Abstract
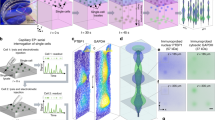
To measure cell-to-cell variation in protein-mediated functions, we developed an approach to conduct ∼103 concurrent single-cell western blots (scWesterns) in ∼4 h. A microscope slide supporting a 30-μm-thick photoactive polyacrylamide gel enables western blotting: settling of single cells into microwells, lysis in situ, gel electrophoresis, photoinitiated blotting to immobilize proteins and antibody probing. We applied this scWestern method to monitor single-cell differentiation of rat neural stem cells and responses to mitogen stimulation. The scWestern quantified target proteins even with off-target antibody binding, multiplexed to 11 protein targets per single cell with detection thresholds of <30,000 molecules, and supported analyses of low starting cell numbers (∼200) when integrated with FACS. The scWestern overcomes limitations of antibody fidelity and sensitivity in other single-cell protein analysis methods and constitutes a versatile tool for the study of complex cell populations at single-cell resolution.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Altschuler, S.J. & Wu, L.F. Cellular heterogeneity: do differences make a difference? Cell 141, 559–563 (2010).
Chang, H.H., Hemberg, M., Barahona, M., Ingber, D.E. & Huang, S. Transcriptome-wide noise controls lineage choice in mammalian progenitor cells. Nature 453, 544–547 (2008).
Raj, A., Rifkin, S.A., Andersen, E. & van Oudenaarden, A. Variability in gene expression underlies incomplete penetrance. Nature 463, 913–918 (2010).
Dalerba, P. et al. Single-cell dissection of transcriptional heterogeneity in human colon tumors. Nat. Biotechnol. 29, 1120–1127 (2011).
Balic, M., Williams, A., Lin, H., Datar, R. & Cote, R.J. Circulating tumor cells: from bench to bedside. Annu. Rev. Med. 64, 31–44 (2013).
Lazzara, M.J. et al. Impaired SHP2-mediated extracellular signal-regulated kinase activation contributes to gefitinib sensitivity of lung cancer cells with epidermal growth factor receptor-activating mutations. Cancer Res. 70, 3843–3850 (2010).
Bendall, S.C. et al. Single-cell mass cytometry of differential immune and drug responses across a human hematopoietic continuum. Science 332, 687–696 (2011).
Zong, C., Lu, S., Chapman, A.R. & Xie, X.S. Genome-wide detection of single-nucleotide and copy-number variations of a single human cell. Science 338, 1622–1626 (2012).
Taniguchi, Y. et al. Quantifying E. coli proteome and transcriptome with single-molecule sensitivity in single cells. Science 329, 533–538 (2010).
Schwanhäusser, B. et al. Global quantification of mammalian gene expression control. Nature 473, 337–342 (2011).
Newman, J.R.S. et al. Single-cell proteomic analysis of S. cerevisiae reveals the architecture of biological noise. Nature 441, 840–846 (2006).
Wei, W. et al. Microchip platforms for multiplex single-cell functional proteomics with applications to immunology and cancer research. Genome Med. 5, 75 (2013).
Ashton, R.S. et al. Astrocytes regulate adult hippocampal neurogenesis through ephrin-B signaling. Nat. Neurosci. 15, 1399–1406 (2012).
Sevecka, M. & MacBeath, G. State-based discovery: a multidimensional screen for small-molecule modulators of EGF signaling. Nat. Methods 3, 825–831 (2006).
Spurrier, B., Ramalingam, S. & Nishizuka, S. Reverse-phase protein lysate microarrays for cell signaling analysis. Nat. Protoc. 3, 1796–1808 (2008).
Stadler, C. et al. Immunofluorescence and fluorescent-protein tagging show high correlation for protein localization in mammalian cells. Nat. Methods 10, 315–323 (2013).
Maecker, H.T. & Trotter, J. Flow cytometry controls, instrument setup, and the determination of positivity. Cytometry A 69, 1037–1042 (2006).
Schulz, K.R., Danna, E.A., Krutzik, P.O. & Nolan, G.P. Single-cell phospho-protein analysis by flow cytometry. Curr. Protoc. Immunol. 96, 8.17 (2012).
