Abstract
Rats with fluorescent markers are of great value for studies that trace lineage-specific development, particularly those assessing the differentiation potential of embryonic stem cells (ESCs). The piggyBac (PB) transposon is widely used for the efficient introduction of genetic modifications into genomes, and has already been successfully used to produce transgenic mice and rats. Here, we generated transgenic rats carrying either the desRed fluorescent protein (RFP) gene or the enhanced green fluorescent protein (eGFP) gene by injecting pronuclei with PB plasmids. We showed that the transgenic rats expressed the RFP or eGFP gene in many organs and had the capability to transmit the marker gene to the next generation through germline integration. In addition, rat embryonic stem cells (ESCs) carrying an RFP reporter gene can be derived from the blastocysts of the transgenic rats. Moreover, the RFP gene can be detected in chimeras derived from RFP ESCs via blastocyst injection. This work suggests that PB-mediated transgenesis is a powerful tool to generate transgenic rats expressing fluorescent proteins with high efficiency, and this technique can be used to derive rat ESCs expressing a reporter protein.
Similar content being viewed by others
Introduction
PiggyBac (PB) is a DNA transposon element that was first isolated from Trichoplusia ni1,2 and has shown an efficient transposition ability in many species3,4,5. Compared with other transposons (e.g., sleeping beauty, Tol2 and MosI), PB can insert with higher transposition activity into mammalian genomes6, which makes it an ideal tool for research on the genetic traits of mammalian cells7,8,9. Therefore, PB has been extensively utilized in studies of genetic function and pathway regulation, such as the reprogramming of somatic cells into pluripotent stem cells10, the discovery of novel regulators that govern the pluripotency of stem cells11 and the generation of a mutation library for genetic screening12.
The rat is an important and useful laboratory species for modeling clinical diseases13. Although the rat genome has been sequenced (Rat Genome Sequencing Project Consortium 2004) and shared (www.ensembl.org/Rattus_norvegicus), little work has been performed to predict genetic functions compared with the mouse. To date, several gene manipulation strategies (such as N-ethyl-N-nitrosourea (ENU)14, zinc-finger nucleases (ZFNs)15 and transcription activator-like (TAL) effector nucleases (TALENs)16) have been used to produce genetically modified rats. Meanwhile, embryo manipulation techniques, including traditional pronuclear injection15, ESC germline transmission17 and haploid ESC intracytoplasmic injection18, have been developed for the production of transgenic rats. However, these methods are difficult to perform and are time-consuming. To date, compared with the thousands of types of transgenic mice, rats carrying fluorescent markers are still lacking in biological research, thus hindering the study of the developmental potential of rats during embryogenesis and differentiation.
Here, we have developed a simple and efficient strategy to generate transgenic rats that carry fluorescent marker genes. We microinjected a modified PB vector carrying either the desRed fluorescent protein (RFP) gene or the enhanced green fluorescent protein (eGFP) gene with a PBase plasmid into the pronuclei of rat zygotes. The RFP and eGFP genes were able to insert into many sites of the rat genome and were expressed in various organs. This stable insertion by the PB system was passed to the next generation through germline transmission. In summary, this study provided a novel method to produce fluorescent rats that can be used as a fluorescent ESC platform to target differentiation potential.
Results
Generation of RFP and eGFP rats with the PB transposon
As a fluorescent marker, the RFP gene has been frequently used in cell and molecular biology19. We designed a system to insert a PB vector containing the RFP or eGFP gene into rat genomes using the traditional pronuclear injection method. As shown in Fig. 1A, donor transgene (which was abbreviated TG) DNA (RFP or eGFP) was inserted into the PB vector adjacent to the EF1α promoter (Fig. 1A). For each injection, 30 ng/μl PB plasmid and 10 ng/μl PBase plasmid were co-injected into the pronuclei of zygotes to test the efficiency of the PB integration (Fig. 1B). The zygotes were obtained from Sprague Dawley (SD) and Dark Agouti (DA) strain rats. Two hundred twenty-three zygotes were manipulated via plasmid injection. One hundred ninety-seven embryos survived (88.3%) and developed to the blastocyst stage (Fig. 1C, Table 1). All blastocysts were transferred back to the uteri of pseudo-pregnant rats, and 98 of them developed to term. Of the generated pups, 44 founder pups (F0) carried the fluorescent markers, among which 26 were SD strain rats (white coat) and 18 were DA strain rats (agouti coat) (Table 1). These pups showed strong red or green fluorescence, as detected by a stereo fluorescence microscope (S165, Leica, Germany). The emission wavelength peaks were detected between 510 nm and 600 nm using in vivo imaging instruments (IVIS, PerkinElmer, USA), indicating that the marker genes had been integrated into the genomes by transposition (Fig. 1D, supplementary Fig. S1A,S1B).
