Abstract
Purpose:
Treacher Collins/Franceschetti syndrome (TCS; OMIM 154500) is a disorder of craniofacial development belonging to the heterogeneous group of mandibulofacial dysostoses. TCS is classically characterized by bilateral mandibular and malar hypoplasia, downward-slanting palpebral fissures, and microtia. To date, three genes have been identified in TCS:,TCOF1, POLR1D, and POLR1C.
Methods:
We report a clinical and extensive molecular study, including TCOF1, POLR1D, POLR1C, and EFTUD2 genes, in a series of 146 patients with TCS. Phenotype–genotype correlations were investigated for 19 clinical features, between TCOF1 and POLR1D, and the type of mutation or its localization in the TCOF1 gene.
Results:
We identified 92/146 patients (63%) with a molecular anomaly within TCOF1, 9/146 (6%) within POLR1D, and none within POLR1C. Among the atypical negative patients (with intellectual disability and/or microcephaly), we identified four patients carrying a mutation in EFTUD2 and two patients with 5q32 deletion encompassing TCOF1 and CAMK2A in particular. Congenital cardiac defects occurred more frequently among patients with TCOF1 mutation (7/92, 8%) than reported in the literature.
Conclusion:
Even though TCOF1 and POLR1D were associated with extreme clinical variability, we found no phenotype–genotype correlation. In cases with a typical phenotype of TCS, 6/146 (4%) remained with an unidentified molecular defect.
Genet Med 18 1, 49–56.
Similar content being viewed by others
Introduction
Mandibulofacial dysostosis (MFD) is defined by abnormal craniofacial development, particularly of the first and second branchial arches. MFD is composed of a group of clinically and genetically heterogeneous disorders. Among MFD, Treacher Collins/Franceschetti syndrome (TCS; OMIM 154500) is the most frequent etiology, with an estimated prevalence of 1/50,000 births.1 TCS is clinically heterogeneous but characterized by typical bilateral facial features, such as malar and mandibular hypoplasia, downward-slanting palpebral fissures, coloboma of the lower lid, and microtia, and it often is associated with conductive hearing loss. Intellectual disability (ID) is rarely reported in the literature,2 and no visceral or skeletal malformations have been reported. TCS is also genetically heterogeneous; to date, three genes have been involved with a dominant (TCOF1,3 POLR1D)4 or a recessive (POLR1D,5 POLR1C)4 autosomal mode of inheritance. TCOF1 is the major gene involved and encodes the nucleolar phosphoprotein Treacle, acting in ribosomal RNA transcription by interacting with upstream binding factor and RNA polymerase I.6 TCOF1 also is involved in the proliferation and differentiation of neural crest cells in the first and second branchial arches during early embryogenesis.7 Heterozygous intragenic deletions in TCOF1 have been recently described as a rare cause of TCS.8,9 To date, no correlations have been established between phenotypic variability and the location of the mutations within TCOF1.2,10
Mutations and intragenic deletions in the POLR1D and POLR1C genes have been observed in a small subset of patients with TCS. POLR1D and POLR1C encode subunits of RNA polymerase I and III, respectively, which are involved in the synthesis of ribosomal RNA precursors and small RNAs.4
Moreover, mutations in EFTUD2 have been identified in patients with MFD and microcephaly or type Guion-Almeida (OMIM 610536),11 which is characterized by progressive and severe microcephaly, ID, and additional malformations such as choanal and aural atresia, cleft palate, congenital heart defects, and esophageal atresia.12 However, Gordon et al.12 and Luquetti et al.13 reported EFTUD2 mutations in patients without microcephaly, suggesting that EFTUD2 could be responsible for MFD and microcephaly (MFDM) as well as other forms of MFD.
In this study we performed molecular screening for mutations in the TCOF1, POLR1D, POLR1C, and EFTUD2 genes and investigated the clinical spectrum of 146 patients assessed for TCS in order to establish phenotype–genotype correlations.
Materials and Methods
Clinical data and DNA samples from a total of 146 patients referred for TCS were assessed for molecular investigation at Lariboisière Hospital, France. Each physician had to complete a form identifying 19 clinical features (details are provided in Table 1 and Supplementary Table S1 online). Based on the molecular identification of the disease, the patients were screened successively for TCOF1, POLR1D, POLR1C, and EFTUD2 using different strategies, namely, direct sequencing by Sanger methods, multiplex ligation-dependent probe amplification analysis, array–comparative genomic hybridization, and denaturing high-pressure liquid chromatography. A statistical analysis using the chi-square test was performed in an attempt to establish any relation between two qualitative variables.14 Written informed consent for genetic testing was obtained from all patients, and specific written informed consent for the publication of photos was obtained from the four patients shown in Figure 1 .
