Abstract
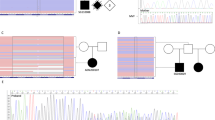
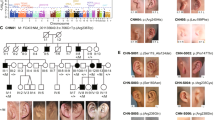
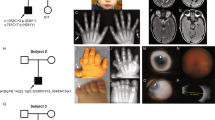
This study describes genomic findings among individuals with both orofacial clefts (OC) and microphthalmia/anophthalmia/coloboma (MAC) recorded in the Brazilian Database on Craniofacial Anomalies (BDCA). Chromosomal microarray analysis (CMA) and Whole Exome Sequencing (WES) were performed in 17 individuals with OC-MAC. Clinical interpretation of molecular findings was based on data available at the BDCA and on re-examination. No copy number variants (CNVs) classified as likely pathogenic or pathogenic were detected by CMA. WES allowed a conclusive diagnosis in six individuals (35.29%), two of them with variants in the CHD7 gene, and the others with variants in the TFAP2A, POMT1, PTPN11, and TP63 genes with the following syndromes: CHARGE, CHD7-spectrum, Branchiooculofacial, POMT1-spectrum, LEOPARD, and ADULT. Variants of uncertain significance (VUS) possibly associated to the phenotypes were found in six other individuals. Among the individuals with VUSes, three individuals presented variants in genes associated to defects of cilia structure and/or function, including DYNC2H1, KIAA0586, WDR34, INTU, RPGRIP1L, KIF7, and LMNA. These results show that WES was the most effective molecular approach for OC-MAC in this cohort. This study also reinforces the genetic heterogeneity of OC-MAC, and the importance of genes related to ciliopathies in this phenotype.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout

Similar content being viewed by others
Data availability
All data are available at www.bipmed.org/brave.
References
Mossey PA, Shaw WC, Munger RG, Murray JC, Murthy J, Little J. Global oral health inequalities: challenges in the prevention and management of orofacial clefts and potential solutions. Adv Dent Res. 2011;23:247–58. https://doi.org/10.1177/0022034511402083.
Saleem K, Zaib T, Sun W, Fu S. Assessment of candidate genes and genetic heterogeneity in human non syndromic orofacial clefts specifically non syndromic cleft lip with or without palate. Heliyon. 2019;5:e03019. https://doi.org/10.1016/j.heliyon.2019.e03019.
Plaisancié J, Ceroni F, Holt R, Zazo Seco C, Calvas P, Chassaing N, et al. Genetics of anophthalmia and microphthalmia. Part 1: non-syndromic anophthalmia/microphthalmia. Hum Genet. 2019;138:799–830. https://doi.org/10.1007/s00439-019-01977-y.
Calzolari E, Pierini A, Astolfi G, Bianchi F, Neville AJ, Rivieri F. Associated anomalies in multi-malformed infants with cleft lip and palate: an epidemiologic study of nearly 6 million births in 23 EUROCAT registries. Am J Med Genet A. 2007;143A:528–37. https://doi.org/10.1002/ajmg.a.31447.
Schraw JM, Benjamin RH, Scott DA, Brooks BP, Hufnagel RB, McLean SD, et al. A comprehensive assessment of co-occurring birth defects among infants with non-syndromic anophthalmia or microphthalmia. Ophthalmic Epidemiol. 2021;28:428–35. https://doi.org/10.1080/09286586.2020.1862244.
Shaw W. Global strategies to reduce the health care burden of craniofacial anomalies: report of WHO meetings on international collaborative research on craniofacial anomalies. Cleft Palate Craniofac J. 2004;41:238–43. https://doi.org/10.1597/03-214.1.
Roos L, Jensen H, Grønskov K, Holst R, Tümer Z. Congenital microphthalmia, anophthalmia and coloboma among live births in Denmark. Ophthalmic Epidemiol. 2016;23:324–30. https://doi.org/10.1080/09286586.2016.1213859.
