Abstract
Traditional approaches for gene mapping from candidate gene studies to positional cloning strategies have been applied for Mendelian disorders. Since 2005, next-generation sequencing (NGS) technologies are improving as rapid, high-throughput and cost-effective approaches to fulfill medical sciences and research demands. Using NGS, the underlying causative genes are directly distinguished via a systematic filtering, in which the identified gene variants are checked for novelty and functionality. During the past 2 years, the role of more than 100 genes has been distinguished in rare Mendelian disorders by means of whole-exome sequencing (WES). Combination of WES with traditional approaches, consistent with linkage analysis, has had the greatest impact on those disorders following autosomal mode of inheritance; in more than 60 identified genes, the causal variants have been transmitted at homozygous or compound heterozygous state. Recent literatures focusing on identified new causal genes in Mendelian disorders using WES are reviewed in the present survey.
Similar content being viewed by others
Introduction
One of the major concerns of medical sciences is finding the causal genes underlying human diseases. New technologies are developed, and progress in elucidating genetic basis of disorders is now one of the most discussing topics in medical genomics. With the advent of next-generation sequencing (NGS) technology, identification of genetic variations that serve as disease causality is progressing at rapid pace, which would improve the disease management either by available treatments or genetic counseling for the children’s health and risk assessment of the relatives.1 Molecular diagnosis, carrier detection, prenatal diagnosis and developing new therapies are the concern for the health care and society. Besides, unraveling the susceptible variants potentiates new interventions and prevention of the disease caused by different risk factors.
Generally, genetic disorders are categorized into monogenic and multifactorial disorders. Monogenic (single gene) disorders include simple and rare disorders. Multifactorial disorders comprise of complex disorders, multiple genes, as well as lifestyle or environmental factors, that are contributed to the disease. Rare genetic disorders have low prevalence of about at most 6.5 out of every 10 000 individuals according to the World Health Organization.2, 3, 4, 5 Tremendous efforts are now being performed for understanding the rare monogenic and complex traits and manifesting the genetic basis of the disease based on exome (all exons in the genome) sequencing.
Up to 27 January 2012, a total of 21 058 entries were reported in OMIM (Online Mendelian Inheritance in Man),6 describing 13 790 genes and 4535 disorders with known molecular basis (http://omim.org/statistics/geneMap). Approximately, 1800 entries had phenotypic descriptions or known loci with unknown molecular bases and 2000 entries have been stated based on suspected Mendelian basis, and mainly the phenotypes are known. Bearing in mind, the Mendelian disorders that explore the novel genetic mechanisms, phenotypic variability, modifier genes, allelic variations and genetic variations of the diseases, may also provide clues in understanding the complex disorders.7, 8 Here, we focus on those diseases that are caused by single genes which their causal variants have been explored by exome sequencing. Sporadic cases are also included in this survey.
From DNA structure report to new NGS reports
Nearly 10 years after the discovery of DNA structure, the first gene was completely sequenced.9 In 1977, Sanger et al.10 and Maxam and Gilbert11 developed initial sequencing methods (Table 1); meanwhile, the majority of human DNA sequence data has been described using the Sanger sequencing and fluorescence-based electrophoresis technologies. With the development of a revolutionary method named PCR by Kary Mullis in the 1980s, molecular genomic field has undergone enormous advances. As a matter of fact, a growing variety of molecular methods, including high-throughput sequencing technologies, has been emerged over the past 7 years. In 2004, the second (next)-generation sequencing methods, massively parallel sequencing platforms, were introduced and the next revolution in molecular genetics, including finding the disease-causing genes, is expected. Most of these platforms rely on sequencing by synthesis, and generation of their clonally clustered amplicons are achieved mainly through in situ polonies, emulsion PCR or bridge PCR. NGS technology affords high speed and throughput, both qualitative and quantitative sequence data, equivalent to the data from human genome project, in 10–20 days. Several different ways are employed in which NGS is being applied for identifying causal gene variant in the rare diseases. Whole-genome sequencing (WGS), whole-exome sequencing (WES), transcriptome sequencing, methylome and other sequencing approaches are applied in NGS systems.
Miller syndrome is the first rare Mendelian disorder from which its causal variants were identified, owing to the development of WES.12 These researchers explained DHODH mutations in three affected pedigrees after filtering against public single-nucleotide polymorphism (SNP) databases and eight HapMap exomes.
There are increasing numbers of reports identifying the causal variants of the diseases. More than 100 causative genes in various Mendelian disorders have been identified by means of exome sequencing. Up to May 2012, a PUBMED search on Mendelian disorders using exome sequencing revealed 102 diseases as summarized in Table 2. A total of 326 exomes have been reported in these successful applications of WES in identifying novel causative genes. In all, 61 out of these 108 identified genes (56.5%) follow autosomal recessive, 40 of them are transmitted as autosomal dominant (37%), one case is X-linked recessive and one case follows X-linked dominant inheritance. About 35.2% of these genes (38 out of 108) have been identified by WES with only one exome, which was confirmed by sequencing of the possible identified genes in other patients. In overall, about three exomes (108/326) might be enough to be sequenced for identification of causative gene of the disease (Table 2). Evidently, we have no information about the unsuccessful WES studies. In addition to gene discovery of dominant and recessive diseases, WES has been used for determining somatic mutations in tumors and rare mutations with moderate effect in common diseases as well as clinical diagnoses.13 As mentioned previously, here, we focus on the single-gene disorders.
Exome sequencing and identifying causal variants
Traditionally, the single-gene disorders were first analyzed based on linkage analyses14, 15 followed by positional cloning; an informative segregation pattern, clear mode of inheritance and enough affected family members could support for gene identification. Homozygosity mapping establishes loci of autosomal recessive disorders.16 More complex forms of single-gene disorders, such as retinitis pigmentosa17 and hearing loss, with different inheritance modes have been reported based on SNP arrays. Allelic association studies of case-control design are suitable for identifying highly associated SNPs with the complex diseases.
Drawbacks and limitations to these approaches that hindered the gene discovery need to be emphasized; there are families with small number of affected individuals, which do not meet the criteria required for classical gene-discovery methods. In addition, finding the causal genes in families fitting the criteria is very difficult in case of expression variability, locus heterogeneity, phenotypic heterogeneity, reduced penetrance or reduced fitness, because in these conditions, the causal effect could hardly be co-segregated with affection status within the family. Exome sequencing permits to overcome these obstacles. Also, there may be several sporadic cases from different families with similar phenotypes, in which exome sequencing interrogates the causal variants. As in Table 2, WES could identify the causal variants with a limited number of patients. Indeed, NGS technologies bring us new sights in unraveling the genetic basis of diseases.
Most pathogenic variants thus far identified are located in highly conserved regions of the genome.18, 19 It is believed that most of the functional variants are located in the coding exons.20 Most (91.8%) of the functional variants of the protein-coding variations are due to nonsense/missense (∼56%), small insertion/deletions (∼24%), splicing (∼10%) and regulatory (∼1.8%) mutations (Human Gene Mutation Database professional 2011).20 Overall, 85% of the disease-causing mutations are estimated to be located at protein-coding regions.20, 21 Accordingly, WES could elucidate at least 78% of causative variants.
Basics of WES
The approach for exome sequencing is based on probe-hybridization method to capture entire exons.22, 23 The whole process is categorized into three steps, namely sample preparation, hybridization and sequencing (Figure 1). Briefly, the first step is sample preparation, in which the genomic DNA is sheared by nebulization or sonication to get desired fragments of about 250 bp. The fragment ends are repaired by T4 DNA ligase. The process of 3′ A-tailing is performed followed by ligation of paired-end adaptor to the fragments. The final step for sample preparation is to amplify the prepared library for a few cycles. To enrich the prepared library, hybridization with a biotinylated oligo library (RNA baits for example, Agilant SureSelect (Santa Clara, CA, USA) (635 250 RNA probes of 120 bp) or DNA baits, for example, NimbleGen22 (Madison, WI, USA) (2.1 million DNA probes of 60–90 bp)) is performed and captured by streptavidin beads. The quality and quantity of the exome library is analyzed by highly sensitive methods, such as Agilent 2100 Bioanalyzer before sequencing step. The exome library is sequenced in paired-end reads for example in Illumina (San Diego, CA, USA) to yield a 75–100 bases per read. Amplification of surface-bound individual fragments using an isothermal bridge amplification method produces clonal clusters of about 1000 identical molecules per cluster; one fragment is, therefore, attached to one surface oligonucleotide, endures cluster generation, and the replicate copies are sequenced to yield one sequence read. When DNA chain is growing, the first step of sequencing procedure is detecting the next added fluorescently labeled base (reversible terminator) by means of a sensitive device like charge-coupled device camera. The terminator is changed to a standard nucleotide by removing the dye. Repeating this cycle, sequentially, determines the next base. About 79% of the reported genes were determined using Illumina sequencing machines (Table 2).