Derivation of GFP- and RFP-labeled rats via the piggyBac transposon.
(A) Diagram of the PB vectors and PBase vector. The PB vector carried either a GFP or an RFP transgene driven by an EF1α promoter and efficiently transported the exogenous genes into the chromosome. (B) Microinjection of the PB and PBase vectors into the pronuclei of male zygotes; scale bar = 50 μm. (C) Images of transgenic embryos developing into blastocysts with strong GFP or RFP expression; scale bar = 100 μm. (D) Images of GFP and RFP rat pups on the DA and SD backgrounds that were generated by PB transposition.
We sacrificed one RFP rat to assess whether the PB transposon had integrated into different organs. All eight organs assessed (the stomach, heart, liver, lung, intestine, kidney, brain and spleen) showed strong red fluorescence (Fig. 2A), indicating that the PB transposon had been introduced efficiently. A PCR analysis of the RFP gene in the eight organs further confirmed this result (Fig. 2B). These data showed that RFP and eGFP (see supplementary Fig. S2A) rats can be efficiently generated using genomic transposition.
Integration of the marker genes and germline transmission.
(A) Images of the organs (stomach, heart, liver, lung, intestine, kidney, brain and spleen) dissected from one RFP transgenic rat (RFP-positive offspring from founder rats); scale bar = 1 mm. (B) PCR analysis of the integration of the desRed gene in different organs shown in panel A. All tested organs from the RFP rats were desRed gene positive compared with the wild type Dark Agouti control (the lane which was marked by DA control). (C) Images of the testes from the RFP founder rat; scale bar = 1 mm. (D) Image of RFP-positive round sperm from the testes of the RFP founder rat; scale bar = 100 μm. (E) Images of normal mature sperm that had separated from the RFP founder rat testes; scale bar = 100 μm. (F) Images of the ovaries from the RFP founder rat; scale bar = 1 mm. (G) Image of RFP-positive oocytes at the germ-vesicle (GV) stage from the ovaries of the RFP founder rat; scale bar = 100 μm. (H) F1 RFP pups generated by mating RFP rats with WT rats.
Germline transmission of RFP and eGFP
We assessed the gametes of the founder rats to determine whether the RFP and eGFP marker genes can be transmitted to the progeny through the germline. The data revealed that testes and germ cells from male rats were RFP-positive under a fluorescence microscope (Fig. 2C,D). However, mature spermatids without a cytoplasm were RFP-negative, indicating that RFP was located inside the cytoplasm (Fig. 2E). Furthermore, we performed a fluorescence-activated cell sorting (FACS) analysis to determine the percentage of RFP-positive cells in haploid germ cells (round spermatids and mature spermatids). In the RFP testes group, 34.4% of the haploid peak gated cells were RFP-positive, whereas no RFP-positive cells were observed in the wild type (WT) control haploid cells (see supplementary Fig. S2B). These data showed that male RFP rats can generate gametes with the RFP modification. In a parallel experiment, female RFP rats were assessed for the formation of RFP germ cells. The data showed that ovaries and germ cells at the mature MII oocyte and germinal vesicle (GV) stages (Fig. 2F,G and supplementary Fig. S2C) were RFP-positive, which implied that female RFP rats can also form gametes with the RFP modification by PB transposon. Furthermore, the RFP-positive rats grew to adulthood, and full-term pups (F1) carrying RFP expression were obtained after crosses with WT rats (Fig. 2H and supplementary Table S1). In this assay, the fluorescent eGFP gene is also an ideal genetic marker for rats (data not shown). In conclusion, the PB-introduced RFP or eGFP marker gene can be stably inherited by the next generation through germline transmission.