Basha M, Demeer B, Revencu N, Helaers R, Theys S, Bou Saba S, et al. Whole exome sequencing identifies mutations in 10% of patients with familial non-syndromic cleft lip and/or palate in genes mutated in well-known syndromes. J Med Genet. 2018;55:449–58. https://doi.org/10.1136/jmedgenet-2017-105110.
Nasreddine G, El Hajj J, Ghassibe-Sabbagh M. Orofacial clefts embryology, classification, epidemiology, and genetics. Mutat Res Rev Mutat Res. 2021;787:108373 https://doi.org/10.1016/j.mrrev.2021.108373.
Harding P, Moosajee M. The molecular basis of human anophthalmia and microphthalmia. J Dev Biol. 2019;7:16 https://doi.org/10.3390/jdb7030016.
Lustosa-Mendes E, Santos APD, Vieira TP, Ribeiro EM, Rezende AA, Fett-Conte AC, et al. Identification of genomic imbalances in oral clefts. J Pediatr. 2021;97:321–8. https://doi.org/10.1016/j.jped.2020.06.005.
Rojnueangnit K, Mikhail FM, Cui X, Yu S, Robin NH. Predictor(s) of abnormal array comparative genomic hybridization results in patients with cleft lip and/or palate. Cleft Palate Craniofac J. 2015;52:724–31. https://doi.org/10.1597/14-088.
Balikova I, de Ravel T, Ayuso C, Thienpont B, Casteels I, Villaverde C, et al. High frequency of submicroscopic chro-mosomal deletions in patients with idiopathic congenital eye malformations. Am J Ophthalmol 2011;151:1087–94. https://doi.org/10.1016/j.ajo.2010.11.025.
Li J, Yang W, Wang YJ, Ma C, Curry CJ, McGoldrick D, et al. Exome sequencing identifies genetic variants in anophthalmia and microphthalmia. Am J Med Genet A. 2022;188:2376–88. https://doi.org/10.1002/ajmg.a.62874.
Volpe-Aquino RM, Monlleó IL, Lustosa-Mendes E, Mora AF, Fett-Conte AC, Félix TM, et al. CranFlow: an application for record-taking and management through the brazilian database on craniofacial anomalies. Birth Defects Res. 2018;110:72–80. https://doi.org/10.1002/bdr2.1123.
Gil-da-Silva-Lopes VL, Tacla MA, Sgardioli IC, Vieira TP, Monlleó IL. Brazil’s craniofacial project: different approaches on orofacial clefts and 22q11.2 deletion syndrome. Am J Med Genet C Semin Med Genet. 2020;184:912–27. https://doi.org/10.1002/ajmg.c.31852.
Riggs ER, Andersen EF, Cherry AM, Kantarci S, Kearney H, Patel A, et al. Technical standards for the interpretation and reporting of constitutional copy-number variants: a joint consensus recommendation of the American College of Medical Genetics and Genomics (ACMG) and the Clinical Genome Resource (ClinGen). Genet Med. 2020;22:245–57. https://doi.org/10.1038/s41436-019-0686-8.
Gonzales PR, Andersen EF, Brown TR, Horner VL, Horwitz J, Rehder CW, et al. Interpretation and reporting of large regions of homozygosity and suspected consanguinity/uniparental disomy, 2021 revision: a technical standard of the American College of Medical Genetics and Genomics (ACMG). Genet Med. 2022;24:255–61. https://doi.org/10.1016/j.gim.2021.10.004.
Highlander. Highlander. 2023. (https://sites.uclouvain.be/highlander)
Faria ACO, Caraciolo MP, Minillo RM, Almeida TF, Pereira SM, Cervato MC, et al. Varstation: a complete and efficient tool to support NGS data analysis. BioRxiv. 2019. https://doi.org/10.1101/833582.
Naslavsky MS, Yamamoto GL, de Almeida TF, Ezquina SAM, Sunaga DY, Pho N, et al. Exonic variants of an elderly cohort of Brazilians in the ABraOM database. Hum Mutat. 2017;38:751–63. https://doi.org/10.1002/humu.23220.