Applying usual filtering to exome-sequencing projects would define novel causal genes for Mendelian disorders; major assumptions about causal genes at these steps are as following: (1) structural variants and other forms of genetic variations are ignored, (2) causal variants are coding, (3) causal variants alter protein sequence and (4) casual variant has almost complete penetrance. A single exome carries about 20 000–30 000 coding SNPs. Over 95% of the variants overlap with data sets depending on ethnicity. Filtering steps narrow down the number of possible disease-associated genes; then, the final variants are limited to those fitting the mode of inheritance. A full color version of this figure is available at the Journal of Human Genetics journal online.
After sequencing, the data is processed in three major steps, including mapping, variant calling and annotation steps (Figure 1). The sequence data is aligned with Burrows–Wheeler Aligner24 tool against a reference sequence such as hg18/hg19 (GRCh37). Next step is calling; data generated by Burrows–Wheeler Aligner in Sequence Alignment Map (SAM) format could be used by SAMTools,25 Genome Analysis Tool Kit,26 and Picard (http://picard.sourceforge.net). SAMTools is used for quality control, short read alignment and variant identification (VarFilter). It processes and sorts the files. Facilitating the short aligned reads (BAM files: binary equivalent SAM format) for fast access is called indexing, which is followed by making a pileup format to facilitate variant calling. The indexed file is visualized by Integrative Genomics Viewer27 or other sequence alignment visualization tools. PCR duplicates are removed using Picard MarkDuplicate and SAMTools. Average coverage and depth of coverage are calculated with Genome Analysis Tool Kit’s Depth of Coverage analysis. ANNOVAR28 is a tool for annotating genetic variants based on the function; the annotation file usually includes gene name, chromosomal position, nucleotide changes, amino-acid changes and description, SIFT (sorting intolerant from tolerant)29 and Polyphen (polymorphism phenotyping)30 values, single-nucleotide polymorphism database ID, allelic frequency of the SNP in 1000 Genome project and sequence quality. VAAST (Variant Annotation, Analysis and Search Tool) incorporated previous amino-acid substitution information with annotation and ranked candidate genes with statistical evaluation, which can be used to list up the candidate genes and variants.31 Most investigators filter the data based on the function of variants. Nearly half of variants are synonymous ones, not considered to be deleterious, which are usually filtered out. Although there are some reports about the causal effect of synonymous variants,32 the probability is very low. The remaining variants are nonsense, missense, indel, splice mutations and other non-coding RNA transcripts. Approximately, 5% of the variants are not reported in the above databases.33 As noted, the variants called based on pathogenic predication of bioinformatics tools, such as SIFT29 and PolyPhen,30 are explored through the annotation. Hence, the pathogenic variants disrupt the protein function or structure in conserved sites. Depending on the knowledge of the affected samples, different analytic frameworks are used to define the causal variant (Figure 2).
Hypothetical frameworks for analyzing single-gene disorders. Combinational analyses could help to determine the probable causative variant. Family-based (a1, a2 and a3), de novo mutation (b) and X-linked variant analysis (c). A full color version of this figure is available at the Journal of Human Genetics journal online.
Calling variants and the candidate gene
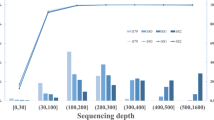
The sequence data are compared with public databases, such as single-nucleotide polymorphism database,34 1000 Genome Project,35 Exome Variant Server (http://evs.gs.washington.edu/EVS) and HapMap.36 It is noticed that individual exome of African-American origin has an average 24 000 single-nucleotide variants, whereas European-Americans origin has a mean 20 000 single-nucleotide variantss.33 Thus, it is inferred from other studies that this number varies depending on the ethnicity and capturing protocols, sequencing platforms, mapping algorithms and variant calling methods. Totally, the number of candidate disease-causing variants is reduced to 100–500 pathogenic variants depending on the study design.18, 36, 37, 38, 39 In a study, it was reported that each genome carries 165 homozygous protein-truncating variants in the diverse pathways.40 Thus, a causal variant cannot be directly identified as the related gene unless integrative genomic analyses are performed as homozygosity mapping, linkage analysis, Sanger sequencing and so on. However, if we study a family with four affected individuals or two or three families, each with at least two affected ones, employing the usual filtering could possibly define a causal gene.41, 42, 43, 44, 45 According to an assumption by Robinson et al.,46 when the same gene is considered as a causality in multi sporadic cases, 5% of the target genes (about 20 000) show rare probable casual variants in all affected individuals, and after sequencing one individual and a usual filtering, nearly 1000 genes would remain as candidate genes. If a second individual is sequenced, only 50 genes (5% of 1000) with variants in both individuals will remain. Sequencing a third affected person predicts less than one gene having a variant in all three affected individuals.46
Frameworks used to analyze single-gene disorders
We, here, exemplify the main approaches for detecting gene variants among the Mendelian disorders. Two main approaches, including family-based and unrelated individual strategies, are explained (Figure 2).
Family-based studies
When a number of affected individuals within a family are sequenced, the shared mutations are selected from the affected members because they harbor the same causal variant. This strategy narrow downs the candidate genes. Non-affected members of the family are sequenced to verify the candidate variations.
Combining the previous knowledge of homozygosity mapping for recessive disorders (Figure 2a2) and linkage studies for dominant (Figure 2a3) and recessive disorders as integrative approaches define the candidate variant. For instance, homozygous regions of the genome detected by SNP array are informative to reduce the number of candidate variants found to be homozygous for a family with recessive inheritance; only those variants in homozygous regions are reliable for pathogenicity. Shared homozygous or compound heterozygous variants are used to find the candidate variant (Figures 2a1 and a2). In a study by Sirmaci et al.,47 the cause of Malpuech–Michels–Mingarelli–Carnevale syndrome in two affected families was identified. An autozygous region on chromosome 3q.27 was identified and exome sequencing confirmed MASP1 mutation co-segregated with the phenotype.
Linkage studies are informative for multiple affected family members with multigenerations and are used in combination with exome sequencing.48 For the dominant disorders, a common heterozygous variant is distinguished among the affected individuals in a family. Using the genome-wide linkage analysis of hereditary diffuse leukoencephalopathy with spheroids affecting central nervous system, Rademakers et al.49 identified 233 candidate genes within the chromosome 5q candidate region and exome sequencing revealed a heterozygous variant in CSF1R in the candidate region, which was confirmed in 13 other affected families with distinct heterozygous mutations.
In case of locus heterogeneity in genetic disorders, such as retinitis pigmentosa, osteogenesis imperfect, hearing loss and so on, different patterns of inheritance may be observed in different families; thus, differentiating the exact clinical descriptions and determining the mode of inheritance would help to find the candidate gene. In a study by Abou Jamra et al.,50 a combination of autozygosity mapping and exome sequencing was applied to identify the pathogenic variants, causing intellectual disability with recessive mode of inheritance in eight affected individuals from three consanguineous families. Using this approach, they identified three causative genes encoding adaptor protein complex 4 within these families.
In case of X-linked pedigrees (Figure 2c), analysis of X-chromosome variants could be helpful; of course, female and male samples are homozygous and hemizygous for autosomal recessive disorders, respectively. In an example, exome sequencing of entire three affected males having an unclassified X-linked lethal congenital malformation syndrome identified a splicing mutation in OFD1 gene.51
Unrelated individual studies
If there are a number of affected cases, but not within the family (sporadic cases), common pathogenic gene could be followed among the samples, assuming no locus heterogeneity among the affected individuals. The approach is called overlap strategy.52 To point out, the cause of Schinzel–Giedion syndrome was identified using this strategy.53 Furthermore, Saitsu et al.54 performed exome sequencing in three unrelated affected individuals with congenital hypomyelination leukoencephalopathy; they found compound heterozygous mutations in POLR3A and POLR3B (encoding RNA polymerase III subunits) in the affected individuals.
