pathogen invasion and colonization, similarly pathogens have devised means to subvert and even exploit their environment.
Quorum Sensing
Bacteria respond to hormone-like molecules called autoinducers to regulate specific target genes, a process known as QS.1, 2 To date, four main categories of cell-to-cell signaling systems have been studied in detail. Gram-negative bacteria communicate in response to autoinducer-1 (AI-1) and autoinducer-3 (AI-3), whereas Gram-positive bacteria use an autoinducing polypeptide system.22 Autoinducer-2 (AI-2) acts as a “universal” signal for interspecies communication and is found in both Gram-negative and Gram-positive cells.23 Because of space constraints, only the AI-1 and AI-3 cell signaling systems will be discussed in this review.
The LuxI/LuxR System
The foundation of QS, AI-1 and its cognate signaling system, the LuxI/LuxR system, was initially discovered in the bioluminescent marine bacterium Vibrio fisheri and its host, the Hawaiian Bobtail Squid Euprymna scolopes.1, 24, 25, 26 LuxI synthesizes AI-1, an N-acyl homoserine lactone (AHL) that can diffuse freely across the bacterial membrane. The simultaneous production of AI-1 by a growing population of bacteria increases the local concentration of AI-1, and at high population density, AI-1 diffuses back into the cell. Inside the cell, AI-1 binds the transcriptional activator LuxR that, in its bound state to AI-1, can activate the transcription of the luxCDABEGH operon.27
Homologs to the LuxI/LuxR system have been identified in many Gram-negative bacteria. For example, the opportunistic human pathogen, Pseudomonas aeruginosa, has two homologous LuxI/LuxR systems: the LasI/LasR system (Figure 1) and the RhlI/RhlR system.28 P. aeruginosa produces two AHLs: 3-oxo-C12 homoserine lactone (HSL) that acts on the las system and C4-HSL that acts on the rhl system. LasI produces 3-oxo-C12-HSL to activate LasR29, 30, 31 that leads to the production of virulence factors like elastase30 and pyoverdin.32 RhlI synthesizes C4-HSL to activate RhlR,33, 34, 35 allowing for the production of rhamnolipid biosurfactants36 and a number of other virulence factors important in biofilm formation and pathogenesis.37
The LasI/LasR quorum-sensing system in Pseudomonas aeruginosa. (a) LasI synthesizes 3-oxo-C12 homoserine lactone (HSL), anN-acyl homoserine lactone (AHL) that freely diffuses across the bacterial membrane at low cell density. (b) At high cell density, 3-oxo-C12 HSL diffuses back into the cell, binds to the transcriptional activator LasR, and induces the expression of virulence genes.
Interestingly, both E. coli and Salmonella typhimurium encode for a LuxR homolog named SdiA but do not have a LuxI homolog.38, 39 The absence of a LuxI homolog indicates that neither E. coli nor S. typhimurium can produce its own AI-1. Instead, presence of SdiA may allow for these bacteria to respond to AI-1 made by other bacteria, including AI-1 produced by the resident flora in the gastrointestinal tract where both of these pathogens colonize.