Towbin, H., Staehelin, T. & Gordon, J. Electrophoretic transfer of proteins from polyacrylamide gels to nitrocellulose sheets: procedure and some applications. Proc. Natl. Acad. Sci. USA 76, 4350–4354 (1979).
Ciaccio, M.F., Wagner, J.P., Chuu, C.P., Lauffenburger, D.A. & Jones, R.B. Systems analysis of EGF receptor signaling dynamics with microwestern arrays. Nat. Methods 7, 148–155 (2010).
Hughes, A.J. & Herr, A.E. Microfluidic western blotting. Proc. Natl. Acad. Sci. USA 109, 21450–21455 (2012).
Shapiro, A.L., Viñuela, E. & Maizel, J.V. Jr. Molecular weight estimation of polypeptide chains by electrophoresis in SDS-polyacrylamide gels. Biochem. Biophys. Res. Commun. 28, 815–820 (1967).
Gallagher, S., Winston, S.E., Fuller, S.A. & Hurrell, J.G. Immunoblotting and immunodetection. Curr. Protoc. Immunol. 83, 8.10 (2008).
Friedman, N., Cai, L. & Xie, X.S. Linking stochastic dynamics to population distribution: an analytical framework of gene expression. Phys. Rev. Lett. 97, 168302 (2006).
Cai, L., Friedman, N. & Xie, X.S. Stochastic protein expression in individual cells at the single molecule level. Nature 440, 358–362 (2006).
Cohen, A.A. et al. Protein dynamics in individual human cells: experiment and theory. PLoS ONE 4, e4901 (2009).
Bendall, S.C., Nolan, G.P., Roederer, M. & Chattopadhyay, P.K. A deep profiler's guide to cytometry. Trends Immunol. 33, 323–332 (2012).
Gage, F.H. et al. Survival and differentiation of adult neuronal progenitor cells transplanted to the adult brain. Proc. Natl. Acad. Sci. USA 92, 11879–11883 (1995).
Peltier, J., O'Neill, A. & Schaffer, D.V. PI3K/Akt and CREB regulate adult neural hippocampal progenitor proliferation and differentiation. Dev. Neurobiol. 67, 1348–1361 (2007).
Jensen, K.J. et al. An ERK-p38 subnetwork coordinates host cell apoptosis and necrosis during coxsackievirus b3 infection. Cell Host Microbe 13, 67–76 (2013).
Huang, C.Y.F. & Ferrell, J.E. Ultrasensitivity in the mitogen-activated protein kinase cascade. Proc. Natl. Acad. Sci. USA 93, 10078–10083 (1996).
Eldar, A. & Elowitz, M.B. Functional roles for noise in genetic circuits. Nature 467, 167–173 (2010).
Chou, Y.H., Khuon, S., Herrmann, H. & Goldman, R.D. Nestin promotes the phosphorylation-dependent disassembly of vimentin intermediate filaments during mitosis. Mol. Biol. Cell 14, 1468–1478 (2003).
Hyder, C.L., Isoniemi, K.O., Torvaldson, E.S. & Eriksson, J.E. Insights into intermediate filament regulation from development to ageing. J. Cell Sci. 124, 1363–1372 (2011).
Sahlgren, C.M. et al. Mitotic reorganization of the intermediate filament protein nestin involves phosphorylation by cdc2 kinase. J. Biol. Chem. 276, 16456–16463 (2001).
Yang, H.Y., Lieska, N., Goldman, A.E. & Goldman, R.D. Colchicine-sensitive and colchicine-insensitive intermediate filament systems distinguished by a new intermediate filament-associated protein, IFAP-70/280kD. Cell Motil. Cytoskeleton 22, 185–199 (1992).
Su, P.H. et al. Identification and cytoprotective function of a novel nestin isoform, Nes-S, in dorsal root ganglia neurons. J. Biol. Chem. 288, 8391–8404 (2013).
Hendrickson, M.L., Rao, A.J., Demerdash, O.N.A. & Kalil, R.E. Expression of nestin by neural cells in the adult rat and human brain. PLoS ONE 6, e18535 (2011).
Marx, V. Finding the right antibody for the job. Nat. Methods 10, 703–707 (2013).
Fujioka, A. et al. Dynamics of the Ras/ERK MAPK cascade as monitored by fluorescent probes. J. Biol. Chem. 281, 8917–8926 (2006).