As in case of Kabuki syndrome, which is a rare disorder worldwide, the majority of cases are sporadic, but parent-to-child transmission has been reported representing the dominant mode of inheritance. WES revealed the causal variants of Kabuki syndrome in 7 out of 10 families in MLL2; follow-up Sanger sequencing of the remaining three families detected mutations in the MLL2 in two of the families, which shows that this would be the main cause of syndrome.39 Other genes, however, may explain the pathogenesis of the condition in the remaining family. One may use this family in conjunction with other new affected families to find the causal variants. In addition, they reported that only 26 out of 43 patients had mutations in MLL2 by Sanger sequencing. The clinical and genetic heterogeneity of the syndrome complicates gene finding, and similar clinically affected cases are helpful to find new genes.
In the absence of number of cases, some integrative strategies are needed depending on their availability. When there is a single affected individual suspected of a recessive disorder, homozygous and compound heterozygous variants are searched namely, as double hit strategy by Gilissen et al.52 WES revealed compound heterozygous mutations in WDR35 gene in a single sporadic case affected with Sensenbrenner syndrome.55 Also, WES of one of the affected sisters with Perrault syndrome manifested two mutations, showing compound heterozygote in the HSD17B4.38
De novo mutations
The previously mentioned frameworks are focused on homogenous diseases. A substantial number of de novo mutations occur sporadically in which cases, mostly fetus, do not survive and so the mutations will be eliminated from the population; thus, these lethal mutations are not usually identified. De novo mutation rate is estimated to be 7.6 × 10−9–2.2 × 10−8 per generation; that is, approximately one in 108 base per haploid genome is mutated spontaneously,56, 57 which could become the causality. For the de novo mutation analysis, the case–parent trios are practical (Figure 2b). Nonpathogenic variants are filtered and then the variants presented in the parents are excluded. There might be a chance of sequencing errors and mapping artefacts, so confirmation by Sanger sequencing with high accuracy should be applied.58 Detection of a de novo mutation is not enough to confirm the causality of the disease. Additional analyses, including replication and functional analyses, should be performed to determine the deleterious or causal variants. Pathogenesity of a variant not only depends on the type and the location of the mutation but also on its functional effects.19
De novo mutation studies have been employed to determine de novo mutations of rare Mendelian disorders, such as Schinzel–Giedion syndrome.53 In addition, causative genetic factors for heterogeneous disorders, such as intellectual disabilities, have been revealed as well.18 Trio-based exome sequencing is demonstrated to be a powerful approach for identifying novel causative genes for sporadic autism spectrum;59 these researchers identified 21 de novo mutations using exome sequencing of 20 sporadic cases of the sporadic autism spectrum.59 More recently, Sanders et al.60 demonstrated, using WES of families affected by autism spectrum disorder, contribution of de novo mutations in brain-expression genes to the risk for these disorders. Iossifov et al.61 sequenced and analyzed the exomes of 343 families with a single individual affected by the autism spectrum and at least one unaffected sibling; they found that gene-disrupting mutations, not missense mutations, are frequent in affected children. In this study, 350–400 genes have been estimated as autism-susceptibility genes.61
Pros and cons
Interpretation of the results obtained by NGS of the Mendelian disorders is of major concern. When using exome sequencing in clinical genetics and medicine, limitation of the approach is evident and experimental design is needed to circumvent the downs. Genetic and phenotypic heterogeneity in different affected individuals make exome sequencing difficult to interpret. Exact clinical examinations and biochemical tests have important roles to distinguish between new syndromes and known ones.
Patients with the same phenotype may not share the same causal variant; indeed they may have distinct variants in a gene as we call allelic heterogeneity. Depending on the clinical information, different strategies or filtering protocols are used for implication of the pathogenicity of a variant. Of course, variants in the reported genes are generally examined at the first step of filtering process. Then, the possible causal variants could be validated by segregating through the family or other cases.
Intensive examinations of variant call are important in exome sequencing; false-positive errors appear as sequencing errors related to mechanical and analytical errors; also short reads generated by NGS would not align perfectly to the appropriate position as a result of paralogous and low copy repeats that may cause errors during calling.62 In the repetitive regions of the genome, misalignment may occur, which could be improved by longer read lengths or higher depth of coverage in those regions.21 Also, false-negative errors could occur because of mechanical and analytic errors due to low coverage, poor capture efficiency and so on. Avoiding false-negative errors is more difficult than false-positive ones; however, it is proposed that the error rate could be estimated by comparing the calls with the test samples, which have been previously called.21, 46 The relation of false-positive and false-negative calls is ‘trade-off’; if we set stringent criteria (high base quality and stringent alignment or and so on), false-positive errors are decreased but false-negative ones are increased, which is important to concern. There are also other problems in the filtering strategies, which may influence data analysis. Filtering out variants with minor allele frequency of <1% may be misleading for recessive disorders. Because the carriers may not show the disease, but still the frequency of the allele is high in the 1000 Genome, which may be excluded in the filtering step because of higher frequency.
Some deleterious variations may be located in the non-coding regions, such as intronic or regulatory regions of the genome, which cannot be called by exome sequencing, whereas WGS covers all the data for genome. WGS is expected to be applied for disease-gene identification in near future; however, the current cost and information burden of WGS need to be circumvented. The huge amount of data generated from WGS comparing with WES provides information on evolutionary-conserved non-coding regions and all variants throughout the genome. Filtering and analyzing these data is challenging. Moreover, the time for analyzing data is increased and larger computational memory is needed for WGS data analysis.
Conclusion
Exome sequencing has evolved the biomedical research. The possible causative genes are directly distinguished using these new sequencing technologies. Up to now, the role of more than 100 genes has been distinguished in rare Mendelian disorders by means of WES, and this statistics is rapidly growing. Combinational approaches, including traditional methods and WES, are easily used for those disorders following autosomal mode of inheritance to define the underlying gene. It is needless to say, new sequencing technologies, such as in Pacific Bioscience and Nanopore, will shed light in this field.63, 64 WES could have a critical role in identifying new genes until the costs for WGS will be decreased.
References
Green, E. D. & Guyer, M. S. Charting a course for genomic medicine from base pairs to bedside. Nature 470, 204–213 (2011).
Reich, D. E. & Lander, E. S. On the allelic spectrum of human disease. Trends Genet. 17, 502–510 (2001).
Lander, E. S. Initial impact of the sequencing of the human genome. Nature 470, 187–197 (2011).
Pritchard, J. K. Are rare variants responsible for susceptibility to complex diseases? Am. J. Hum. Genet. 69, 124–137 (2001).
Gorlov, I. P., Gorlova, O. Y., Sunyaev, S. R., Spitz, M. R. & Amos, C. I. Shifting paradigm of association studies: value of rare single-nucleotide polymorphisms. Am. J. Hum. Genet. 82, 100–112 (2008).
McKusick, V. A. Mendelian Inheritance in Man and its online version, OMIM. Am. J. Hum. Genet. 80, 588–604 (2007).
Antonarakis, S. E. & Beckmann, J. S. Mendelian disorders deserve more attention. Nat. Rev. Genet. 7, 277–282 (2006).
Peltonen, L., Perola, M., Naukkarinen, J. & Palotie, A. Lessons from studying monogenic disease for common disease. Hum Mol Genet 15Spec No 1, R67–R74.
Holley, R. W. The nucleotide sequence of a nucleic acid. Sci. Am. 214, 30–39 (1966).
Sanger, F., Nicklen, S. & Coulson, A. R. DNA sequencing with chain-terminating inhibitors. Proc. Natl Acad. Sci. USA 74, 5463–5467 (1977).
Maxam, A. M. & Gilbert, W. A new method for sequencing DNA. Proc. Natl Acad. Sci. USA 74, 560–564 (1977).
Ng, S. B., Buckingham, K. J., Lee, C., Bigham, A. W., Tabor, H. K., Dent, K. M. et al. Exome sequencing identifies the cause of a mendelian disorder. Nat. Genet. 42, 30–35 (2010).
Lindhurst, M. J., Sapp, J. C., Teer, J. K., Johnston, J. J., Finn, E. M., Peters, K. et al. A mosaic activating mutation in AKT1 associated with the Proteus syndrome. N. Engl. J. Med. 365, 611–619 (2011).
Riordan, J. R., Rommens, J. M., Kerem, B., Alon, N., Rozmahel, R., Grzelczak, Z. et al. Identification of the cystic fibrosis gene: cloning and characterization of complementary DNA. Science 245, 1066–1073 (1989).
Gusella, J. F., Wexler, N. S., Conneally, P. M., Naylor, S. L., Anderson, M. A., Tanzi, R. E. et al. A polymorphic DNA marker genetically linked to Huntington's disease. Nature 306, 234–238 (1983).