Pathogenesis in the Gut
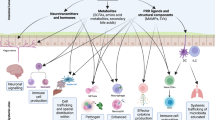
The gastrointestinal tract is a diverse and dynamic environment, home to large communities of bacterial flora, and constantly threatened by opportunistic and pathogenic microbes. Within the gastrointestinal tract, bacteria communicate with one another and with their host to coordinate expression of key genes. Hormonal communication between the host and microorganisms is termed inter-kingdom signaling.40 One example of an inter-kingdom signaling system is the AI-3/epinephrine/norepinephrine signaling system.4
Commensal E. coli, as well as the intestinal bacterial species EHEC, enteropathogenic E. coli, Klebsiella pneumoniae, Shigella sp., Salmonella sp., and Enterobacter cloacae, produce AI-3.41 The mammalian hormones epinephrine and norepinephrine are also found in the intestine.42 Although the primary roles of epinephrine and norepinephrine are to modulate smooth muscle contraction, submucosal blood flow, and chloride and potassium secretion,43 enteric pathogens like EHEC O157:H7 can hijack these host hormones to induce virulence genes and promote colonization in the intestine.4
The QseBC (quorum sensing E. coli regulators B and C) two-component QS system can detect AI-3, epinephrine, and norepinephrine.44 Upon sensing its signal, the histidine sensor kinase QseC stimulates an intrinsic autophosphorylation activity that results in phosphorylation of a conserved histidine residue present in the cytoplasm (Figure 2a). The phosphate group is then transferred to a conserved aspartate residue on three response regulators: QseB, KdpE, and QseF.45 Once phosphorylated, the response regulators differentially regulate gene expression. QseB activates the master flagellar regulator genes flhDC to regulate flagella biosynthesis and motility.46 KdpE controls potassium uptake, osmolarity, and the formation of AE (attaching and effacing) lesions.45 QseF regulates the bacterial SOS stress response, as well as the formation of AE lesions.45 The genes responsible for AE lesion formation are contained within the LEE (locus of enterocyte effacement), a chromosomal pathogenicity island responsible for much of EHEC's virulence; in addition, the LEE encodes for a type III secretion system and secreted effectors.47, 48, 49
The epinephrine/norepinephrine/autoinducer-3 (AI-3) interkingdom signaling system. (a) The histidine sensor kinase QseC senses the mammalian hormones epinephrine and norepinephrine and the bacterial autoinducer AI-3. Activation of QseC stimulates an intrinsic autophosphorylation activity, allowing QseC to then transfer its phosphate group to one of three response regulators: QseB, QseF, and KdpE. The response regulators differentially regulate gene expression. (b) Epinephrine and norepinephrine bind to mammalian adrenergic receptors and have a role in smooth muscle control, submucosal blood flow, and chloride (Cl) and potassium (K) secretion. It remains unknown if AI-3 can bind to mammalian adrenergic receptors and alter mammalian function.
QseC is a functional analog of an adrenergic receptor, and its activity can be blocked by the α-adrenergic antagonist phentolamine.44 Interestingly, the α2-adrenergic receptor is expressed at high levels in the proximal and transverse colon of the human intestine, the site at which EHEC and commensal flora colonize.50 It remains unknown if AI-3 can bind to mammalian adrenergic receptors (Figure 2b). If so, this could be another mechanism by which EHEC exploits the intestinal environment to promote its colonization.
The importance of QseC is highlighted by its conservation in a number of bacterial species, including Salmonella sp., Shigella flexneri, Vibrio parahaemolyticus, Yersinia pestis, and Francisella tularensis.51 In a mutant strain of S. typhimurium deficient in QseC, S. typhimurium has impaired flagellar motility and reduced invasion and survival in macrophages.52 Furthermore, mice deficient in dopamine β-hydroxylase that are unable to produce epinephrine or norepinephrine have different susceptibility to Salmonella infection, indicating a role for AI-3/epinephrine/norepinephrine in inter-kingdom signaling.52
Pathogenesis in the lungs