Integration of the PB transposon in the rat genome
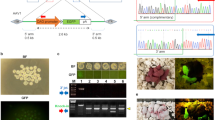
We investigated the insertion sites of the PB transposon in the genomes of the RFP and eGFP rats. Among the 10 insertion sites, 5 were inserted into an intron and 5 were inserted into intergenic sites (Fig. 3A). The five genes with PB insertion in an intron were Macf1, Tyw1, Ptpn3, Dact1 and Rad51b, which are located on chromosomes 5, 12, 5, 6 and 6, respectively (Fig. 3B). Some of the rats had a single copy insertion, but many others had multiple copy insertions (Fig. 3C). We explored the genomes of 21 RFP rats and 24 eGFP rats by inverse PCR to evaluate the efficiency of the transgene transfer (see supplementary Fig. S3A). Our data showed that the PB transposon could efficiently deliver the exogenous genes into the rat genome (see supplementary Fig. S3B,S3C). A southern blot assay was performed to identify the number of PB transposon copies. Our data showed that the PB transposon could integrate into the genome at multiple sites (Fig. 3D), which is consistent with the inverse PCR result (see supplementary Fig. S3B). Furthermore, we explored the entire genome to test whether the PBase vector integrated. Although PBase integration occurred in both RFP- and eGFP-positive rats, some of the offspring were PBase-free, with stable integration of the marker genes (Fig. 3E and supplementary Fig. S3D).
Analysis of PB insertions in the generated pups.
(A) Summary of the PB transposon integration sites. Five rats exhibited an insertion in an intron region and another five exhibited an insertion in an intergenic region. (B) Exact insertion sites of ten independent rats. (C) Inverse PCR analysis of the transposition sites in the transgenic rats. (D) Detection of the number of PB transposon copies in an RFP-positive offspring. (E) Exploration of the integration of PBase in the transgenic rats; each band indicated the random integration of the PBase gene.
RFP-positive ESCs derived from the PB-integrated rats
We next performed rat ESC derivation experiments using RFP blastocysts harvested from the PB-integrated DA strain rats. RFP was stably expressed in both E4.5 blastocysts and derivative outgrowths after 5 days of culture in N2B27 medium supplemented with “2i” (PD0325901 and CHIR99021) and leukemia inhibitory factor (LIF)20,21 (Fig. 4A). Standard rat ESC cell lines that expressed RFP were established and passaged in vitro for long periods. The expression of alkaline phosphatase (Fig. 4B) and pluripotent marker genes (Fig. 4C) indicated that the PB insertion did not affect the core pathways regulating pluripotency. We assessed whether RFP would be silenced during the ESCs culture procedure using FACS analysis. The results indicated that the percentage of RFP-positive cells at passage five was 98.7% (Fig. 4D) and was 97.8% after passage 12, with no subsequent significant decrease (Fig. 4E). Hence, PB integration can stably introduce RFP genes into the genome of the derivative ESCs, and the expression remained constant.
Derivation of ESCs from the RFP rats.
(A) RFP-positive ESCs derived from E4.5 blastocysts of RFP rats. Top panel: rat blastocysts; middle panel: outgrowth (day 5); and bottom panel: established rat ESCs; scale bar = 50 μm. (B) Alkaline phosphatase (AP) staining of RFP-rat ESC colonies; scale bar = 50 μm. (C) RT-PCR analysis of pluripotent marker gene expression in the derived RFP-ESCs. (D) Percentages of RFP-positive cells (PBES1-1) at the initial passage (P5, 98.7%) were detected by FACS analysis; scale bar = 50 μm. (E) Percentages of RFP-positive cells in PBES1-1 cells after 12 passages (P17, 97.8%); scale bar = 50 μm. (F) Chimeric rat pups derived from the RFP-rat ESCs. The chimeric rats showed a high contribution of RFP-rat ESCs, according to the coat color. (G) Expression of the desRed gene in the chimeric pups. The chimeric pups showed a high level of RFP expression compared with the negative control.