Lander, E. S. & Botstein, D. Homozygosity mapping: a way to map human recessive traits with the DNA of inbred children. Science 236, 1567–1570 (1987).
Pomares, E., Riera, M., Permanyer, J., Mendez, P., Castro-Navarro, J., Andres-Gutierrez, A. et al. Comprehensive SNP-chip for retinitis pigmentosa-leber congenital amaurosis diagnosis: new mutations and detection of mutational founder effects. Eur. J. Hum. Genet. 18, 118–124 (2010).
Vissers, L. E., de Ligt, J., Gilissen, C., Janssen, I., Steehouwer, M., de Vries, P. et al. A de novo paradigm for mental retardation. Nat. Genet. 42, 1109–1112 (2010).
Cooper, D. N., Chen, J. M., Ball, E. V., Howells, K., Mort, M., Phillips, A. D. et al. Genes, mutations, and human inherited disease at the dawn of the age of personalized genomics. Hum. Mutat. 31, 631–655 (2011).
Botstein, D. & Risch, N. Discovering genotypes underlying human phenotypes: past successes for mendelian disease, future approaches for complex disease. Nat. Genet. 33 (Suppl), 228–237 (2003).
Majewski, J., Schwartzentruber, J., Lalonde, E., Montpetit, A. & Jabado, N. What can exome sequencing do for you? J. Med. Genet. 48, 580–589 (2011).
Hodges, E., Xuan, Z., Balija, V., Kramer, M., Molla, M. N., Smith, S. W. et al. Genome-wide in situ exon capture for selective resequencing. Nat. Genet. 39, 1522–1527 (2007).
Gnirke, A., Melnikov, A., Maguire, J., Rogov, P., LeProust, E. M., Brockman, W. et al. Solution hybrid selection with ultra-long oligonucleotides for massively parallel targeted sequencing. Nat. Biotechnol. 27, 182–189 (2009).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
Li, H., Handsaker, B., Wysoker, A., Fennell, T., Ruan, J., Homer, N. et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
McKenna, A., Hanna, M., Banks, E., Sivachenko, A., Cibulskis, K., Kernytsky, A. et al. The Genome Analysis Toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res. 20, 1297–1303 (2010).
Robinson, J. T., Thorvaldsdottir, H., Winckler, W., Guttman, M., Lander, E. S., Getz, G. et al. Integrative genomics viewer. Nat. Biotechnol. 29, 24–26 (2011).
Wang, K., Li, M. & Hakonarson, H. ANNOVAR: functional annotation of genetic variants from high-throughput sequencing data. Nucleic Acids Res. 38, e164 (2010).
Kumar, P., Henikoff, S. & Ng, P. C. Predicting the effects of coding non-synonymous variants on protein function using the SIFT algorithm. Nat. Protoc. 4, 1073–1081 (2009).
Adzhubei, I. A., Schmidt, S., Peshkin, L., Ramensky, V. E., Gerasimova, A., Bork, P. et al. A method and server for predicting damaging missense mutations. Nat. Methods 7, 248–249 (2010).
Yandell, M., Huff, C., Hu, H., Singleton, M., Moore, B., Xing, J., Jorde, L. B. et al. A probabilistic disease-gene finder for personal genomes. Genome Res. 21, 1529–1542 (2011).
Sauna, Z. E. & Kimchi-Sarfaty, C. Understanding the contribution of synonymous mutations to human disease. Nat. Rev. Genet. 12, 683–691 (2011).
Bamshad, M. J., Ng, S. B., Bigham, A. W., Tabor, H. K., Emond, M. J., Nickerson, D. A. et al. Exome sequencing as a tool for Mendelian disease gene discovery. Nat. Rev. Genet. 12, 745–755 (2011).
Sherry, S. T., Ward, M. H., Kholodov, M., Baker, J., Phan, L., Smigielski, E. M. et al. dbSNP: the NCBI database of genetic variation. Nucleic Acids Res. 29, 308–311 (2001).
Consortium, G. P. A map of human genome variation from population-scale sequencing. Nature 467, 1061–1073 (2010).
Altshuler, D. M., Gibbs, R. A., Peltonen, L., Dermitzakis, E., Schaffner, S. F., Yu, F. et al. Integrating common and rare genetic variation in diverse human populations. Nature 467, 52–58 (2010).
Musunuru, K., Pirruccello, J. P., Do, R., Peloso, G. M., Guiducci, C., Sougnez, C. et al. Exome sequencing, ANGPTL3 mutations, and familial combined hypolipidemia. N. Engl. J. Med. 363, 2220–2227 (2010).
Pierce, S. B., Walsh, T., Chisholm, K. M., Lee, M. K., Thornton, A. M., Fiumara, A. et al. Mutations in the DBP-deficiency protein HSD17B4 cause ovarian dysgenesis, hearing loss, and ataxia of Perrault syndrome. Am. J. Hum. Genet. 87, 282–288 (2010).
Ng, S. B., Bigham, A. W., Buckingham, K. J., Hannibal, M. C., McMillin, M. J., Gildersleeve, H. I. et al. Exome sequencing identifies MLL2 mutations as a cause of Kabuki syndrome. Nat. Genet. 42, 790–793 (2010).
Pelak, K., Shianna, K. V., Ge, D., Maia, J. M., Zhu, M., Smith, J. P. et al. The characterization of twenty sequenced human genomes. PLoS Genet 6, e1001111 (2010).
Zangen, D., Kaufman, Y., Zeligson, S., Perlberg, S., Fridman, H., Kanaan, M. et al. XX ovarian dysgenesis is caused by a PSMC3IP/HOP2 mutation that abolishes coactivation of estrogen-driven transcription. Am. J. Hum. Genet. 89, 572–579 (2011).
Pierson, T. M., Adams, D., Bonn, F., Martinelli, P., Cherukuri, P. F., Teer, J. K. et al. Whole-exome sequencing identifies homozygous AFG3L2 mutations in a spastic ataxia-neuropathy syndrome linked to mitochondrial m-AAA proteases. PLoS Genet 7, e1002325 (2011).
Puente, X. S., Quesada, V., Osorio, F. G., Cabanillas, R., Cadinanos, J., Fraile, J. M. et al. Exome sequencing and functional analysis identifies BANF1 mutation as the cause of a hereditary progeroid syndrome. Am. J. Hum. Genet. 88, 650–656 (2011).
Wang, J. L., Yang, X., Xia, K., Hu, Z. M., Weng, L., Jin, X. et al. TGM6 identified as a novel causative gene of spinocerebellar ataxias using exome sequencing. Brain 133 (Part 12), 3510–3518 (2010).
Sun, Y., Almomani, R., Aten, E., Celli, J., van der Heijden, J., Venselaar, H. et al. Terminal osseous dysplasia is caused by a single recurrent mutation in the FLNA gene. Am. J. Hum. Genet. 87, 146–153 (2010).
Robinson, P. N., Krawitz, P. & Mundlos, S. Strategies for exome and genome sequence data analysis in disease-gene discovery projects. Clin. Genet. 80, 127–132 (2011).
Sirmaci, A., Walsh, T., Akay, H., Spiliopoulos, M., Sakalar, Y. B., Hasanefendioglu-Bayrak, A. et al. MASP1 mutations in patients with facial, umbilical, coccygeal, and auditory findings of Carnevale, Malpuech, OSA, and Michels syndromes. Am. J. Hum. Genet. 87, 679–686 (2010).
Depienne, C., Bouteiller, D., Meneret, A., Billot, S., Groppa, S., Klebe, S. et al. RAD51 haploinsufficiency causes congenital mirror movements in humans. Am. J. Hum. Genet. 90, 301–307 (2012).
Rademakers, R., Baker, M., Nicholson, A. M., Rutherford, N. J., Finch, N., Soto-Ortolaza, A. et al. Mutations in the colony stimulating factor 1 receptor (CSF1R) gene cause hereditary diffuse leukoencephalopathy with spheroids. Nat. Genet. 44, 200–205 (2011).
Abou Jamra, R., Philippe, O., Raas-Rothschild, A., Eck, S. H., Graf, E., Buchert, R. et al. Adaptor protein complex 4 deficiency causes severe autosomal-recessive intellectual disability, progressive spastic paraplegia, shy character, and short stature. Am. J. Hum. Genet. 88, 788–795 (2011).