P. aeruginosa is an opportunistic human pathogen that colonizes the lungs of cystic fibrosis patients. P. aeruginosa produces two AHLs as QS signaling molecules, 3-oxo-C12-HSL and C4-HSL, as discussed previously, that act on the las and rhl systems, respectively.29, 30, 31, 33, 34, 35 In addition to the AHL-based QS systems, P. aeruginosa uses an autoinducer regulatory system based on 2-alkyl-4(1H)-quinolones (AQs). The AQ biosynthetic enzymes of P. aeruginosa enable the generation of a diverse range of related AQ molecules: HQNO (2-heptyl-4-hydroxyquinoline N-oxide), HHQ (4-hydroxy-2-heptylquinoline), and PQS (Pseudomonas quinolone signal (3,4-dihydroxy-2-heptylquinoline)).53, 54 PQS induces the expression of pqsABCDE55 and is required for the expression of phzA1-G1, the gene responsible for pyocyanin production. Release of pyocyanin induces neutrophil apoptosis and epithelial cell damage,56 allowing P. aeruginosa to subvert immune surveillance and gain residency in the lungs. PQS also controls the expression of the lasB (elastase) gene,57 and a synergistic effect is achieved in the presence of both PQS and C4-HSL.58 Furthermore, PQS increases expression of the lecA gene that encodes for PA-I lectin (PA-IL).59
In addition to bacterial autoinducers, P. aeruginosa activates virulence genes in response to the opioid dynorphin.60 Endogenous opioids are among the first signals released during times of stress.61 Similar to the ability of EHEC to respond to the stress hormones epinephrine and norepinephrine, P. aeruginosa exploits the host during a weakened state for its own benefit. Dynorphin synergizes with PQS to increase expression of pqsABCDE and induce expression of the AQs HQNO and HHQ. Additionally, dynorphin enhances virulence against the probiotic Lactobacillus species and the nematode Caenorhabditis elegans.60
In addition to P. aeruginosa, AQs have been identified in a number of species of Burkholderia, including B. ambifaria, B. thailandensis, and B. pseudomallei. These organisms produce 3-methyl derivatives of PHQ, HHQ, and NHQ termed 4-hydroxy-3-methyl-2-alkylquinolones.62 Because AQs also act as antibiotics, as evidenced by the ability of AQs produced by P. aeruginosa to inhibit the growth of Staphylococcus aureus and Candida albicans,63 it remains to be determined if Burkholderia sp. use AQs solely as antibiotics or if they also use AQs as signaling molecules.
Conclusions
Bacterial communities reside on the skin and on every mucosal surface in the human body. Although many bacteria benefit the host's well-being, opportunistic and pathogenic bacteria await times of stress to exploit their host. Defining pathways unique to opportunistic and pathogenic bacteria will allow for the development of therapies that target specifically the virulence associated with these bacteria.
Since Alexander Fleming's discovery of the antibiotic penicillin in 1928, a number of additional antibiotics have been isolated from living organisms or synthesized. Unfortunately, because of the intense selective pressure placed on bacteria by antibiotics, a number of multidrug-resistant bacteria have emerged. Pathogens utilize an array of virulence strategies to promote colonization and cause disease in their hosts, including expression of adhesins, production of toxins, and secretion of effectors through specialized secretion systems. An alternative strategy to antibiotics is the development of antimicrobial drugs that target microbial virulence instead of growth.64 Several such drugs are being tested in both laboratory and clinical settings against pathogens like Bacillus anthracis,65, 66 S. typhimurium, EHEC, F. tularensis,51, 64 S. aureus,67 and P. aeruginosa.68, 69
Because many of the QS pathways are conserved across bacterial species, a single therapy could be designed to target multiple pathogens. However, much research remains to be done to evaluate the risk of such therapies on resident flora that may also signal through these pathways. It is crucial that we seek new therapies against microbial pathogens as the incidence of resistance to current antibiotics rapidly rises. As we gain further understanding of the complex relationship among the host, symbiotic microbes, and invading pathogens, we will be better able to rationally design therapies that specifically target virulence traits incurred by pathogens.
References
Nealson, K.H., Platt, T. & Hastings, J.W. Cellular control of the synthesis and activity of the bacterial luminescent system. J. Bacteriol. 104, 313–322 (1970).
Fuqua, C., Winans, S.C. & Greenberg, E.P. Census and consensus in bacterial ecosystems: the LuxR-LuxI family of quorum-sensing transcriptional regulators. Annu. Rev. Microbiol. 50, 727–751 (1996).