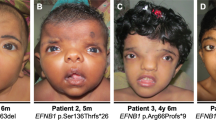
Front or lateral view of four patients from our series. (a) Patient with a heterozygous missense mutation (c.4009_4016delAGCAGCAG, p.Ser1337Glufs*59) in TCOF1 (patient 68; Supplementary Table S1 online). (b) Patient with an intragenic deletion of TCOF1 localized in the 3′ untranslated region (noncoding exon 26) (patient 92; Supplementary Table S1 online). (c) Patient with a missense mutation (c.259C>T, p.Arg87*) in POLR1D (patient 100; Supplementary Table S1 online). (d) Patient with complete deletion of POLR1D (patient 103; Supplementary Table S1 online).
For more details, see the Supplementary Materials and Methods online.
Results
The clinical and molecular data regarding TCOF1, POLR1D, and EFTUD2 genetic variations of the 146 patients from our series are reported in Supplementary Table S1 online.
Clinical data
The ratio of males to females for each gene was not significantly different (33% males for TCOF1, 50% for POLR1D, and 50% for EFTUD2; P = 0.81). The majority of patients were from France and its overseas departments, such as Guadeloupe and the Reunion Islands (76%), but we also analyzed samples from Belgium, Hungary, Lithuania, Morocco, Portugal, Spain, and Switzerland (24%).
As reported in the literature, we observed extreme intrafamilial variability in several families with either TCOF1 or POLR1D mutations. We were not surprised to identify an inherited mutation in TCOF1 in two of six patients referred as sporadic cases (patients 9 and 54; Supplementary Table S1 online) and an inherited mutation in POLR1D in the four others (patients 97, 98, 100, and 101; Supplementary Table S1 online).
All the patients with mutations in TCOF1, and for whom we collected enough clinical data, presented with downward-slanting palpebral fissures (70/70), and 99% presented with malar hypoplasia (70/71) ( Table 1 , Figures 1 and 2 ). Lower eyelid coloboma was present in 65% of the patients (43/66), cleft palate in 22% (14/64), and choanal atresia in 14% (8/56) compared with 50, 33, and 28%, respectively, in the study by Teber et al.2 ( Table 1 ). Furthermore, ID was present in only one patient, compared with 12% in the series reported by Teber et al. A cardiac malformation was observed in seven patients, namely, two cases of patent ductus arteriosus, four atrial septal defects, and one atrial septal defect combined with hypertrophic cardiomyopathy. In addition, one patient presented with nail hypoplasia. Mean severity score for patients with mutation in TCOF1 was 9.2 in the study by Teber et al. and 7.5 in this study (maximum score 17; mild ≤8, severe ≥9) (Supplementary Table S2 online; see details in Supplementary Materials and Methods online).
All patients with mutations in POLR1D presented with mandibular hypoplasia (7/7) and deafness (5/5), but none presented with choanal atresia, cardiac or renal malformations, anomaly of the limbs, or microcephaly ( Table 1 , Figure 2 ). In addition, no patients required intubation or tracheostoma during the neonatal period. Severe scoliosis was observed in one patient. Mean severity score was mild for both studies: 8.5 for Teber et al.2 and 7 in our investigation (maximum score 17; mild ≤8, severe ≥9).
Molecular studies
We identified 92 patients carrying a TCOF1 heterozygous molecular abnormality. Among the TCOF1 mutations, 47 were novel (Supplementary Table S1 online), and 85/92 (92%) of them were predicted to lead to a premature stop codon as a result of frameshift mutations (44 deletions, 9 duplications, 2 insertions, 3 insertions/deletions), nonsense substitutions (n = 18), and splice-site mutations (n = 9) (Supplementary Table S1 online). Moreover, three heterozygous missense substitutions were detected (Supplementary Table S1 online). In accordance with the findings of Teber et al.,2 we observed that the frameshift mutations were the most frequent mutation encountered (58/92, 63%; P < 0.0001). The common 5-bp deletion in exon 24 (p.Lys1457Glufs) occurred in 7/92 patients (8%) in our series (Supplementary Table S1 online). Moreover, we identified four heterozygous intragenic microdeletions in TCOF1, namely, a 60-kb deletion encompassing the promoter region, as well as exon 1 and 2, and three deletions encompassing exon 11, 24, and 25, respectively (Supplementary Table S1 online). In addition, two patients exhibited large deletions (262 kb and 1 Mb) detected by array–comparative genomic hybridization15 and encompassing several genes, including TCOF1 (Supplementary Table S1 online).