Tsurusaki, Y., Kosho, T., Hatasaki, K., Narumi, Y., Wakui, K., Fukushima, Y. et al. Exome sequencing in a family with an X-linked lethal malformationsyndrome: clinical consequences of hemizygous truncating OFD1 mutations in male patients. Clin. Genet doi:10.1111/j.1399-0004.2012.01885.x (2012).
Gilissen, C., Hoischen, A., Brunner, H. G. & Veltman, J. A. Disease gene identification strategies for exome sequencing. Eur. J. Hum. Genet. 20, 490–497 (2011).
Hoischen, A., van Bon, B. W., Gilissen, C., Arts, P., van Lier, B., Steehouwer, M. et al. De novo mutations of SETBP1 cause Schinzel-Giedion syndrome. Nat. Genet. 42, 483–485 (2010).
Saitsu, H., Osaka, H., Sasaki, M., Takanashi, J., Hamada, K., Yamashita, A. et al. Mutations in POLR3A and POLR3B encoding RNA Polymerase III subunits cause an autosomal-recessive hypomyelinating leukoencephalopathy. Am. J. Hum. Genet. 89, 644–651 (2011).
Gilissen, C., Arts, H. H., Hoischen, A., Spruijt, L., Mans, D. A., Arts, P. et al. Exome sequencing identifies WDR35 variants involved in Sensenbrenner syndrome. Am. J. Hum. Genet. 87, 418–423 (2010).
Roach, J. C., Glusman, G., Smit, A. F., Huff, C. D., Hubley, R., Shannon, P. T. et al. Analysis of genetic inheritance in a family quartet by whole-genome sequencing. Science 328, 636–639 (2010).
Lynch, M. Rate molecular spectrum, and consequences of human mutation. Proc. Natl Acad. Sci. USA 107, 961–968 (2010).
Gilissen, C., Hoischen, A., Brunner, H. G. & Veltman, J. A. Unlocking Mendelian disease using exome sequencing. Genome Biol. 12, 228 (2011).
O'Roak, B. J., Deriziotis, P., Lee, C., Vives, L., Schwartz, J. J., Girirajan, S. et al. Exome sequencing in sporadic autism spectrum disorders identifies severe de novo mutations. Nat. Genet. 43, 585–589 (2011).
Sanders, S. J., Murtha, M. T., Gupta, A. R., Murdoch, J. D., Raubeson, M. J., Willsey, A. J. et al. De novo mutations revealed by whole-exome sequencing are strongly associated with autism. Nature 485, 237–241 (2012).
Iossifov, I., Ronemus, M., Levy, D., Wang, Z., Hakker, I., Rosenbaum, J. et al. De novo gene disruptions in children on the autistic spectrum. Neuron 74, 285–299 (2012).
Koboldt, D. C., Ding, L., Mardis, E. R. & Wilson, R. K. Challenges of sequencing human genomes. Brief Bioinform. 11, 484–498 (2010).
Eid, J., Fehr, A., Gray, J., Luong, K., Lyle, J., Otto, G. et al. Real-time DNA sequencing from single polymerase molecules. Science 323, 133–138 (2009).
Branton, D., Deamer, D. W., Marziali, A., Bayley, H., Benner, S. A., Butler, T. et al. The potential and challenges of nanopore sequencing. Nat. Biotechnol. 26, 1146–1153 (2008).
Watson, J. D. & Crick, F. H. Molecular structure of nucleic acids; a structure for deoxyribose nucleic acid. Nature 171, 737–738 (1953).
Botstein, D., White, R. L., Skolnick, M. & Davis, R. W. Construction of a genetic linkage map in man using restriction fragment length polymorphisms. Am. J. Hum. Genet. 32, 314–331 (1980).
Mullis, K., Faloona, F., Scharf, S., Saiki, R., Horn, G. & Erlich, H. Specific enzymatic amplification of DNA in vitro: the polymerase chain reaction. Cold Spring Harb. Symp. Quant. Biol. 51, 263–273 (1986).
Smith, L. M., Sanders, J. Z., Kaiser, R. J., Hughes, P., Dodd, C., Connell, C. R. et al. Fluorescence detection in automated DNA sequence analysis. Nature 321, 674–679 (1986).
Royer-Pokora, B., Kunkel, L. M., Monaco, A. P., Goff, S. C., Newburger, P. E., Baehner, R. L. et al. Cloning the gene for an inherited human disorder—chronic granulomatous disease—on the basis of its chromosomal location. Nature 322, 32–38 (1986).
Cohen, D., Chumakov, I. & Weissenbach, J. A first-generation physical map of the human genome. Nature 366, 698–701 (1993).
Fleischmann, R. D., Adams, M. D., White, O., Clayton, R. A., Kirkness, E. F., Kerlavage, A. R. et al. Whole-genome random sequencing and assembly of Haemophilus influenzae Rd. Science 269, 496–512 (1995).
Dunham, I., Shimizu, N., Roe, B. A., Chissoe, S., Hunt, A. R., Collins, J. E. et al. The DNA sequence of human chromosome 22. Nature 402, 489–495 (1999).
Adams, M. D., Celniker, S. E., Holt, R. A., Evans, C. A., Gocayne, J. D., Amanatides, P. G. et al. The genome sequence of Drosophila melanogaster. Science 287, 2185–2195 (2000).
Myers, E. W., Sutton, G. G., Smith, H. O., Adams, M. D. & Venter, J. C. On the sequencing and assembly of the human genome. Proc. Natl Acad. Sci. USA 99, 4145–4146 (2002).
Venter, J. C., Adams, M. D., Myers, E. W., Li, P. W., Mural, R. J., Sutton, G. G. et al. The sequence of the human genome. Science 291, 1304–1351 (2001).
Lander, E. S., Linton, L. M., Birren, B., Nusbaum, C., Zody, M. C., Baldwin, J. et al. Initial sequencing and analysis of the human genome. Nature 409, 860–921 (2001).
Margulies, M., Egholm, M., Altman, W. E., Attiya, S., Bader, J. S., Bemben, L. A. et al. Genome sequencing in microfabricated high-density picolitre reactors. Nature 437, 376–380 (2005).
Wheeler, D. A., Srinivasan, M., Egholm, M., Shen, Y., Chen, L., McGuire, A. et al. The complete genome of an individual by massively parallel DNA sequencing. Nature 452, 872–876 (2008).
Ng, S. B., Turner, E. H., Robertson, P. D., Flygare, S. D., Bigham, A. W., Lee, C. et al. Targeted capture and massively parallel sequencing of 12 human exomes. Nature 461, 272–276 (2009).
Bolze, A., Byun, M., McDonald, D., Morgan, N. V., Abhyankar, A., Premkumar, L. et al. Whole-exome-sequencing-based discovery of human FADD deficiency. Am. J. Hum. Genet. 87, 873–881 (2010).
Walsh, T., Shahin, H., Elkan-Miller, T., Lee, M. K., Thornton, A. M., Roeb, W. et al. Whole exome sequencing and homozygosity mapping identify mutation in the cell polarity protein GPSM2 as the cause of nonsyndromic hearing loss DFNB82. Am. J. Hum. Genet. 87, 90–94 (2010).
Haack, T. B., Danhauser, K., Haberberger, B., Hoser, J., Strecker, V., Boehm, D. et al. Exome sequencing identifies ACAD9 mutations as a cause of complex I deficiency. Nat. Genet. 42, 1131–1134 (2010).
Krawitz, P. M., Schweiger, M. R., Rodelsperger, C., Marcelis, C., Kolsch, U., Meisel, C. et al. Identity-by-descent filtering of exome sequence data identifies PIGV mutations in hyperphosphatasia mental retardation syndrome. Nat. Genet. 42, 827–829 (2010).
Bilguvar, K., Ozturk, A. K., Louvi, A., Kwan, K. Y., Choi, M., Tatli, B. et al. Whole-exome sequencing identifies recessive WDR62 mutations in severe brain malformations. Nature 467, 207–210 (2010).
Caliskan, M., Chong, J. X., Uricchio, L., Anderson, R., Chen, P., Sougnez, C. et al. Exome sequencing reveals a novel mutation for autosomal recessive non-syndromic mental retardation in the TECR gene on chromosome 19p13. Hum. Mol. Genet. 20, 1285–1289 (2011).
Zuchner, S., Dallman, J., Wen, R., Beecham, G., Naj, A., Farooq, A. et al. Whole-exome sequencing links a variant in DHDDS to retinitis pigmentosa. Am. J. Hum. Genet. 88, 201–206 (2011).