Telford, G. et al. The Pseudomonas aeruginosa quorum-sensing signal molecule N-(3-oxododecanoyl)-L-homoserine lactone has immunomodulatory activity. Infect. Immun. 66, 36–42 (1998).
Sperandio, V., Torres, A.G., Jarvis, B., Nataro, J.P. & Kaper, J.B. Bacteria-host communication: the language of hormones. Proc. Natl. Acad. Sci. USA 100, 8951–8956 (2003).
Hooper, L.V. & Gordon, J.I. Commensal host-bacterial relationships in the gut. Science 292, 1115–1118 (2001).
Eckburg, P.B. et al. Diversity of the human intestinal microbial flora. Science 308, 1635–1638 (2005).
Backhed, F., Ley, R.E., Sonnenburg, J.L., Peterson, D.A. & Gordon, J.I. Host-bacterial mutualism in the human intestine. Science 307, 1915–1920 (2005).
Palmer, C., Bik, E.M., DiGiulio, D.B., Relman, D.A. & Brown, P.O. Development of the human infant intestinal microbiota. PLoS Biol. 5, e177 (2007).
McGuckin, M.A., Eri, R., Simms, L.A., Florin, T.H. & Radford-Smith, G. Intestinal barrier dysfunction in inflammatory bowel diseases. Inflamm. Bowel. Dis. 15, 100–113 (2009).
Guarner, F. & Malagelada, J.R. Gut flora in health and disease. Lancet 361, 512–519 (2003).
Lehrer, R.I., Ganz, T. & Selsted, M.E. Defensins: endogenous antibiotic peptides of animal cells. Cell 64, 229–230 (1991).
Yang, D., Chertov, O. & Oppenheim, J.J. Participation of mammalian defensins and cathelicidins in anti-microbial immunity: receptors and activities of human defensins and cathelicidin (LL-37). J. Leukoc. Biol. 69, 691–697 (2001).
Hazlett, L. & Wu, M. Defensins in innate immunity. Cell Tissue Res. (2010). (e-pub ahead of print 21 August 2010).
Bowdish, D.M., Davidson, D.J. & Hancock, R.E. Immunomodulatory properties of defensins and cathelicidins. Curr. Top Microbiol. Immunol. 306, 27–66 (2006).
Hooper, L.V., Stappenbeck, T.S., Hong, C.V. & Gordon, J.I. Angiogenins: a new class of microbicidal proteins involved in innate immunity. Nat. Immunol. 4, 269–273 (2003).
Lehotzky, R.E. et al. Molecular basis for peptidoglycan recognition by a bactericidal lectin. Proc. Natl. Acad. Sci. USA 107, 7722–7727 (2010).
Cash, H.L., Whitham, C.V., Behrendt, C.L. & Hooper, L.V. Symbiotic bacteria direct expression of an intestinal bactericidal lectin. Science 313, 1126–1130 (2006).
Schauber, J. et al. Expression of the cathelicidin LL-37 is modulated by short chain fatty acids in colonocytes: relevance of signalling pathways. Gut 52, 735–741 (2003).
Carey, C.M., Kostrzynska, M., Ojha, S. & Thompson, S. The effect of probiotics and organic acids on Shiga-toxin 2 gene expression in enterohemorrhagic Escherichia coli O157:H7. J. Microbiol. Methods 73, 125–132 (2008).
Claesson, M.J. et al. Multireplicon genome architecture of Lactobacillus salivarius. Proc. Natl. Acad. Sci. USA 103, 6718–6723 (2006).
Dunne, C. et al. In vitro selection criteria for probiotic bacteria of human origin: correlation with in vivo findings. Am. J. Clin. Nutr. 73, 386S–392S (2001).
Parker, C.T. & Sperandio, V. Cell-to-cell signalling during pathogenesis. Cell Microbiol. 11, 363–369 (2009).