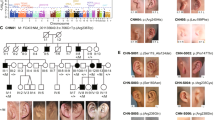
Among patients with a molecular abnormality in TCOF1, 27/92 (29%) were familial cases and 47/92 (51%) were referred as sporadic cases. De novo mutations were confirmed in 29/92 patients (32%) (Supplementary Table S1 online). In familial cases 18 patients inherited the mutation from their mother and six from their father, including one father presenting with a somatic mosaicism (patient 9; Supplementary Table S1 online and Figure 3 ), confirmed in DNA from blood, urine, and saliva (about 30% of mosaicism).
(a) Sequencing electropherograms and (b) denaturing high-pressure liquid chromatography graphs obtained by conventional genetic analysis using genomic DNA from circulating leukocytes. The patient affected by Treacher Collins/Franceschetti syndrome harbored the heterozygous deletion c.389_390delAG in TCOF1, and her father’s sequence was characterized by a somatic mosaicism (about 30%).
In an attempt to identify an association between the clinical features observed in the patients and a specific coding domain of the TCOF1 gene, we distinguished two different domains in TCOF1, namely, the LisH domain from exon 1 to 2 and the Treacle domain from exons 2 to 24. We identified only four mutations in the LisH domain; the majority of the mutations were localized in the Treacle domain (89/92, 97%; P < 0.0001) and, except for a hotspot in exon 24 (17/92, 18%), mutations were spread throughout the gene (P < 0.0001) (Supplementary Table S1 online).
Molecular analysis of POLR1D gene retrieved a molecular abnormality in nine patients, including seven mutations and two intragenic microdeletions. All seven mutations were located in exon 3, including two nonsense mutations, three missense mutations, and two deletions (Supplementary Table S1 online). The mutations were all heterozygous, except the homozygous c.163C>G (p.Leu55Val) mutation, which was inherited in a recessive manner.5 We also identified two large deletions of 65 and 362 kb encompassing a part of or the whole POLR1D gene, namely Chr13:g.(28163240 _28170998)_(28235492_28240863)del and Chr13:g.(27 995 500 _27 999747)_(28 362 012 _28 365086)del (NCBI build 37.3). We did not identify pathological variants in POLR1C in the 45 remaining patients with no mutations in TCOF1 and POLR1D. In conclusion, we found a positive result in TCOF1 or in POLR1D in 101/146 patients (69%) referred for molecular genetics analysis of TCS.
The EFTUD2 gene was analyzed in 11 patients with suspected MFDM or MFD Guion-Almeida type 1. Four patients in the cohort had a molecular abnormality in EFTUD2 (Supplementary Table S1 online): missense mutation (n = 1), nonsense mutation (n = 1), duplication (n = 1), deletion (n = 1). The nonsense mutation was previously reported in the same patient by Lines et al.11 (patient 105; Supplementary Table S1 online). A de novo origin of the mutation was confirmed for two patients.
Phenotype–genotype correlations
For each clinical feature (n = 19), we studied whether the proportion of one clinical sign in patients with a mutation in TCOF1 was different from the proportion of this clinical sign in patients with a mutation in POLR1D. We did not observe any significant association between any of the clinical features and the two genes studied (P values for all 19 clinical signs were >0.05; data not shown) ( Figure 1 ). In the same way, we did not find any significant association between the genotype and the degree of severity expressed as the score by either Teber et al.2 (i.e., 62% of patients with a TCOF1 mutation were coded as severe (50/81) compared with 57% for those with POLR1D mutations (4/7); P = 1) or in this study (i.e., 32% of patients with a TCOF1 mutation were coded as severe (26/81) compared with 29% for those with POLR1D mutations (2/7); P = 1). No specific association occurred when the TCS phenotype was scored as “typical” or “atypical” when the score could be determined (i.e., 89% of patients with a TCOF1 mutation were coded as typical (72/81) compared with 86% for those with POLR1D mutations (6/7); P = 0.58) (Supplementary Table S1 online).