Becker, J., Semler, O., Gilissen, C., Li, Y., Bolz, H. J., Giunta, C. et al. Exome sequencing identifies truncating mutations in human SERPINF1 in autosomal-recessive osteogenesis imperfecta. Am. J. Hum. Genet. 88, 362–371 (2011).
Glazov, E. A., Zankl, A., Donskoi, M., Kenna, T. J., Thomas, G. P., Clark, G. R. et al. Whole-exome re-sequencing in a family quartet identifies POP1 mutations as the cause of a novel skeletal dysplasia. PLoS Genet. 7, e1002027 (2011).
Sloan, J. L., Johnston, J. J., Manoli, I., Chandler, R. J., Krause, C., Carrillo-Carrasco, N. et al. Exome sequencing identifies ACSF3 as a cause of combined malonic and methylmalonic aciduria. Nat. Genet. 43, 883–886 (2011).
O’Sullivan, J., Bitu, C. C., Daly, S. B., Urquhart, J. E., Barron, M. J., Bhaskar, S. S. et al. Whole-exome sequencing identifies FAM20A mutations as a cause of amelogenesis imperfecta and gingival hyperplasia syndrome. Am. J. Hum. Genet. 88, 616–620 (2011).
Vissers, L. E., Lausch, E., Unger, S., Campos-Xavier, A. B., Gilissen, C., Rossi, A. et al. Chondrodysplasia and abnormal joint development associated with mutations in IMPAD1, encoding the Golgi-resident nucleotide phosphatase, gPAPP. Am. J. Hum. Genet. 88, 608–615 (2011).
Gotz, A., Tyynismaa, H., Euro, L., Ellonen, P., Hyotylainen, T., Ojala, T. et al. Exome sequencing identifies mitochondrial alanyl-tRNA synthetase mutations in infantile mitochondrial cardiomyopathy. Am. J. Hum. Genet. 88, 635–642 (2011).
Tariq, M., Belmont, J. W., Lalani, S., Smolarek, T. & Ware, S. M. SHROOM3 is a novel candidate for heterotaxy identified by whole exome sequencing. Genome Biol. 12, R91 (2011).
Snape, K., Hanks, S., Ruark, E., Barros-Nunez, P., Elliott, A., Murray, A. et al. Mutations in CEP57 cause mosaic variegated aneuploidy syndrome. Nat. Genet. 43, 527–529 (2011).
Theis, J. L., Sharpe, K. M., Matsumoto, M. E., Chai, H. S., Nair, A. A., Theis, J. D. et al. Homozygosity mapping and exome sequencing reveal GATAD1 mutation in autosomal recessive dilated cardiomyopathy. Circ. Cardiovasc. Genet. 4, 585–594 (2011).
Takata, A., Kato, M., Nakamura, M., Yoshikawa, T., Kanba, S., Sano, A. et al. Exome sequencing identifies a novel missense variant in RRM2B associated with autosomal recessive progressive external ophthalmoplegia. Genome Biol. 12, R92 (2011).
Aldahmesh, M. A., Khan, A. O., Mohamed, J. Y., Alkuraya, H., Ahmed, H., Bobis, S. et al. Identification of ADAMTS18 as a gene mutated in Knobloch syndrome. J. Med. Genet. 48, 597–601 (2011).
Doi, H., Yoshida, K., Yasuda, T., Fukuda, M., Fukuda, Y., Morita, H. et al. Exome sequencing reveals a homozygous SYT14 mutation in adult-onset, autosomal-recessive spinocerebellar ataxia with psychomotor retardation. Am. J. Hum. Genet. 89, 320–327 (2011).
Shaheen, R., Faqeih, E., Sunker, A., Morsy, H., Al-Sheddi, T., Shamseldin, H. E. et al. Recessive mutations in DOCK6, encoding the guanidine nucleotide exchange factor DOCK6, lead to abnormal actin cytoskeleton organization and Adams-Oliver syndrome. Am. J. Hum. Genet. 89, 328–333 (2011).
Sanna-Cherchi, S., Burgess, K. E., Nees, S. N., Caridi, G., Weng, P. L., Dagnino, M. et al. Exome sequencing identified MYO1E and NEIL1 as candidate genes for human autosomal recessive steroid-resistant nephrotic syndrome. Kidney Int. 80, 389–396 (2011).
Barak, T., Kwan, K. Y., Louvi, A., Demirbilek, V., Saygi, S., Tuysuz, B. et al. Recessive LAMC3 mutations cause malformations of occipital cortical development. Nat. Genet. 43, 590–594 (2011).
Galmiche, L., Serre, V., Beinat, M., Assouline, Z., Lebre, A. S., Chretien, D. et al. Exome sequencing identifies MRPL3 mutation in mitochondrial cardiomyopathy. Hum. Mutat. 32, 1225–1231 (2011).
Ozgul, R. K., Siemiatkowska, A. M., Yucel, D., Myers, C. A., Collin, R. W., Zonneveld, M. N. et al. Exome sequencing and cis-regulatory mapping identify mutations in MAK, a gene encoding a regulator of ciliary length, as a cause of retinitis pigmentosa. Am. J. Hum. Genet. 89, 253–264 (2011).
Hanson, D., Murray, P. G., O’Sullivan, J., Urquhart, J., Daly, S., Bhaskar, S. S. et al. Exome sequencing identifies CCDC8 mutations in 3-M syndrome, suggesting that CCDC8 contributes in a pathway with CUL7 and OBSL1 to control human growth. Am. J. Hum. Genet. 89, 148–153 (2011).
Kalay, E., Yigit, G., Aslan, Y., Brown, K. E., Pohl, E., Bicknell, L. S. et al. CEP152 is a genome maintenance protein disrupted in Seckel syndrome. Nat. Genet. 43, 23–26 (2010).
Bjursell, M. K., Blom, H. J., Cayuela, J. A., Engvall, M. L., Lesko, N., Balasubramaniam, S. et al. Adenosine kinase deficiency disrupts the methionine cycle and causes hypermethioninemia, encephalopathy, and abnormal liver function. Am. J. Hum. Genet. 89, 507–515 (2011).
Bredrup, C., Saunier, S., Oud, M. M., Fiskerstrand, T., Hoischen, A., Brackman, D. et al. Ciliopathies with skeletal anomalies and renal insufficiency due to mutations in the IFT-A gene WDR19. Am. J. Hum. Genet. 89, 634–643 (2011).
Aldahmesh, M. A., Mohamed, J. Y., Alkuraya, H. S., Verma, I. C., Puri, R. D., Alaiya, A. A. et al. Recessive mutations in ELOVL4 cause ichthyosis, intellectual disability, and spastic quadriplegia. Am. J. Hum. Genet. 89, 745–750 (2011).
Dauber, A., Nguyen, T. T., Sochett, E., Cole, D. E., Horst, R., Abrams, S. A. et al. Genetic defect in CYP24A1, the vitamin D 24-hydroxylase gene, in a patient with severe infantile hypercalcemia. J. Clin. Endocrinol. Metab. 97, E268–E274 (2011).
Logan, C. V., Lucke, B., Pottinger, C., Abdelhamed, Z. A., Parry, D. A., Szymanska, K. et al. Mutations in MEGF10, a regulator of satellite cell myogenesis, cause early onset myopathy, areflexia, respiratory distress and dysphagia (EMARDD). Nat. Genet. 43, 1189–1192 (2011).
Albers, C. A., Cvejic, A., Favier, R., Bouwmans, E. E., Alessi, M. C., Bertone, P. et al. Exome sequencing identifies NBEAL2 as the causative gene for gray platelet syndrome. Nat. Genet. 43, 735–737 (2011).
de Greef, J. C., Wang, J., Balog, J., den Dunnen, J. T., Frants, R. R., Straasheijm, K. R. et al. Mutations in ZBTB24 are associated with immunodeficiency, centromeric instability, and facial anomalies syndrome type 2. Am. J. Hum. Genet. 88, 796–804 (2011).
Sergouniotis, P. I., Davidson, A. E., Mackay, D. S., Li, Z., Yang, X., Plagnol, V. et al. Recessive mutations in KCNJ13, encoding an inwardly rectifying potassium channel subunit, cause leber congenital amaurosis. Am. J. Hum. Genet. 89, 183–190 (2011).
Erlich, Y., Edvardson, S., Hodges, E., Zenvirt, S., Thekkat, P., Shaag, A. et al. Exome sequencing and disease-network analysis of a single family implicate a mutation in KIF1A in hereditary spastic paraparesis. Genome Res. 21, 658–664 (2011).