Schauder, S., Shokat, K., Surette, M.G. & Bassler, B.L. The LuxS family of bacterial autoinducers: biosynthesis of a novel quorum-sensing signal molecule. Mol. Microbiol. 41, 463–476 (2001).
Engebrecht, J., Nealson, K. & Silverman, M. Bacterial bioluminescence: isolation and genetic analysis of functions from Vibrio fischeri. Cell 32, 773–781 (1983).
Engebrecht, J. & Silverman, M. Identification of genes and gene products necessary for bacterial bioluminescence. Proc. Natl. Acad. Sci. USA 81, 4154–4158 (1984).
Ruby, E.G. Lessons from a cooperative, bacterial-animal association: the Vibrio fischeri-Euprymna scolopes light organ symbiosis. Annu. Rev. Microbiol. 50, 591–624 (1996).
Devine, J.H., Shadel, G.S. & Baldwin, T.O. Identification of the operator of the lux regulon from the Vibrio fischeri strain ATCC7744. Proc. Natl. Acad. Sci. USA 86, 5688–5692 (1989).
de Kievit, T.R. & Iglewski, B.H. Bacterial quorum sensing in pathogenic relationships. Infect. Immun. 68, 4839–4849 (2000).
Gambello, M.J. & Iglewski, B.H. Cloning and characterization of the Pseudomonas aeruginosa lasR gene, a transcriptional activator of elastase expression. J. Bacteriol. 173, 3000–3009 (1991).
Passador, L., Cook, J.M., Gambello, M.J., Rust, L. & Iglewski, B.H. Expression of Pseudomonas aeruginosa virulence genes requires cell-to-cell communication. Science 260, 1127–1130 (1993).
Pearson, J.P. et al. Structure of the autoinducer required for expression of Pseudomonas aeruginosa virulence genes. Proc. Natl. Acad. Sci. USA 91, 197–201 (1994).
Stintzi, A., Evans, K., Meyer, J.M. & Poole, K. Quorum-sensing and siderophore biosynthesis in Pseudomonas aeruginosa: lasR/lasI mutants exhibit reduced pyoverdine biosynthesis. FEMS Microbiol. Lett. 166, 341–345 (1998).
Latifi, A. et al. Multiple homologues of LuxR and LuxI control expression of virulence determinants and secondary metabolites through quorum sensing in Pseudomonas aeruginosa PAO1. Mol. Microbiol. 17, 333–343 (1995).
Pearson, J.P., Passador, L., Iglewski, B.H. & Greenberg, E.P. A second N-acylhomoserine lactone signal produced by Pseudomonas aeruginosa. Proc. Natl. Acad. Sci. USA 92, 1490–1494 (1995).
Winson, M.K. et al. Multiple N-acyl-L-homoserine lactone signal molecules regulate production of virulence determinants and secondary metabolites in Pseudomonas aeruginosa. Proc. Natl. Acad. Sci. USA 92, 9427–9431 (1995).
Ochsner, U.A. & Reiser, J. Autoinducer-mediated regulation of rhamnolipid biosurfactant synthesis in Pseudomonas aeruginosa. Proc. Natl. Acad. Sci. USA 92, 6424–6428 (1995).
Pearson, J.P., Pesci, E.C. & Iglewski, B.H. Roles of Pseudomonas aeruginosa las and rhl quorum-sensing systems in control of elastase and rhamnolipid biosynthesis genes. J. Bacteriol. 179, 5756–5767 (1997).
Michael, B., Smith, J.N., Swift, S., Heffron, F. & Ahmer, B.M. SdiA of Salmonella enterica is a LuxR homolog that detects mixed microbial communities. J. Bacteriol. 183, 5733–5742 (2001).
Hughes, D.T. et al. Chemical sensing in mammalian host-bacterial commensal associations. Proc. Natl. Acad. Sci. USA 107, 9831–9836 (2010).
Hughes, D.T. & Sperandio, V. Inter-kingdom signalling: communication between bacteria and their hosts. Nat. Rev. Microbiol. 6, 111–120 (2008).