In the same way, according to the type of mutation in TCOF1 (frameshift, nonsense, splice, missense, and intragenic microdeletion), the severity of the TCS phenotype was not significantly different, whatever the mutation (i.e., in those with typical versus atypical phenotypes, respectively, 63% (45/72) versus 78% (7/9) had frameshift mutations, 22% (16/72) versus 0% had nonsense mutations, 8% (6/72) versus 0% had splice mutations, 4% (3/72) versus 11% (1/9) had intragenic microdeletions, and 3% (2/72) versus 11% (1/9) had missense mutations; P = 1). There were no significant correlations for any of the clinical features of patients (P > 0.05 for each clinical feature; data not shown). We noticed that the three patients with missense mutations in TCOF1 had a mild phenotype for both scores, but this was not statistically significant.
Regarding TCOF1, the localization of the mutations in the LisH or Treacle domains had no effects on clinical features (P > 0.05 for each clinical feature; data not shown) or the degree of severity of TCS (P = 0.38 and P = 1). However, we observed a significantly smaller proportion of patients with severe scores using the scoring system described by Teber et al.2 (6/17, 59%) among the patients with a mutation in the exon 24 of TCOF1 compared with the other locations in the gene (P = 0.04). These data have not been confirmed using functional severity based on our own score(P = 0.12).
Among the seven patients with a cardiac defect, six had a frameshift mutation and one had a nonsense mutation (patients 2, 9, 32, 33, 38, 59, and 82; Supplementary Table S1 online). The mutations were located all along the TCOF1 gene.
We identified a large deletion encompassing TCOF1 in two patients presenting with typical facial features of TCS and ID.15 The sizes of the deletion were 262 kb and 1 Mb, and the minimal critical region also encompassed the CDX1, SLC6A7, CAMK2A, ARSI, and CD74 genes.
In addition, we identified 19 different exonic single-nucleotide polymorphism (SNPs) in TCOF1 (Supplementary Table S3 online); all were reported in the dbSNP database (http://www.ncbi.nlm.nih.gov/projects/SNP) at different frequencies. There was no difference between the frequencies of SNPs in patients and controls for all SNPs (P > 0.05; data not shown). We specially studied the intronic SNP c.-108-238C>T (rs28372960) in the promoter region, which was already described as a functional SNP leading to reduced TCOF1 expression.16 rs28372960 was found in six patients with a TCOF1 mutation, and we were not able to observe any significant correlation between the presence of SNP c.-108-238C>T (rs28372960) and the clinical findings (P > 0.05 for any clinical feature; data not shown) or the severity of the disease.
All patients carrying a mutation in EFTUD2 presented with microcephaly (4/4), ID (4/4), malar and mandibular hypoplasia (4/4), deafness (4/4), downward slanting palpebral fissures (3/3), and microtia (2/2) ( Figure 2 ). One patient presented with esophageal atresia and one with a complex cardiac malformation involving an atrial septal defect, patent ductus arteriosus, and a bicuspid aortic valve. None of them had coloboma of the lower lid, projection of scalp hair onto the lateral cheek, renal malformation, or anomaly of the limbs.
In total, among 146 total patients, we observed 94, 9, and 4 patients with a molecular defect in TCOF1, POLR1D, and EFTUD2, respectively. Among those 107 patients with identified molecular defect, 17 patients were atypical, including the two with a large deletion encompassing TCOF1 and the four with a mutation in EFTUD2. Among the negative patients, 33/39 (85%) were atypical.
Discussion
The aim of this study was to report the clinical and molecular data of a large cohort of 146 patients with TCS who were screened for TCOF1, POLR1D, and POLR1C. TCS is genetically and phenotypically heterogeneous, but to date no phenotype–genotype correlation has been established. In previous studies the diagnosis of TCS was usually established according to typical clinical facial features, namely, malar and mandibular hypoplasia, downward-slanting palpebral fissures, coloboma of the lower lid, and microtia without consideration of extrafacial features.17 Molecular diagnosis was not previously systematically performed because the only gene identified in 1996 was TCOF1, which causes a form of TCS inherited in an autosomal-dominant manner.3 Moreover, studies correlating clinical and molecular data from a series of patients with TCS are rare in the literature. In this study we analyzed clinical data, including additional features not previously described, and molecular data of the three genes known to be involved in the syndrome.