We produced chimeras by injecting blastocysts to examine the differentiation potential of these RFP-positive rat ESCs (see supplementary Fig. S4A). In two independent donor cell lines, 39 offspring were derived from 86 reconstructed and transferred embryos (Table S2). Ten chimeric rats were generated, with the contributions distinguished by coat color (Fig. 4F) and fluorescence detection (Fig. 4G). In summary, genomically integrated, pluripotent RFP-positive ESCs could be derived from PB-integrated RFP rats. This technique enables the generation of a supply of fluorescently labeled rat ESCs for future research.
In this study, we developed a simple and efficient method to generate transgenic rats that carry fluorescent marker genes. Our results showed that RFP and eGFP rats can be produced by microinjection of PB vectors carrying RFP or eGFP genes into the pronuclei of zygotes. We further showed that the PB system and fluorescence genes can synergistically form a powerful tool for genetic research in mammals.
Discussion
Sleeping beauty was the first DNA-based transposon system used for genomic engineering in mammalian cells. However, because PB was proven to efficiently deliver genes in mice in 200522, it has been widely used in gene modification. Without requiring DNA synthesis, the PB transposon can excise itself by forming a hairpin structure2, which provides a seamless excision with no “footprint”. Recently, the combination of CRISPR/Cas9 and the PB transposase system was used to produce a model to track and transform neocortical progenitors and provided a new strategy to study genetic function23,24. The protocol of co-injecting PB and PBase-encoding mRNA into pronuclei was specifically developed to improve the efficiency of producing transgenic animals and prevent re-transposition events, and can obtain average transformation frequencies of 80%25. Cytoplasmic microinjections of hyPBase mRNA and pPB-CAG-TagRFP DNA showed that 94.4% of blastocysts were TagRFP-positive26. However, it is a leap from the cellular level to an individual animal, and the injection of the PBase mRNA has great potential as an easier and highly effective method to generate transgenic animals. In this study, PBase did not show active state in the generated eGFP/RFP transgenic rats, which might relate to powerful reproductive ability of rodents. But application of this technique in larger farm animals warrants more investigations. To date, the PB system has proven itself as a versatile genetic tool for various applications, such as mutagenesis and transgenesis. Compared with sleeping beauty and other conventional viral vectors, such as retrovirus AAV and adenovirus, the PB system has a larger cargo size and a protein domain fusion that can flexibly modify the transposase to achieve site-directed integration2.
The PB transposon is preferentially biased toward transcription units, particularly in regions that contain genes. We found that out of 10 randomly selected rats, five had insertion sites in introns and five had insertion sites in intergenic regions. This system produced fluorescent rats with no deficiencies. In addition, some rats had a single copy insertion, whereas others had multiple insertions. As PB is prone to integrate near or within coding units, we have not clearly determined the reason why the rats did not carry insertion sites in exons or promoter regions.
Our study provided a new method to generate genetically modified rats expressing a fluorescent protein in different organs. RFP pups were produced by mating RFP rats with WT rats, which indicated that the fluorescent marker genes can be stably inherited by the next generation through germline transmission. This technique will enable the production of a supply of fluorescently labeled rat ESCs for further studies.
Methods
Rats
Sprague-Dawley (SD), Fisher 344 (F344) and Dark Agouti (DA) strain rats and CF-1 strain mice were purchased from the Beijing Vital River Laboratory Animal Center. All experiments involving animals were conducted according to the Guidelines for the Care and Use of Laboratory Animals established by the Beijing Association for Laboratory Animal Science and approved under the Animal Ethics Committee of the Institute of Zoology, Chinese Academy of Sciences (1 Beichen West Road, Chaoyang District, Beijing, P. R. China).
Construction of the vectors
Basic PB (PB533-A) and transpose vectors were purchased from SBI System Biosciences and then modified. The desRed (RFP) or enhanced green fluorescent protein (eGFP) genes were inserted into the PB533A-1 vector at the EcoRI and BamHI sites.