Clayton-Smith, J., O'Sullivan, J., Daly, S., Bhaskar, S., Day, R., Anderson, B. et al. Whole-exome-sequencing identifies mutations in histone acetyltransferase gene KAT6B in individuals with the Say-Barber-Biesecker variant of Ohdo syndrome. Am. J. Hum. Genet. 89, 675–681 (2011).
Chen, W. J., Lin, Y., Xiong, Z. Q., Wei, W., Ni, W., Tan, G. H. et al. Exome sequencing identifies truncating mutations in PRRT2 that cause paroxysmal kinesigenic dyskinesia. Nat. Genet. 43, 1252–1255 (2011).
Simpson, M. A., Irving, M. D., Asilmaz, E., Gray, M. J., Dafou, D., Elmslie, F. V. et al. Mutations in NOTCH2 cause Hajdu-Cheney syndrome, a disorder of severe and progressive bone loss. Nat. Genet. 43, 303–305 (2011).
Isidor, B., Lindenbaum, P., Pichon, O., Bezieau, S., Dina, C., Jacquemont, S. et al. Truncating mutations in the last exon of NOTCH2 cause a rare skeletal disorder with osteoporosis. Nat. Genet. 43, 306–308 (2011).
Hoischen, A., van Bon, B. W., Rodriguez-Santiago, B., Gilissen, C., Vissers, L. E., de Vries, P. et al. De novo nonsense mutations in ASXL1 cause Bohring–Opitz syndrome. Nat. Genet. 43, 729–731 (2011).
Boyden, E. D., Campos-Xavier, A. B., Kalamajski, S., Cameron, T. L., Suarez, P., Tanackovich, G. et al. Recurrent dominant mutations affecting two adjacent residues in the motor domain of the monomeric kinesin KIF22 result in skeletal dysplasia and joint laxity. Am. J. Hum. Genet. 89, 767–772 (2011).
Min, B. J., Kim, N., Chung, T., Kim, O. H., Nishimura, G., Chung, C. Y. et al. Whole-exome sequencing identifies mutations of KIF22 in spondyloepimetaphyseal dysplasia with joint laxity, leptodactylic type. Am. J. Hum. Genet. 89, 760–766 (2011).
Noskova, L., Stranecky, V., Hartmannova, H., Pristoupilova, A., Baresova, V., Ivanek, R. et al. Mutations in DNAJC5, encoding cysteine-string protein alpha, cause autosomal-dominant adult-onset neuronal ceroid lipofuscinosis. Am. J. Hum. Genet. 89, 241–252 (2011).
Sirmaci, A., Spiliopoulos, M., Brancati, F., Powell, E., Duman, D., Abrams, A. et al. Mutations in ANKRD11 cause KBG syndrome, characterized by intellectual disability, skeletal malformations, and macrodontia. Am. J. Hum. Genet. 89, 289–294 (2011).
Dickinson, R. E., Griffin, H., Bigley, V., Reynard, L. N., Hussain, R., Haniffa, M. et al. Exome sequencing identifies GATA-2 mutation as the cause of dendritic cell, monocyte, B and NK lymphoid deficiency. Blood 118, 2656–2658 (2011).
Zimprich, A., Benet-Pages, A., Struhal, W., Graf, E., Eck, S. H., Offman, M. N. et al. A mutation in VPS35, encoding a subunit of the retromer complex, causes late-onset Parkinson disease. Am. J. Hum. Genet. 89, 168–175 (2011).
Vilarino-Guell, C., Wider, C., Ross, O. A., Dachsel, J. C., Kachergus, J. M., Lincoln, S. J. et al. VPS35 mutations in Parkinson disease. Am. J. Hum. Genet. 89, 162–167 (2011).
Klein, C. J., Botuyan, M. V., Wu, Y., Ward, C. J., Nicholson, G. A., Hammans, S. et al. Mutations in DNMT1 cause hereditary sensory neuropathy with dementia and hearing loss. Nat. Genet. 43, 595–600 (2011).
Norton, N., Li, D., Rieder, M. J., Siegfried, J. D., Rampersaud, E., Zuchner, S. et al. Genome-wide studies of copy number variation and exome sequencing identify rare variants in BAG3 as a cause of dilated cardiomyopathy. Am. J. Hum. Genet. 88, 273–282 (2011).
Shi, Y., Li, Y., Zhang, D., Zhang, H., Lu, F., Liu, X. et al. Exome sequencing identifies ZNF644 mutations in high myopia. PLoS Genet. 7, e1002084 (2011).
Bowne, S. J., Humphries, M. M., Sullivan, L. S., Kenna, P. F., Tam, L. C., Kiang, A. S. et al. A dominant mutation in RPE65 identified by whole-exome sequencing causes retinitis pigmentosa with choroidal involvement. Eur. J. Hum. Genet. 19, 1074–1081 (2011).
Weedon, M. N., Hastings, R., Caswell, R., Xie, W., Paszkiewicz, K., Antoniadi, T. et al. Exome sequencing identifies a DYNC1H1 mutation in a large pedigree with dominant axonal Charcot-Marie-Tooth disease. Am. J. Hum. Genet. 89, 308–312 (2011).
Zhou, C., Zang, D., Jin, Y., Wu, H., Liu, Z., Du, J. et al. Mutation in ribosomal protein L21 underlies hereditary hypotrichosis simplex. Hum. Mutat. 32, 710–714 (2011).
Le Goff, C., Mahaut, C., Wang, L. W., Allali, S., Abhyankar, A., Jensen, S. et al. Mutations in the TGFbeta binding-protein-like domain 5 of FBN1 are responsible for acromicric and geleophysic dysplasias. Am. J. Hum. Genet. 89, 7–14 (2011).
Le Goff, C., Mahaut, C., Abhyankar, A., Le Goff, W., Serre, V., Afenjar, A. et al. Mutations at a single codon in Mad homology 2 domain of SMAD4 cause Myhre syndrome. Nat. Genet. 44, 85–88 (2011).
Tsurusaki, Y., Osaka, H., Hamanoue, H., Shimbo, H., Tsuji, M., Doi, H. et al. Rapid detection of a mutation causing X-linked leucoencephalopathy by exome sequencing. J. Med. Genet. 48, 606–609 (2011).
Shamseldin, H. E., Faden, M. A., Alashram, W. & Alkuraya, F. S. Identification of a novel DLX5 mutation in a family with autosomal recessive split hand and foot malformation. J. Med. Genet. 49, 16–20 (2012).
Khan, K., Logan, C. V., McKibbin, M., Sheridan, E., Elcioglu, N. H., Yenice, O. et al. Next generation sequencing identifies mutations in Atonal homolog 7 (ATOH7) in families with global eye developmental defects. Hum. Mol. Genet. 21, 776–783 (2012).
Zhang, Z., Xia, W., He, J., Ke, Y., Yue, H., Wang, C. et al. Exome sequencing identifies SLCO2A1 mutations as a cause of primary hypertrophic osteoarthropathy. Am. J. Hum. Genet. 90, 125–132 (2012).
Mitchell, K., O'Sullivan, J., Missero, C., Blair, E., Richardson, R., Anderson, B. et al. Exome sequence identifies RIPK4 as the bartsocas—Papas syndrome locus. Am. J. Hum. Genet. 90, 69–75 (2012).
Walne, A., Dokal, A., Plagnol, V., Beswick, R., Kirwan, M., de la Fuente, J. et al. Exome sequencing identifies MPL as a causative gene in familial aplastic anemia. Haematologica 97, 524–528 (2012).
Cabral, R. M., Kurban, M., Wajid, M., Shimomura, Y., Petukhova, L. & Christiano, A. M. Whole-exome sequencing in a single proband reveals a mutation in the CHST8 gene in autosomal recessive peeling skin syndrome. Genomics 99, 202–208 (2012).
Mayr, J. A., Haack, T. B., Graf, E., Zimmermann, F. A., Wieland, T., Haberberger, B. et al. Lack of the mitochondrial protein acylglycerol kinase causes Sengers syndrome. Am. J. Hum. Genet. 90, 314–320 (2012).
Boyden, L. M., Choi, M., Choate, K. A., Nelson-Williams, C. J., Farhi, A., Toka, H. R. et al. Mutations in kelch-like 3 and cullin 3 cause hypertension and electrolyte abnormalities. Nature 482, 98–102 (2012).