Clinical data
The most commonly occurring features in patients with TCS with mutations in TCOF1 in our series were downward-slanting palpebral fissures (100%) and malar hypoplasia (99%), as observed in a previous study.2 Conductive deafness was also a very frequent feature and was observed in 91% of the patients. Mandibular hypoplasia was absent in nine patients, who were probably affected with a very mild form that did not require further computed tomography or radiographic explorations. Indeed, the first and second branchial arches are affected in TCS, suggesting at least an impact on the mandible, varying from the absence of any sign to very severe micrognathia. Lower eyelid coloboma was less frequent but considered more specific to TCS syndrome (65%). Asymmetry was also quite frequent (53%), in contrast to a previous description of typical symmetrical facial features.17 We observed only one patient with ID and a mutation in TCOF1 (patient 62) in our series, compared with three patients described by Teber et al.2 When considering the incidence of ID in the general population, these data confirm that ID is not a feature of TCS. Functional issues secondary to mandibular hypoplasia and choanal atresia were evaluated by the need for a nasogastric tube or gastrostomy and for intubation or tracheostoma in the neonatal period ( Table 1 ). We observed in each situation that 20 and 28% of patients needed feeding or respiratory support, respectively, compared with 11% who needed respiratory support in the study by Teber et al.2 ( Table 1 ). Cleft palate was less frequent in our study (22%) compared with those by Teber et al.2 (33%) and Splendore et al.18 (28%) ( Table 1 ). Surprisingly, we observed a high rate of congenital cardiac defects in patients with mutations in TCOF1 (8%), which was never described in the literature. This suggests that cardiac malformation is not a rare feature of TCS. We recommend systematic screening echocardiograms for patients with TCS.
Patients with mutations in POLR1D had mild features and had no life-threatening complications. However, no statistical analysis was possible in this group because of the limited sample size. Larger series would be necessary to confirm these data.
Molecular data
TCOF1 was involved in 92/146 (63%) of all referrals in this study, but it was altered in 72/84 (86%) of patients with typical features. In the literature, the percentage of patients with mutations in TCOF1 ranged from 53% (97/182)8 to 93% (26/28).2,8,18 In the study by Bowman et al.,8 the proportion increased from 53 to 71% if patients with a high clinical suspicion of TCS are considered, and in the study by Teber et al.2 it increased from 61 to 78%.
TCOF1 has not been previously reported to contain mutational hotspots. Splendore et al.18,19 observed a clustering of pathogenic mutations in exons 10, 15, 16, 23, and 24. In this study we identified 47 novel mutations spread throughout the TCOF1 gene and only one clustering located in exons 23 and 24, which contained 27% of the mutations. The common 5-bp deletion in exon 24 (p.Lys1457Glufs) was present in 7% of patients in our series compared with 20% of patients in the study by Splendore et al.18 and none in the study by Teber et al.2 We hypothesize that the distinct geographic origin of the patients between studies explains such difference (Brazilian versus European centers), given that population in our study was quite diverse.
As described in the literature, the majority of TCOF1 mutations were frameshift mutations,8 yielding to haploinsufficiency of the Treacle protein.20,21,22
Bowman et al.8 also described five large deletions in TCOF1 in patients with typical TCS, although the full extent of each deletion was not known because they involved either the first or last exon. In this series we identified four intragenic microdeletions within TCOF1 in patients with typical TCS and two large deletions in 5q32 encompassing TCOF1 and neighboring genes in patients with TCS and ID.15 Even if we suggest distinguishing those two entities and set aside the large deletions, we recommend searching for all types of deletion by multiplex ligation-dependent probe amplification in cases of typical facial features of TCS, despite whether they are associated with ID.