Richards S, Aziz N, Bale S, Bick D, Das S, Gastier-Foster J, et al. Standards and guidelines for the interpretation of sequence variants: a joint consensus recommendation of the American College of Medical Genetics and Genomics and the Association for Molecular Pathology. Genet Med. 2015;17:405–24. https://doi.org/10.1038/gim.2015.30.
Miller DT, Adam MP, Aradhya S, Biesecker LG, Brothman AR, Carter NP, et al. Consensus statement: chromosomal microarray is a first-tier clinical diagnostic test for individuals with developmental disabilities or congenital anomalies. Am J Hum Genet. 2010;86:749–64. https://doi.org/10.1016/j.ajhg.2010.04.006.
Hale CL, Niederriter AN, Green GE, Martin DM. Atypical phenotypes associated with pathogenic CHD7 variants and a proposal for broadening CHARGE syndrome clinical diagnostic criteria. Am J Med Genet A. 2016;170A:344–54. https://doi.org/10.1002/ajmg.a.37435.
Rinne T, Brunner HG, van Bokhoven H. p63-Associated Disorders. Cell Cycle. 2007;6:262–8. https://doi.org/10.4161/cc.6.3.3796.
Thanikachalam S, Hodapp E, Chang TC, Morel Swols D, Cengiz FB, Guo S, et al. Spectrum of genetic variants associated with anterior segment dysgenesis in South Florida. Genes. 2020;11:350 https://doi.org/10.3390/genes11040350.
Sarkozy A, Digilio MC, Dallapiccola B. Leopard syndrome. Orphanet J Rare Dis. 2008;3:13 https://doi.org/10.1186/1750-1172-3-13.
Vajsar J, Baskin B, Swoboda K, Biggar DW, Schachter H, Ray PN. Walker-warburg syndrome with POMT1 mutations can be associated with cleft lip and cleft palate. Neuromuscul Disord. 2008;18:675–7. https://doi.org/10.1016/j.nmd.2008.05.014.
Johnson B, Ouyang K, Frank L, Truty R, Rojahn S, Morales A, et al. Systematic use of phenotype evidence in clinical genetic testing reduces the frequency of variants of uncertain significance. Am J Med Genet A. 2022;188:2642–51. https://doi.org/10.1002/ajmg.a.62779.
Wawrocka A, Walczak-Sztulpa J, Pawlak M, Gotz-Wieckowska A, Krawczynski MR. Non-syndromic anophthalmia/microphthalmia can be caused by a PORCN variant inherited in X-linked recessive manner. Am J Med Genet A. 2021;185:250–5. https://doi.org/10.1002/ajmg.a.61938.
Jin SC, Homsy J, Zaidi S, Lu Q, Morton S, DePalma SR, et al. Contribution of rare inherited and de novo variants in 2,871 congenital heart disease probands. Nat Genet. 2017;49:1593–601. https://doi.org/10.1038/ng.3970.
Azevedo CMS, Machado RA, Martelli-Júnior H, Reis SRA, Persuhn DC, Coletta RD, et al. Exploring GRHL3 polymorphisms and SNP‐SNP interactions in the risk of non‐syndromic oral clefts in the Brazilian population. Oral Dis. 2019;26:145–51. https://doi.org/10.1111/odi.13204.
Dąbrowska J, Biedziak B, Szponar-Żurowska A, Budner M, Jagodziński PP, Płoski R, et al. Identification of novel susceptibility genes for non-syndromic cleft lip with or without cleft palate using NGS-based multigene panel testing. Mol Genet Genom. 2022;297:1315–27. https://doi.org/10.1007/s00438-022-01919-w.