Pronuclear microinjection
The PB plasmid (30 ng/μl) carrying the RFP or eGFP genes and the PBase vector (10 ng/μl) were co-injected into pronuclei of zygotes harvested from 0.5 dpc (days post-coitus) SD and DA strain rats to produce transgenic embryos. The reconstructed embryos were transferred to the oviducts of pseudo-pregnant female rats.
Inverse PCR and genotyping
The schematic of the inverse PCR was illustrated in Fig. S3A. Briefly, the genomic DNA from each sample was extracted using the MicroElute Genomic DNA kit (Omega, USA) and digested with BstYI for 16 hours at 37 °C. The BstYI enzyme was then inactivated at 80 °C for 30 min. The ligation reaction conditions were 4 °C for 16 hours after the direct addition of T4 DNA Ligase (NEB, USA) into the inactivated digestion reaction. The PCR experiments (primers are listed in the supplementary information; see supplementary Table S3.) were performed under the following conditions: 95 °C for 5 min, followed by 25 cycles of 95 °C for 30 sec, 58 °C for 30 sec and 72 °C for 2 min, and 72 °C for 10 min for terminal replication (first round); and 95 °C for 5 min, followed by 32 cycles of 95 °C for 30 sec, 60 °C for 30 sec, and 72 °C for 90 sec, and 72 °C for 10 min for terminal replication (second round). The products of the inverse PCR were cloned into the pMD™18-T vector (TaKaRa, China) and then sequenced by Sanger sequencing with the M13F primer. The sequence files were analyzed and aligned using the BLAST tool (NCBI, USA) on the official website.
The fluorescent cassettes in the transgenic rats were amplified in a PCR assay under the following conditions: 95 °C for 5 min, 32 cycles of 95 °C for 30 sec, 59 °C for 30 sec, and 72 °C for 1 min, followed by 72 °C for 10 min.
A pair of primers (supplementary Table S3) was designed to amplify the PBase DNA in the genome, under the following conditions: 95 °C for 5 min, 32 cycles of 95 °C for 30 sec, 60 °C for 30 sec, and 72 °C for 1 min, followed by 72 °C for 10 min.
Southern blot
Genomic DNA from the transgenic rats was digested with BamHI and EcoRV at 37 °C for 16 hours, and then separated on 0.8% agarose gels prior to Southern analysis. The probe was synthesized using the Prime-a-Gene Labeling kit (Promega, lot:0000182475), which used the Neo cassette as the template, and the probe was labeled with alpha-P32dATP.
ESC derivation
Blastocysts labeled with RFP or GFP were seeded on mitomycin-C-treated mouse embryonic fibroblasts (MEFs) and cultured in the previously reported standard “2i” rat ESC medium, which was N2B27 supplemented with “2i” (PD0325901 and CHIR99021) and leukemia inhibitory factor (LIF)27. Outgrowths were picked manually and trypsinized into single cells with 0.05% trypsin-EDTA. The cells from each outgrowth (one cell line) were cultured in a new well of the plate in rat ESC medium on feeder cells. The medium was changed daily for each cell line, and the cell lines were passaged every other day.
AP staining and karyotype analysis
AP staining was performed according to the standard manufacturer’s instructions using the alkaline phosphatase kit. The results were observed under an inverted microscope (DMi-8, Leica, Germany). Karyotype analysis was performed according to standard methods28.
Chimera production
Rat blastocysts were collected from 4.5 dpc F344 strain female rats and injected with 10 to 12 RFP- or GFP-labeled ESCs. The reconstructed embryos were cultured in mR1ECM (246 mOsM)29 in a 37 °C incubator with 5% CO2 for 30 min. Images of the reconstructed blastocysts were captured on an inverted microscope (DMi-8, Leica, Germany). The chimeric embryos were transferred to the uteri of pseudo-pregnant female SD rats. The chimeric rats were identified by coat color or expression of the fluorescent proteins.
Additional Information
How to cite this article: Li, T. et al. Efficient Production of Fluorescent Transgenic Rats using the piggyBac Transposon. Sci. Rep. 6, 33225; doi: 10.1038/srep33225 (2016).
