Abstract
Quantitative cell motility studies are necessary for understanding biophysical processes, developing models for cell locomotion and for drug discovery. Such studies are typically performed by controlling environmental conditions around a lens-based microscope, requiring costly instruments while still remaining limited in field-of-view. Here we present a compact cell monitoring platform utilizing a wide-field (24 mm2) lensless holographic microscope that enables automated single-cell tracking of large populations that is compatible with a standard laboratory incubator. We used this platform to track NIH 3T3 cells on polyacrylamide gels over 20 hrs. We report that, over an order of magnitude of stiffness values, collagen IV surfaces lead to enhanced motility compared to fibronectin, in agreement with biological uses of these structural proteins. The increased throughput associated with lensfree on-chip imaging enables higher statistical significance in observed cell behavior and may facilitate rapid screening of drugs and genes that affect cell motility.
Similar content being viewed by others
Introduction
Cell migration is an important process that is vital to both the development and maintenance of healthy organisms, such as during wound healing and to the progression of various diseases including cancer metastasis. As such, this process is complex and often dictated by a wide variety of internal and external stimuli. Examples of known external factors are mechanical forces acting on the cell, environmental conditions such as temperature and pH, presence of chemical factors such as chemoattractants and the extent of cell-cell contact determined by the local cell density. Internally, cell motility can be affected by the general state of the cell, i.e., health, mitotic phase, degree of differentiation in stem cells, etc.1,2.
The general mechanism behind cell movement on 2D surfaces involves protrusion of a leading edge forming new adhesions, followed by cell contraction and release of rear adhesions3, although besides this fibroblast-like movement an amoeboid mechanism has also been described4. Migration on 3D substrates appears to be quite different than both of these mechanisms. Since cell motility involves coordination of many biochemical signals and because there may be extreme variability between cells, the exact manner by which stimuli affect migration is unclear. Therefore, gaining an accurate quantification of motility for large populations of cells under controlled conditions would be highly useful in further developing cell migration models and may aid in developing therapeutic strategies for diseases that are either dependent on, or are caused by the misregulation of cell motility5.
In order to perform such quantitative studies on large cell populations efficiently, both imaging and cell tracking must be automated. This generally involves taking time-lapse videos using phase or bright-field optical microscopy and subsequently processing these videos with image analysis software. Several other methods have also been developed for studying cell motility. Force cytometry has been used to study the tracks that cells form on substrates during locomotion5. Phagokinetic track assays in which cells ingest markers present on the substrate as they travel, leaving behind empty spaces have also been used6,7. Both methods require markers, however. Additionally, the former focuses more on characterizing the mechanism behind cell locomotion rather than quantifying cell motility and the latter requires complex preparation steps, may influence cell behavior and state and leaves open the possibility of masked cell movements once enough markers are gone. For these reasons, time-lapse videos form the simplest and one of the most informative types of assays.
Currently, all such time-lapse videos are created using standard bench-top optical microscopes. Although image resolution is inherently good and time resolution can be optimized with advanced cameras, standard lens-based microscopy platforms have several disadvantages. First, such assays must be performed on a microscope and since it is important to monitor cell motility over the course of many hours or full days, extensive microscope occupation may be necessary posing a practical challenge to many labs. Secondly, an incubator environment must be created at the microscope. Both the temperature and carbon-dioxide content must be carefully adjusted and this requires bulky and relatively expensive microscope incubators and chambers. Finally, the throughput of such lens-based imaging assays is limited by the field-of-view (FOV) of traditional microscope objectives, which is typically smaller than 5 mm2.
Over the last few years, computational lensfree on-chip imaging has emerged and promises to be a versatile, cost-effective, field-portable and yet powerful biomedical imaging tool both in labs and at remote sites8,9. Based on digital in-line holography, this on-chip imaging technique was shown to be an alternative tool for various clinical and telemedicine applications, addressing diagnosis of diseases such as malaria10, automated cell counting11, water quality monitoring12, blood analysis13 as well as imaging cytometry14,15,16. In this work, we demonstrate that lensfree on-chip imaging can provide a solution to wide-field cell motility tracking that can overcome many of the above outlined shortcomings of lens-based optical microscopy methods.
Unlike traditional microscopy-based time-lapse video recording, lensfree on-chip imaging, with its extreme compactness, allows cell motility experiments to be conducted right inside of a conventional incubator eliminating the need for a custom-designed microscope chamber. Its large FOV (e.g., ~24 mm2) enables imaging of many more cells per experiment than it is possible with bench-top optical microscopes (which for example have a FOV of <4 mm2 with a 10× objective-lens). Further, the lensfree on-chip imaging design is such that the major component is an optoelectronic sensor array (e.g., a CMOS imager chip), resulting in a significantly more cost-effective imaging device compared to a conventional light microscope. Although holographic image reconstruction algorithms can be used with lensfree on-chip imaging to recover fine image details providing diffraction-limited resolution over a large FOV17, the raw digital holograms of the cells can also be very useful for cell tracking. Since these raw holographic images yield simplified cell signatures that vary minimally with time and position, they lend themselves to correlation-based image analysis allowing for relatively simpler cell tracking algorithms.