Walters, M., Sircili, M.P. & Sperandio, V. AI-3 synthesis is not dependent on luxS in Escherichia coli. J. Bacteriol. 188, 5668–5681 (2006).
Eldrup, E. & Richter, E.A. DOPA, dopamine, and DOPAC concentrations in the rat gastrointestinal tract decrease during fasting. Am. J. Physiol. Endocrinol. Metab. 279, E815–822 (2000).
Horger, S., Schultheiss, G. & Diener, M. Segment-specific effects of epinephrine on ion transport in the colon of the rat. Am. J. Physiol. 275, G1367–1376 (1998).
Clarke, M.B., Hughes, D.T., Zhu, C., Boedeker, E.C. & Sperandio, V. The QseC sensor kinase: a bacterial adrenergic receptor. Proc. Natl. Acad. Sci. USA 103, 10420–10425 (2006).
Hughes, D.T., Clarke, M.B., Yamamoto, K., Rasko, D.A. & Sperandio, V. The QseC adrenergic signaling cascade in Enterohemorrhagic E. coli (EHEC). PLoS Pathog. 5, e1000553 (2009).
Clarke, M.B. & Sperandio, V. Transcriptional regulation of flhDC by QseBC and sigma (FliA) in enterohaemorrhagic Escherichia coli. Mol. Microbiol. 57, 1734–1749 (2005).
Elliott, S.J., Yu, J. & Kaper, J.B. The cloned locus of enterocyte effacement from enterohemorrhagic Escherichia coli O157:H7 is unable to confer the attaching and effacing phenotype upon E. coli K-12. Infect. Immun. 67, 4260–4263 (1999).
Jarvis, K.G. et al. Enteropathogenic Escherichia coli contains a putative type III secretion system necessary for the export of proteins involved in attaching and effacing lesion formation. Proc. Natl. Acad. Sci. USA 92, 7996–8000 (1995).
Mellies, J.L., Elliott, S.J., Sperandio, V., Donnenberg, M.S. & Kaper, J.B. The Per regulon of enteropathogenic Escherichia coli: identification of a regulatory cascade and a novel transcriptional activator, the locus of enterocyte effacement (LEE)-encoded regulator (Ler). Mol. Microbiol. 33, 296–306 (1999).
Valet, P. et al. Characterization and distribution of alpha 2-adrenergic receptors in the human intestinal mucosa. J. Clin. Invest. 91, 2049–2057 (1993).
Rasko, D.A. et al. Targeting QseC signaling and virulence for antibiotic development. Science 321, 1078–1080 (2008).
Moreira, C.G., Weinshenker, D. & Sperandio, V. QseC mediates Salmonella enterica serovar typhimurium virulence in vitro and in vivo. Infect. Immun. 78, 914–926 (2010).
Lepine, F., Deziel, E., Milot, S. & Rahme, L.G. A stable isotope dilution assay for the quantification of the Pseudomonas quinolone signal in Pseudomonas aeruginosa cultures. Biochim. Biophys. Acta. 1622, 36–41 (2003).
Lepine, F., Milot, S., Deziel, E., He, J. & Rahme, L.G. Electrospray/mass spectrometric identification and analysis of 4-hydroxy-2-alkylquinolines (HAQs) produced by Pseudomonas aeruginosa. J. Am. Soc. Mass Spectrom. 15, 862–869 (2004).
Wade, D.S. et al. Regulation of Pseudomonas quinolone signal synthesis in Pseudomonas aeruginosa. J. Bacteriol. 187, 4372–4380 (2005).
Lau, G.W., Hassett, D.J., Ran, H. & Kong, F. The role of pyocyanin in Pseudomonas aeruginosa infection. Trends Mol. Med. 10, 599–606 (2004).