References
Cary, L. C. et al. Transposon mutagenesis of baculoviruses: analysis of Trichoplusia ni transposon IFP2 insertions within the FP-locus of nuclear polyhedrosis viruses. Virology 172, 156–169, doi: 10.1016/0042-6822(89)90117-7 (1989).
Yusa, K. piggyBac Transposon. Microbiology Spectrum 3, doi: 10.1128/microbiolspec.MDNA3-0028-2014 (2015).
Lobo, N., Li, X. & Fraser, M. J. Jr. Transposition of the piggyBac element in embryos of Drosophila melanogaster, Aedes aegypti and Trichoplusia ni. Molecular & general genetics: MGG 261, 803–810, doi: 10.1007/s004380050024 (1999).
Grossman, G. L., Rafferty, C. S., Fraser, M. J. & Benedict, M. Q. The piggyBac element is capable of precise excision and transposition in cells and embryos of the mosquito, Anopheles gambiae. Insect biochemistry and molecular biology 30, 909–914 doi: 10.1016/S0965-1748(00)00092-8 (2000).
Ren, X., Han, Z. & Miller, T. A. Excision and transposition of piggyBac transposable element in tobacco budworm embryos. Archives of insect biochemistry and physiology 63, 49–56, doi: 10.1002/arch.20140 (2006).
Wu, S. C. et al. piggyBac is a flexible and highly active transposon as compared to sleeping beauty, Tol2, and Mos1 in mammalian cells. Proc Natl Acad Sci USA 103, 15008–15013, doi: 10.1073/pnas.0606979103 (2006).
Ding, S. et al. Efficient transposition of the piggyBac (PB) transposon in mammalian cells and mice. Cell 122, 473–483, doi: 10.1016/j.cell.2005.07.013 (2005).
Wu, S., Ying, G., Wu, Q. & Capecchi, M. R. Toward simpler and faster genome-wide mutagenesis in mice. Nature genetics 39, 922–930, doi: 10.1038/ng2060 (2007).
Li, W. et al. Androgenetic haploid embryonic stem cells produce live transgenic mice. Nature 490, 407–411, doi: 10.1038/nature11435 (2012).
Woltjen, K. et al. piggyBac transposition reprograms fibroblasts to induced pluripotent stem cells. Nature 458, 766–770, doi: 10.1038/nature07863 (2009).
Li, X. et al. Calcineurin-NFAT signaling critically regulates early lineage specification in mouse embryonic stem cells and embryos. Cell stem cell 8, 46–58, doi: 10.1016/j.stem.2010.11.027 (2011).
Leeb, M., Dietmann, S., Paramor, M., Niwa, H. & Smith, A. Genetic exploration of the exit from self-renewal using haploid embryonic stem cells. Cell stem cell 14, 385–393, doi: 10.1016/j.stem.2013.12.008 (2014).
Jacob, H. J. Functional genomics and rat models. Genome Res 9, 1013–1016 (1999).
Zan, Y. et al. Production of knockout rats using ENU mutagenesis and a yeast-based screening assay. Nat Biotechnol 21, 645–651, doi: 10.1038/nbt830 (2003).
Geurts, A. M. et al. Knockout rats via embryo microinjection of zinc-finger nucleases. Science 325, 433, doi: 10.1126/science.1172447 (2009).
Tesson, L. et al. Knockout rats generated by embryo microinjection of TALENs. Nat Biotechnol 29, 695–696, doi: 10.1038/nbt.1940 (2011).
Tong, C., Li, P., Wu, N. L., Yan, Y. & Ying, Q. L. Production of p53 gene knockout rats by homologous recombination in embryonic stem cells. Nature 467, 211–213, doi: 10.1038/nature09368 (2010).
Li, W. et al. Genetic modification and screening in rat using haploid embryonic stem cells. Cell Stem Cell 14, 404–414, doi: 10.1016/j.stem.2013.11.016 (2014).
Kain, S. R. et al. Green fluorescent protein as a reporter of gene expression and protein localization. BioTechniques 19, 650–655 (1995).