Schmidts M, Arts HH, Bongers EMHF, Yap Z, Oud MM, Antony D, et al. Exome sequencing identifies DYNC2H1 mutations as a common cause of asphyxiating thoracic dystrophy (Jeune syndrome) without major polydactyly, renal or retinal involvement. J Med Genet. 2013a;50:309–23. https://doi.org/10.1136/jmedgenet-2012-101284.
Vilboux T, Doherty DA, Glass IA, Parisi MA, Phelps IG, Cullinane AR, et al. Molecular genetic findings and clinical correlations in 100 patients with Joubert syndrome and related disorders prospectively evaluated at a single center. Genet Med. 2017;19:875–82. https://doi.org/10.1038/gim.2016.204.
Schmidts M, Vodopiutz J, Christou-Savina S, Cortés Claudio R, McInerney-Leo AM, Emes RD, et al. Mutations in the Gene Encoding IFT Dynein Complex Component WDR34 Cause Jeune Asphyxiating Thoracic Dystrophy. Am J Hum Genet. 2013;93:932–44. https://doi.org/10.1016/j.ajhg.2013.10.003.
Toriyama M, Lee C, Taylor SP, Duran I, Cohn DH, Bruel AL, et al. The ciliopathy-associated CPLANE proteins direct basal body recruitment of intraflagellar transport machinery. Nat Genet. 2016;48:648–56. https://doi.org/10.1038/ng.3558.
Delous M, Baala L, Salomon R, Laclef C, Vierkotten J, Tory K, et al. The ciliary gene RPGRIP1L is mutated in cerebello-oculo-renal syndrome (Joubert syndrome type B) and Meckel syndrome. Nat Genet. 2007;39:875–81. https://doi.org/10.1038/ng2039.
Dafinger C, Liebau MC, Elsayed SM, Hellenbroich Y, Boltshauser E, Korenke GC, et al. Mutations in KIF7 link Joubert syndrome with Sonic Hedgehog signaling and microtubule dynamics. J Clin Investig. 2011;121:2662–7. https://doi.org/10.1172/JCI43639.
Doherty D, Parisi MA, Finn LS, Gunay-Aygun M, Al-Mateen M, Bates D, et al. Mutations in 3 genes (MKS3, CC2D2A and RPGRIP1L) cause COACH syndrome (Joubert syndrome with congenital hepatic fibrosis). J Med Genet. 2010;47:8–21. https://doi.org/10.1136/jmg.2009.067249.
Kortüm F, Chyrek M, Fuchs S, Albrecht B, Gillessen-Kaesbach G, Mütze U, et al. Hallermann-Streiff Syndrome: No Evidence for a Link to Laminopathies. Mol Syndromol. 2011;2:27–34. https://doi.org/10.1159/000334317.
Niceta M, Dentici ML, Ciolfi A, Marini R, Barresi S, Lepri FR, et al. Co-occurrence of mutations in KIF7 and KIAA0556 in Joubert syndrome with ocular coloboma, pituitary malformation and growth hormone deficiency: a case report and literature review. BMC Pediatr. 2020;20:120 https://doi.org/10.1186/s12887-020-2019-0.
de Araujo TK, Secolin R, Félix TM, de Souza LT, Fontes MÍB, Monlleó IL, et al. A multicentric association study between 39 genes and nonsyndromic cleft lip and palate in a Brazilian population. J Craniomaxillofac Surg. 2016;44:16–20. https://doi.org/10.1016/j.jcms.2015.07.026.
Putoux A, Thomas S, Coene KLM, Davis EE, Alanay Y, Ogur G, et al. KIF7 mutations cause fetal hydrolethalus and acrocallosal syndromes. Nat Genet. 2011;43:601–6. https://doi.org/10.1038/ng.826.
Cela P, Hampl M, Shylo NA, Christopher KJ, Kavkova M, Landova M, et al. Ciliopathy protein Tmem107 plays multiple roles in craniofacial development. J Dent Res. 2018;97:108–17. https://doi.org/10.1177/0022034517732538.