Pesci, E.C. et al. Quinolone signaling in the cell-to-cell communication system of Pseudomonas aeruginosa. Proc. Natl. Acad. Sci. USA 96, 11229–11234 (1999).
McKnight, S.L., Iglewski, B.H. & Pesci, E.C. The Pseudomonas quinolone signal regulates rhl quorum sensing in Pseudomonas aeruginosa. J. Bacteriol. 182, 2702–2708 (2000).
Diggle, S.P. et al. The Pseudomonas aeruginosa quinolone signal molecule overcomes the cell density-dependency of the quorum sensing hierarchy, regulates rhl-dependent genes at the onset of stationary phase and can be produced in the absence of LasR. Mol. Microbiol. 50, 29–43 (2003).
Zaborina, O. et al. Dynorphin activates quorum sensing quinolone signaling in Pseudomonas aeruginosa. PLoS Pathog. 3, e35 (2007).
Peterson, P.K., Molitor, T.W. & Chao, C.C. The opioid-cytokine connection. J. Neuroimmunol. 83, 63–69 (1998).
Vial, L. et al. Burkholderia pseudomallei, B. thailandensis, and B. ambifaria produce 4-hydroxy-2-alkylquinoline analogues with a methyl group at the 3 position that is required for quorum-sensing regulation. J. Bacteriol. 190, 5339–5352 (2008).
Machan, Z.A., Taylor, G.W., Pitt, T.L., Cole, P.J. & Wilson, R. 2-Heptyl-4-hydroxyquinoline N-oxide, an antistaphylococcal agent produced by Pseudomonas aeruginosa. J. Antimicrob. Chemother. 30, 615–623 (1992).
Rasko, D.A. & Sperandio, V. Anti-virulence strategies to combat bacteria-mediated disease. Nat. Rev. Drug. Discov. 9, 117–128 (2010).
Karginov, V.A., Nestorovich, E.M., Moayeri, M., Leppla, S.H. & Bezrukov, S.M. Blocking anthrax lethal toxin at the protective antigen channel by using structure-inspired drug design. Proc. Natl. Acad. Sci. USA 102, 15075–15080 (2005).
Shoop, W.L. et al. Anthrax lethal factor inhibition. Proc. Natl. Acad. Sci. USA 102, 7958–7963 (2005).
Kiran, M.D. et al. Discovery of a quorum-sensing inhibitor of drug-resistant staphylococcal infections by structure-based virtual screening. Mol. Pharmacol. 73, 1578–1586 (2008).
Lesic, B. et al. Inhibitors of pathogen intercellular signals as selective anti-infective compounds. PLoS Pathog. 3, 1229–1239 (2007).
Muh, U. et al. Novel Pseudomonas aeruginosa quorum-sensing inhibitors identified in an ultra-high-throughput screen. Antimicrob. Agents Chemother. 50, 3674–3679 (2006).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declared no conflict of interest.
PowerPoint slides
Rights and permissions
About this article
Cite this article
Curtis, M., Sperandio, V. A complex relationship: the interaction among symbiotic microbes, invading pathogens, and their mammalian host. Mucosal Immunol 4, 133–138 (2011). https://doi.org/10.1038/mi.2010.89
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/mi.2010.89
This article is cited by
-
Enhancement of immune response against Bordetella spp. by disrupting immunomodulation
Scientific Reports (2019)
-
Mechanistic insights into the host-microbe interaction and pathogen exclusion mediated by the Mucus-binding protein of Lactobacillus plantarum
Scientific Reports (2018)
-
The rumen microbiome as a reservoir of antimicrobial resistance and pathogenicity genes is directly affected by diet in beef cattle
Microbiome (2017)
-
Interactions between the microbiota and pathogenic bacteria in the gut
Nature (2016)
-
The Role of the QseC Quorum-Sensing Sensor Kinase in Epinephrine-Enhanced Motility and Biofilm Formation by Escherichia coli
Cell Biochemistry and Biophysics (2014)