Yu, J.H. & Schaffer, D.V. Selection of novel vesicular stomatitis virus glycoprotein variants from a peptide insertion library for enhanced purification of retroviral and lentiviral vectors. J. Virol. 80, 3285–3292 (2006).
Peltier, J. & Schaffer, D.V. Viral packaging and transduction of adult hippocampal neural progenitors. Methods Mol. Biol. 621, 103–116 (2010).
Hughes, A.J., Lin, R.K.C., Peehl, D.M. & Herr, A.E. Microfluidic integration for automated targeted proteomic assays. Proc. Natl. Acad. Sci. USA 109, 5972–5977 (2012).
Hughes, A.J. & Herr, A.E. Quantitative enzyme activity determination with zeptomole sensitivity by microfluidic gradient-gel zymography. Anal. Chem. 82, 3803–3811 (2010).
Selinummi, J. et al. Bright field microscopy as an alternative to whole cell fluorescence in automated analysis of macrophage images. PLoS ONE 4, e7497 (2009).
Acknowledgements
We acknowledge E. Connelly for assistance with cell culture and conventional western blots, K. Heydari at the UC Berkeley Cancer Research Laboratory Flow Cytometry Facility for assistance with interfacing with FACS; R. Lin, L. Bugaj, E. Woods and C. Nilson for critical discussion; and the QB3 Biomolecular Nanotechnology Center (BNC) at UC Berkeley for partial infrastructure support. A.J.H. was funded as a 2013 Siebel Scholar. D.P.S. was funded as a 2014 Siebel Scholar and by a California Institute for Regenerative Medicine training grant (TG2-01164). A.E.H. an Alfred P. Sloan Research Fellow in chemistry. This work was supported by a US National Institutes of Health (NIH) New Innovator Award (DP2OD007294 to A.E.H.), a Medical Research Program Grant from the W.M. Keck Foundation (to A.E.H.), a UC Berkeley Bakar Fellowship (to A.E.H.) and an NIH research project grant (R01ES020903 to D.V.S.).
Author information
Authors and Affiliations
Contributions
A.J.H. and A.E.H. designed the scWestern. A.J.H., A.E.H., D.P.S. and D.V.S. designed experiments. A.J.H., Z.X. and C.-C.K. performed scWesterns and calibration experiments. D.P.S. performed NSC culture, stimulation and differentiation; conventional western blots; flow cytometry; and plate-based immunocytochemistry. A.J.H. and D.P.S. performed in-microwell ICC. A.J.H., D.P.S. and Z.X. performed confocal microscopy. Z.X. and A.J.H. designed software and performed analysis for cell/microwell scoring, immunoprobing quality control and fluorescence quantitation. Z.X. performed fluid dynamics simulations. Validation data were analyzed by A.J.H., D.P.S. and Z.X. All authors wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
A.J.H., Z.X., C.-C.K. and A.E.H. are inventors on pending patents related to scWestern blot methods. A.E.H. holds equity interest in Zephyrus Biosciences.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–27, Supplementary Table 1 and Supplementary Notes 1–8 (PDF 13747 kb)
Supplementary Software
Software and readme files for automated cell/microwell counting and scWestern blot analysis. (ZIP 26 kb)
EGFP+ NSC lysis, protein separation, and immobilization
Inverted fluorescence microscopy video at 10x magnification showing EGFP+ NSC lysis, protein separation upon application of electric field (dark moving bands), and UV immobilization steps in a scWestern gel. Black frames in the video correspond to a period of UV exposure of the scWestern gel after protein separation. Note retention of EGFP in the scWestern gel at the end of the UV exposure step. Vertical extent of the field of view is approximately 700 microns. (MP4 1616 kb)
Rights and permissions
About this article
Cite this article
Hughes, A., Spelke, D., Xu, Z. et al. Single-cell western blotting. Nat Methods 11, 749–755 (2014). https://doi.org/10.1038/nmeth.2992
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth.2992
This article is cited by
-
Decoding cancer insights: recent progress and strategies in proteomics for biomarker discovery
Journal of Proteins and Proteomics (2024)
-
Optimized data-independent acquisition approach for proteomic analysis at single-cell level
Clinical Proteomics (2022)
-
Cyclic microchip assay for measurement of hundreds of functional proteins in single neurons
Nature Communications (2022)