Morbidoni V, Agolini E, Slep KC, Pannone L, Zuccarello D, Cassina M, et al. Biallelic mutations in the TOGARAM1 gene cause a novel primary ciliopathy. J Med Genet. 2021;58:526–33. https://doi.org/10.1136/jmedgenet-2020-106833.
Sadler TW. Establishing the embryonic axes: prime time for teratogenic insults. J Cardiovasc Dev Dis. 2017;4:15 https://doi.org/10.3390/jcdd4030015.
Thiel C, Kessler K, Giessl A, Dimmler A, Shalev SA, von der Haar S, et al. NEK1 mutations cause short-rib polydactyly syndrome type majewski. Am J Hum Genet. 2011;88:106–14. https://doi.org/10.1016/j.ajhg.2010.12.004.
Rankin J, Auer-Grumbach M, Bagg W, Colclough K, Nguyen TD, Fenton-May J, et al. Extreme phenotypic diversity and nonpenetrance in families with the LMNA gene mutation R644C. Am J Med Genet A. 2008;146A:1530–42. https://doi.org/10.1002/ajmg.a.32331.
Acknowledgements
This study would not be possible without the dedication of the patients and families and our colleagues from BDCA participant centers.The authors would like to thank the following BDCA participant centers, which provided samples from patients included in this study: Centro de atenção aos defeitos da face (CADEFI), Faculdade de Medicina de São José do Rio Preto (FAMERP), Centrinho Prefeito Luiz Gomes-Joinville, Universidade Federal do Rio Grande do Norte (UFRN), and Hospital de Clínicas de Porto Alegre. We are also indebted to Gabriela Roldão Correia-Costa for her technical support.
Funding
This study was partially supported by the National Council for Scientific and Technological Development (CNPq) (#408504/2018-8 and #309782/2020-1), São Paulo Research Foundation (FAPESP #2018 / 21370-4) and Coordenação de Aperfeiçoamento de Pessoal de Nível Superior – Brasil (CAPES #001), and by the Fonds de la Recherche Scientifique - FNRS Grant J.0228.20 (to MV). The authors thank the Genomics Platform of University of Louvain for the access to the Next Generation Sequencing data analyses cluster. Eleonore Pairet is a Research Fellow (ASP) grantee of the Fonds de la Recherche Scientifique – FNRS. We also thank the National Lottery, Belgium and the Foundation Against Cancer (2010-101), Belgium for their support to the Genomics Platform of University of Louvain and de Duve Institute, as well as the Fonds de la Recherche Scientifique - FNRS Eguipment Grant U.N035.17 for the «Big data analysis cluster for NGS at UCLouvain».
Author information
Authors and Affiliations
Contributions
MAT performed clinical evaluation, clinical interpretation of molecular findings and wrote the manuscript; MMC performed laboratory tests, CMA and WES data analyses through Varstation and revised the manuscript; EP performed the WES data analyses through Highlander and variant evaluation, and wrote the manuscript; ILM collaborated with the design of the study and revised the manuscript; EMR performed clinical evaluation and revised the manuscript; ELM performed clinical evaluation and revised the manuscript; RH developed the data analysis program used for WES analysis; TPV collaborated with the design of the study, CMA analysis and revised the manuscript; MV supervised the WES studies, provided funding and revised the manuscript; VLGSL designed the study, performed clinical evaluation, provided funding, performed clinical interpretation of molecular findings, and revised the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethical approval
The study was conducted in accordance with the Declaration of Helsinki and approved by the Ethics Committee of University of Campinas (protocol numbers: 35316314.9.1001.5404 and 85020018.8.0000.5404). Informed consent was obtained from all subjects involved in the study. Written informed consent has been obtained from the patients to publish this paper.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Atique Tacla, M., de Mello Copelli, M., Pairet, E. et al. Molecular investigation in individuals with orofacial clefts and microphthalmia-anophthalmia-coloboma spectrum. Eur J Hum Genet (2023). https://doi.org/10.1038/s41431-023-01488-5
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41431-023-01488-5