Buehr, M. et al. Capture of authentic embryonic stem cells from rat blastocysts. Cell 135, 1287–1298, doi: 10.1016/j.cell.2008.12.007 (2008).
Li, P. et al. Germline competent embryonic stem cells derived from rat blastocysts. Cell 135, 1299–1310, doi: 10.1016/j.cell.2008.12.006 (2008).
Woodard, L. E. & Wilson, M. H. piggyBac-ing models and new therapeutic strategies. Trends in Biotechnology 33, 525–533, doi: 10.1016/j.tibtech.2015.06.009.
Chen, F., Rosiene, J., Che, A., Becker, A. & LoTurco, J. Tracking and transforming neocortical progenitors by CRISPR/Cas9 gene targeting and piggyBac transposase lineage labeling. Development 142, 3601–3611, doi: 10.1242/dev.118836 (2015).
van Boxtel, R. & Cuppen, E. Rat traps: filling the toolbox for manipulating the rat genome. Genome Biol 11, 217, doi: 10.1186/gb-2010-11-9-217 (2010).
Jang, C. W. & Behringer, R. R. Transposon-mediated transgenesis in rats. CSH Protoc 2007, pdb prot4866, doi: 10.1101/pdb.prot4866 (2007).
Suzuki, S., Tsukiyama, T., Kaneko, T., Imai, H. & Minami, N. A hyperactive piggyBac transposon system is an easy-to-implement method for introducing foreign genes into mouse preimplantation embryos. The Journal of Reproduction and Development 61, 241–244, doi: 10.1262/jrd.2014-157 (2015).
Buehr, M. et al. Capture of Authentic Embryonic Stem Cells from Rat Blastocysts. Cell 135, 1287–1298, doi: 10.1016/j.cell.2008.12.007 (2008).
Li, T. et al. Derivation of germline competent rat embryonic stem cells from DA rats. J Genet Genomics 39, 603–606, doi: 10.1016/j.jgg.2012.06.006 (2012).
Oh, S. H., Miyoshi, K. & Funahashi, H. Rat oocytes fertilized in modified rat 1-cell embryo culture medium containing a high sodium chloride concentration and bovine serum albumin maintain developmental ability to the blastocyst stage. Biol Reprod 59, 884–889 (1998).
Acknowledgements
We appreciate the discussion with all of the members in Dr. Zhou’s lab and Dr. Li’s lab. This work was supported by grants from the National High Technology Research and Development Program (2015AA020307) and the National Science and Technology Support Program (2014BAI02B01, 2015BAI08B02) from the Ministry of Science and Technology of the PRC. We thank Qing Meng, Qin Hua, and Shiwen Li from the State Key Laboratory of Reproductive Biology (SKLRB) for their help with FACS and confocal laser scanning microscopy. We also thank Liandi Lei and Luzheng Xu from the Medical and Health Analysis Center Molecular Imaging Laboratory, Peking University for their help with the animal fluorescence imaging experiments.
Author information
Authors and Affiliations
Contributions
T.L., W.L. and Q.Z. designed the study; T.L., L.S., J.M., X.W., M.W., X.Z., L.W. and Y.L. collected and analyzed the data; T.L., L.S. and W.L. wrote the paper. All authors revised and edited the manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Li, T., Shuai, L., Mao, J. et al. Efficient Production of Fluorescent Transgenic Rats using the piggyBac Transposon. Sci Rep 6, 33225 (2016). https://doi.org/10.1038/srep33225
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep33225
This article is cited by
-
Optimization of piggyBac Transposon System Electrotransfection in Sheep Fibroblasts
Molecular Biotechnology (2023)
-
Application of transposon systems in the transgenesis of bovine somatic and germ cells
BMC Veterinary Research (2022)
-
Long-term health and germline transmission in transgenic cattle following transposon-mediated gene transfer
BMC Genomics (2018)
-
Development of genome engineering technologies in cattle: from random to specific
Journal of Animal Science and Biotechnology (2018)
-
From engineering to editing the rat genome
Mammalian Genome (2017)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.