Gibson, W. T., Hood, R. L., Zhan, S. H., Bulman, D. E., Fejes, A. P., Moore, R. et al. Mutations in EZH2 cause Weaver syndrome. Am. J. Hum. Genet. 90, 110–118 (2012).
Simpson, M. A., Deshpande, C., Dafou, D., Vissers, L. E., Woollard, W. J., Holder, S. E. et al. De Novo mutations of the gene encoding the histone acetyltransferase KAT6B cause genitopatellar syndrome. Am. J. Hum. Genet. 90, 290–294 (2012).
Campeau, P. M., Kim, J. C., Lu, J. T., Schwartzentruber, J. A., Abdul-Rahman, O. A., Schlaubitz, S. et al. Mutations in KAT6B, encoding a histone acetyltransferase, cause genitopatellar syndrome. Am. J. Hum. Genet. 90, 282–289 (2012).
Bochukova, E., Schoenmakers, N., Agostini, M., Schoenmakers, E., Rajanayagam, O., Keogh, J. M. et al. A mutation in the thyroid hormone receptor alpha gene. N. Engl. J. Med. 366, 243–249 (2012).
Hood, R. L., Lines, M. A., Nikkel, S. M., Schwartzentruber, J., Beaulieu, C., Nowaczyk, M. J. et al. Mutations in SRCAP, Encoding SNF2-Related CREBBP Activator Protein, Cause Floating-Harbor Syndrome. Am. J. Hum. Genet. 90, 308–313 (2012).
Montenegro, G., Rebelo, A. P., Connell, J., Allison, R., Babalini, C., D’Aloia, M. et al. Mutations in the ER-shaping protein reticulon 2 cause the axon-degenerative disorder hereditary spastic paraplegia type 12. J. Clin. Invest. 122, 538–544 (2012).
Ostergaard, P., Simpson, M. A., Mendola, A., Vasudevan, P., Connell, F. C., van Impel, A. et al. Mutations in KIF11 cause autosomal-dominant microcephaly variably associated with congenital lymphedema and chorioretinopathy. Am. J. Hum. Genet. 90, 356–362 (2012).
Jones, M. A., Ng, B. G., Bhide, S., Chin, E., Rhodenizer, D., He, P. et al. DDOST mutations identified by whole-exome sequencing are implicated in congenital disorders of glycosylation. Am. J. Hum. Genet. 90, 363–368 (2012).
Lines, M. A., Huang, L., Schwartzentruber, J., Douglas, S. L., Lynch, D. C., Beaulieu, C. et al. Haploinsufficiency of a spliceosomal GTPase encoded by EFTUD2 causes mandibulofacial dysostosis with microcephaly. Am. J. Hum. Genet. 90, 369–377 (2012).
Harms, M. B., Sommerville, R. B., Allred, P., Bell, S., Ma, D., Cooper, P. et al. Exome sequencing reveals DNAJB6 mutations in dominantly-inherited myopathy. Ann. Neurol. 71, 407–416 (2012).
Audo, I., Bujakowska, K., Orhan, E., Poloschek, C. M., Defoort-Dhellemmes, S., Drumare, I. et al. Whole-exome sequencing identifies mutations in GPR179 leading to autosomal-recessive complete congenital stationary night blindness. Am. J. Hum. Genet. 90, 321–330 (2012).
Hussain, M. S., Baig, S. M., Neumann, S., Nurnberg, G., Farooq, M., Ahmad, I. et al. A truncating mutation of CEP135 causes primary microcephaly and disturbed centrosomal function. Am. J. Hum. Genet. 90, 871–878 (2012).
Kirwan, M., Walne, A. J., Plagnol, V., Velangi, M., Ho, A., Hossain, U. et al. Exome sequencing identifies autosomal-dominant SRP72 mutations associated with familial aplasia and myelodysplasia. Am. J. Hum. Genet. 90, 888–892 (2012).
Lee, H., Graham, J. M., Rimoin, D. L., Lachman, R. S., Krejci, P., Tompson, S. W. et al. Exome sequencing identifies PDE4D mutations in acrodysostosis. Am. J. Hum. Genet. 90, 746–751 (2012).
Lin, Z., Chen, Q., Lee, M., Cao, X., Zhang, J., Ma, D. et al. Exome sequencing reveals mutations in TRPV3 as a cause of Olmsted syndrome. Am. J. Hum. Genet. 90, 558–564 (2012).
Fiskerstrand, T., Arshad, N., Haukanes, B. I., Tronstad, R. R., Pham, K. D., Johansson, S. et al. Familial diarrhea syndrome caused by an activating GUCY2C mutation. N. Engl. J. Med. 366, 1586–1595 (2012).
Bernier, F. P., Caluseriu, O., Ng, S., Schwartzentruber, J., Buckingham, K. J., Innes, A. M. et al. Haploinsufficiency of SF3B4, a component of the pre-mrna spliceosomal complex, causes Nager syndrome. Am. J. Hum. Genet. 90, 925–933 (2012).
Spiegel, R., Pines, O., Ta-Shma, A., Burak, E., Shaag, A., Halvardson, J. et al. Infantile cerebellar-retinal degeneration associated with a mutation in mitochondrial aconitase, ACO2. Am. J. Hum. Genet. 90, 518–523 (2012).
Zankl, A., Duncan, E. L., Leo, P. J., Clark, G. R., Glazov, E. A., Addor, M. C. et al. Multicentric carpotarsal osteolysis is caused by mutations clustering in the amino-terminal transcriptional activation domain of MAFB. Am. J. Hum. Genet. 90, 494–501 (2012).
Tsurusaki, Y., Okamoto, N., Ohashi, H., Kosho, T., Imai, Y., Hibi-Ko, Y. et al. Mutations affecting components of the SWI/SNF complex cause Coffin-Siris syndrome. Nat. Genet. 44, 376–378 (2012).
Srour, M., Schwartzentruber, J., Hamdan, F. F., Ospina, L. H., Patry, L., Labuda, D. et al. Mutations in C5ORF42 cause Joubert syndrome in the French Canadian population. Am. J. Hum. Genet. 90, 693–700 (2012).
Polvi, A., Linnankivi, T., Kivela, T., Herva, R., Keating, J. P., Makitie, O. et al. Mutations in CTC1, encoding the CTS telomere maintenance complex component 1, cause cerebroretinal microangiopathy with calcifications and cysts. Am. J. Hum. Genet. 90, 540–549 (2012).
Schossig, A., Wolf, N. I., Fischer, C., Fischer, M., Stocker, G., Pabinger, S. et al. Mutations in ROGDI cause Kohlschutter-Tonz syndrome. Am. J. Hum. Genet. 90, 701–707 (2012).
Santen, G. W., Aten, E., Sun, Y., Almomani, R., Gilissen, C., Nielsen, M. et al. Mutations in SWI/SNF chromatin remodeling complex gene ARID1B cause Coffin-Siris syndrome. Nat. Genet. 44, 379–380 (2012).
Nakazawa, Y., Sasaki, K., Mitsutake, N., Matsuse, M., Shimada, M., Nardo, T. et al. Mutations in UVSSA cause UV-sensitive syndrome and impair RNA polymerase IIo processing in transcription-coupled nucleotide-excision repair. Nat. Genet. 44, 586–592 (2012).
Austin, E. D., Ma, L., Leduc, C., Berman Rosenzweig, E., Borczuk, A., Phillips, J. A. et al. Whole exome sequencing to identify a novel gene (caveolin-1) associated with human pulmonary arterial hypertension. Circ. Cardiovasc. Genet. 44, 586–592 (2012).
Acknowledgements
We would like to thank the staff of Division of Human Genetics at National Institute of Genetics, Mishima, Shizuoka, Japan.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Rabbani, B., Mahdieh, N., Hosomichi, K. et al. Next-generation sequencing: impact of exome sequencing in characterizing Mendelian disorders. J Hum Genet 57, 621–632 (2012). https://doi.org/10.1038/jhg.2012.91
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/jhg.2012.91
Keywords
This article is cited by
-
Isolated proteinuria due to CUBN homozygous mutation – challenging the investigative paradigm
BMC Nephrology (2019)
-
Denoising of Aligned Genomic Data
Scientific Reports (2019)
-
Selective blockade of the lyso-PS lipase ABHD12 stimulates immune responses in vivo
Nature Chemical Biology (2018)
-
CXorf56, a dendritic neuronal protein, identified as a new candidate gene for X-linked intellectual disability
European Journal of Human Genetics (2018)
-
High-throughput mouse phenomics for characterizing mammalian gene function
Nature Reviews Genetics (2018)