Regarding the inheritance of the TCS for TCOF1 and POLR1D mutations, 55/101 (54%) of the patients from our series were referred as isolated cases. This value was in accordance with the study by Trainor et al.,23 in which 60% of cases were thought to arise as the result of a de novo mutation, based only on family history. However, we were able to confirm a de novo origin of the mutation in only 30/101 (30%) of the apparently sporadic patients. Concerning the remaining patients either DNA samples from parents were lacking for confirmation or inheritance from an asymptomatic parent was identified. This occurred in 6/55 (11%) of patients with TCS with a mutation in TCOF1 or POLR1D that were thought to be sporadic cases. These data suggest verifying systematically the inheritance of the mutation, even if the parents are asymptomatic. In addition, we described the second case of TCOF1 somatic mosaicism in an individual clinically unaffected with TCS. The first patient was previously described by Shoo et al.24 in an asymptomatic mother with a heterozygous mutation in leukocytes, hair root bulb, buccal mucosa, urine, and stool, but not in skin fibroblasts.
Mutations in POLR1D were inherited from an asymptomatic parent in 4/9 cases, compared with 2/92 for TCOF1. Moreover, mutations in POLR1C and the mutation p.Leu55Val in POLR1D are inherited in an autosomal-recessive manner.4,5 Consequently, genetic counseling is more difficult for sporadic cases, with the possibility that one of the “asymptomatic” parents is a carrier or that inheritance occurs in autosomal-recessive manner.
It is interesting to note that we did not identify any mutations in POLR1C, whereas Dauwerse et al.4 identified mutations in both alleles of POLR1C in 3/252 individuals with TCS (1%). To our knowledge, no other patient with POLR1C mutations has been reported in the literature. Examining whether POLR1C variants are carried by patients with particular geographic or ethnic origins would be interesting.
Phenotype–genotype correlations
As described in the literature, we observed in this study an extreme intrafamilial clinical variability.2,23,25 These new data also suggested that the prevalence of carriers of TCOF1 mutations in the population with a very mild phenotype was higher than estimated.
The degree of TCS severity was usually based on the overall appearance of the facial gestalt.10 Teber et al.2 developed another scoring system based on clinical facial features, resulting in an evaluation of the facial features severity of patients. We suggested evaluating the severity based on functional impairment and level of intervention required, namely intubation/tracheostoma, nasogastric tube/gastrostomy, choanal atresia, and conductive hearing loss (Supplementary Table S2 online; see Supplementary Materials and Methods online). We observed no correlation between the degree of severity and the gene involved or the type of mutation. Regarding the location of the mutation in the TCOF1 gene, we observed that patients with a mutation in exon 24 of TCOF1 had a lower severity score using the scoring system described by Teber et al. Because this result has not been confirmed with our score, we conclude that patients with a mutation in exon 24 have less severe facial features but do not have fewer functional consequences. We suggest further research with a larger cohort to confirm or invalidate these data.
Despite the high rate of SNPs in the TCOF1 gene,18 we did not observe any association between these polymorphic variants and the 19 clinical features considered. The functional SNP in the promoter region of TCOF116 does not seem to be associated with clinical severity according to the score calculated by Teber et al.2 and our score. These data suggest the implication of other additional factors.
In addition, Teber et al.2 demonstrated the absence of any correlation between the TCS phenotype and the location of the mutation within the TCOF1 gene or its biological consequence (missense versus premature termination), with the exception of conductive hearing loss, which occurred at a lower frequency in patients with mutations of the 3′ part of the open reading frame of TCOF1.2 Similarly, we did not observe any phenotype–genotype correlation between the type of gene involved (TCOF1/POLR1D), the type of mutation in TCOF1 (missense versus premature termination), and the location of the mutation within TCOF1, including mutations of the 3′ part of the open reading frame.
The small number of patients carrying POLR1D mutations precluded any statistical analysis. We noticed, however, that the nine patients described in this study had a typical and mild TCS phenotype. Moreover, in four familial cases parents were described as asymptomatic by the clinician, confirming the milder phenotype in patients with a mutation in POLR1D.
Negative patients
Among the 45 remaining patients with no mutation in TCOF1, POLR1D, and POLR1C, we observed 6 patients with typical TCS (13%) and 39 with atypical TCS (87%). Among the atypical patients, four (10%) had a mutation in EFTUD2, and two (5%) had a large 5q32 deletion encompassing TCOF1 ( Figure 4 ). These data confirmed that MFDM or MFD Guion-Almeida type 1 syndrome is a differential diagnosis for TCS in patients presenting with atypical features. Among the 146 patients of the cohort, 6 negative patients had typical TCS (4%), suggesting the implication of other genes.
(a) Mutation rate and distinction of typical and atypical patients. We identified a molecular defect in 73% of all patients assessed. Among the negative patients, 6/39 (15%) had a typical Treacher Collins/Franceschetti syndrome (TCS) phenotype and 33/39 (85%) had an atypical TCS phenotype, including microcephaly or intellectual disability. (b) Among typical patients, 78/84 (93%) were mutated in TCOF1 or POLR1D and 6/84 (7%) remained unsolved (equivalent to 4% of the cohort (6/146)). (c) Among atypical patients, 6/39 (15%) had a mutation in EFTUD2 or had a 5q32 deletion, and 33/39 (85%) remained unsolved.
In conclusion, we reported hitherto the largest series of patients with TCS with clinical and molecular data concerning TCOF1, POLR1D, and POLR1C. The molecular diagnosis rate for TCOF1 and POLR1D was 93% (78/84) in patients with typical TCS ( Figure 4 ). We concluded that a diagnosis of TCS should be discussed as soon as the common clinical diagnostic criteria are observed. However, some atypical clinical findings (congenital cardiac defects or facial asymmetry) do not exclude a diagnosis of TCS. We observed that congenital cardiac defects were not unusual in TCS and should be systematically evaluated. In addition, patients with a mutation in POLR1D seem to have a milder phenotype. We did not observe any phenotype–genotype correlations, confirming previous findings. We recommend molecular diagnosis for each patient with TCS and their parents to provide appropriate genetic counseling. Indeed, there is the possibility of a recessive form if POLR1D or POLR1C are involved, which leads to a 25% recurrence risk. In addition, we confirm the possible occurrence of somatic mosaicism regarding TCOF1 and nonpenetrance within asymptomatic parents regarding TCOF1 and POLR1D mutations, which leads to a recurrence risk up to 50%. Despite the identification of three genes responsible for TCS, we observed that 4% of patients in the cohort were affected with typical TCS without molecular abnormalities, suggesting the implication of additional genes.
Disclosure
The authors declare no conflict of interest.
References
Gorlin RJ, Cohen MM, Hennekam RCM . Syndromes of the Head and Neck, 4th edn. Oxford University Press: New York, 2001.
Teber OA, Gillessen-Kaesbach G, Fischer S, et al. Genotyping in 46 patients with tentative diagnosis of Treacher Collins syndrome revealed unexpected phenotypic variation. Eur J Hum Genet 2004;12:879–890.
Group TTCSC. Positional cloning of a gene involved in the pathogenesis of Treacher Collins syndrome. Nat Genet 1996;12:130–136.
Dauwerse JG, Dixon J, Seland S, et al. Mutations in genes encoding subunits of RNA polymerases I and III cause Treacher Collins syndrome. Nat Genet 2011;43:20–22.
Schaefer E, Collet C, Genevieve D, et al. Autosomal recessive POLR1D mutation with decrease of TCOF1 mRNA is responsible for Treacher Collins syndrome. Genet Med 2014;16:720–724.
Valdez BC, Henning D, So RB, Dixon J, Dixon MJ. The Treacher Collins syndrome (TCOF1) gene product is involved in ribosomal DNA gene transcription by interacting with upstream binding factor. Proc Natl Acad Sci USA 2004;101:10709–10714.
Dixon J, Jones NC, Sandell LL, et al. Tcof1/Treacle is required for neural crest cell formation and proliferation deficiencies that cause craniofacial abnormalities. Proc Natl Acad Sci USA 2006;103:13403–13408.
Bowman M, Oldridge M, Archer C, et al. Gross deletions in TCOF1 are a cause of Treacher-Collins-Franceschetti syndrome. Eur J Hum Genet 2012;20:769–777.
Beygo J, Buiting K, Seland S, et al. First Report of a Single Exon Deletion in TCOF1 Causing Treacher Collins Syndrome. Mol Syndromol 2012;2:53–59.
Edwards SJ, Gladwin AJ, Dixon MJ. The mutational spectrum in Treacher Collins syndrome reveals a predominance of mutations that create a premature-termination codon. Am J Hum Genet 1997;60:515–524.
Lines MA, Huang L, Schwartzentruber J, et al.; FORGE Canada Consortium. Haploinsufficiency of a spliceosomal GTPase encoded by EFTUD2 causes mandibulofacial dysostosis with microcephaly. Am J Hum Genet 2012;90:369–377.
Gordon CT, Petit F, Oufadem M, et al. EFTUD2 haploinsufficiency leads to syndromic oesophageal atresia. J Med Genet 2012;49:737–746.
Luquetti DV, Hing AV, Rieder MJ, et al. “Mandibulofacial dysostosis with microcephaly” caused by EFTUD2 mutations: expanding the phenotype. Am J Med Genet A 2013;161A:108–113.
Griffiths AJF, Miller JH, Suzuki DT, Lewontin RC, Gelbart WM. An Introduction to Genetic Analysis. WH Freeman: New York, 2000.
Vincent M, Collet C, Verloes A, et al. Large deletions encompassing the TCOF1 and CAMK2A genes are responsible for Treacher Collins syndrome with intellectual disability. Eur J Hum Genet 2014;22:52–56.
Masotti C, Armelin-Correa LM, Splendore A, et al. A functional SNP in the promoter region of TCOF1 is associated with reduced gene expression and YY1 DNA-protein interaction. Gene 2005;359:44–52.
Posnick JC . Treacher Collins syndrome: perspectives in evaluation and treatment. J Oral Maxillofac Surg 1997;55:1120–1133.
Splendore A, Silva EO, Alonso LG, et al. High mutation detection rate in TCOF1 among Treacher Collins syndrome patients reveals clustering of mutations and 16 novel pathogenic changes. Hum Mutat 2000;16:315–322.
Splendore A, Jabs EW, Passos-Bueno MR . Screening of TCOF1 in patients from different populations: confirmation of mutational hot spots and identification of a novel missense mutation that suggests an important functional domain in the protein treacle. J Med Genet 2002;39:493–495.
Isaac C, Marsh KL, Paznekas WA, et al. Characterization of the nucleolar gene product, treacle, in Treacher Collins syndrome. Mol Biol Cell 2000;11:3061–3071.
Gonzales B, Henning D, So RB, Dixon J, Dixon MJ, Valdez BC. The Treacher Collins syndrome (TCOF1) gene product is involved in pre-rRNA methylation. Hum Mol Genet 2005;14:2035–2043.
Masotti C, Ornelas CC, Splendore-Gordonos A, et al. Reduced transcription of TCOF1 in adult cells of Treacher Collins syndrome patients. BMC Med Genet 2009;10:136.
Trainor PA, Dixon J, Dixon MJ. Treacher Collins syndrome: etiology, pathogenesis and prevention. Eur J Hum Genet 2009;17:275–283.
Shoo BA, McPherson E, Jabs EW. Mosaicism of a TCOF1 mutation in an individual clinically unaffected with Treacher Collins syndrome. Am J Med Genet A 2004;126A:84–88.
Dixon J, Ellis I, Bottani A, Temple K, Dixon MJ. Identification of mutations in TCOF1: use of molecular analysis in the pre- and postnatal diagnosis of Treacher Collins syndrome. Am J Med Genet A 2004;127A:244–248.
Acknowledgements
The authors thank the patients and the family members for their support. Part of this work was supported by the French Franceschetti-Treacher Collins association Coline (http://netcoline.org), by the research programme “Programme Hospitalier de Recherche Clinique Régional” Languedoc-Roussillon, and by the “Plan National Maladies Rares 2011–2014” from the French “Direction Générale de l’Organisation des Soins” (DGOS).
Author information
Authors and Affiliations
Corresponding author
Supplementary information
Supplementary Table S1
(PDF 217 kb)
Supplementary Table S2
(PDF 58 kb)
Supplementary Table S3
(PDF 55 kb)
Supplementary Materials and Methods
(DOC 45 kb)
Rights and permissions
About this article
Cite this article
Vincent, M., Geneviève, D., Ostertag, A. et al. Treacher Collins syndrome: a clinical and molecular study based on a large series of patients. Genet Med 18, 49–56 (2016). https://doi.org/10.1038/gim.2015.29
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/gim.2015.29
Keywords
This article is cited by
-
Association between craniofacial anomalies, intellectual disability and autism spectrum disorder: Western Australian population-based study
Pediatric Research (2022)
-
TCOF1 pathogenic variants identified by Whole-exome sequencing in Chinese Treacher Collins syndrome families and hearing rehabilitation effect
Orphanet Journal of Rare Diseases (2019)
-
Rare ribosomopathies: insights into mechanisms of cancer
Nature Reviews Cancer (2019)