As a demonstration of our lensfree imaging platform's cell tracking capability, we tracked the movements of 3T3 cells on ECM-coated hydrogel substrates of varying stiffness values over 20 hours with a sampling period of 1 minute; see Figure 1 for our lensfree microscope design. We processed these videos with custom-written software to extract on average >47 cell paths from each experiment and assigned effective diffusivity coefficients to each sample based on the average distance traveled by the cells over time. These experiments indicate that 3T3 cells exhibit increased motility with increasing substrate stiffness in the tested stiffness range (0.4 kPa, 4 kPa and 40 kPa) on substrates coated with either fibronectin or collagen type IV, but also that specific ECM plays an important role in their absolute degree of motility. The automated operation and wide FOV of such a compact on-chip imaging platform make it a promising tool for efficient tracking of large cell populations. The cost-effective format of this technology also provides a framework for future developments of parallelized rapid screening platforms for drugs and genes that affect cell motility.
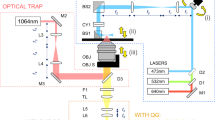
(a) A photograph of the lensfree compact on-chip microscope with a tray for loading samples (photograph taken by I.P.) (b) Cross-sectional view of (a). An LED light source, filtered by an aperture, illuminates the sample. The light scattered by the samples interferes with unscattered light at the CMOS sensor creating in-line holograms. The entire area of the CMOS sensor chip is used for imaging, yielding a FOV of 24 mm2. Power and data communication is supplied to the device through a USB cable. Relay used for triggering the image capture is not shown.
Results
Automated tracking of cell motility across a large FOV
To demonstrate our on-chip imaging platform's ability to track individual cells seeded on a substrate in low to medium confluency, we applied it to quantify the effects of substrate stiffness on cell motility. Specifically, we used it to track movements of NIH 3T3 mouse fibroblast cells on polyacrylamide gels with Young's moduli of 0.4 kPa, 4 kPa and 40 kPa, coated either with fibronectin (FN) or collagen type IV (Col-IV) over 20 hrs. For each combination of Young's modulus value and protein coating, we conducted 3 separate experiments, each of which produced two-dimensional positions of the tracked cells, every minute, for the duration of the experiments, i.e., 20 hrs. Finely resolved cell paths were reconstructed from these positions. Both the initial cell detection, carried out in the first frame of each video and all cell tracking in subsequent frames was automated using 2D intensity-based correlation calculations as detailed in the Methods Section.
In each of these 18 experiments, we achieved high N for the number of individual cells that could be successfully tracked (Figure 2). The average N from all these experiments was 47.5 and no single experiment recorded an N less than 32. Previously reported time-lapse microscopy based migration assays performed variably, with some studies reporting as few as 5–15 cells per experiment18,19 while more recently, Nenasheva et al. reported a new technique using dark-field microscopy that was able to track larger numbers of cells20. All of these studies, however, required a bench-top and lens-based incubated microscope. Our overall tracking success, defined as the percentage of initially detected cells that were successfully tracked throughout the entire video (i.e., for 20 hrs), was between 30–50% for highly motile 40 kPa and 4 kPa samples and it increased to 56–80% for the less motile 0.4 kPa samples. This tracking efficiency is summarized in Figure 2.
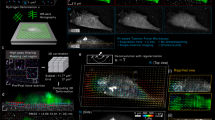
Tracking cells with lens-free on-chip imaging.
(a) Representative images of cell paths constructed from coordinates obtained automatically with our image-processing algorithm. The green ‘X’ marks the initial position of a cell and a red ‘O’ encircles the cell's final position. Each path consists of 1200 steps (sampled each minute for a duration of 20 hrs) and is plotted on the final frame of the video. (b) Number of successfully tracked cells per video. 18 total videos were recorded: three independent videos for each combination of stiffness (40 kPa, 4 kPa, 0.4 kPa) and ECM coating (fibronectin, collagen type IV). Each video yielded at least 32 and as many as 78 individual cell paths. Obstacles to successful tracking of cells included spatial artifacts present on the sample, cell deaths, cells leaving the FOV and some cell-division events.
Measurement of the cell diffusion coefficient (D)
One measure of the cell motility is average cell migration velocity. However, since this measure is highly dependent on the sampling rate, i.e., movements along a random path that may retrace itself will lead to different average migration speeds depending on the delay between frames, an alternative measure must be developed. While we include measurements of cell velocity, track lengths and mean directional persistence for ease of comparison to past studies, we also report an effective diffusion coefficient for each cell population, analogous to the diffusion coefficient in a mass-transfer setting describing particle diffusion in a fluid, but in two dimensions.
In making this analysis, we assume that the cells are executing a random walk. This assumption is justified since all the experimental conditions are designed to avoid influencing the cell motion: there are no gradients in stiffness or cell density, no chemo-attractants and substrate boundary effects are mitigated since the imaging FOV (~24 mm2) captures only the central ~22% of the gels' total areas. Additionally, we reasonably assume that the distance between the cells is large enough to prevent the contact between the cells since only 120–150 cells (producing holographic interference patterns approximately 80 μm in diameter per cell) are uniformly distributed across our large FOV at the start of the videos. Calculated diffusivity coefficients in our experiments did not show significant variation between sampling periods of 5 mins, 2, mins or 1 min (data not shown).
Since the cells are necessarily adhered to the hydrogel surface, their motion is limited to two dimensions. A two-dimensional diffusion coefficient can be extracted from the mean displacement expression <r> = √(4Dt) which can be re-written as the mean squared displacement (MSD) equation <r2> = 4Dt where <r2> is the mean of the displacement squared, t is time and D is the effective diffusivity21,22 with units of μm2 per minute. Post-processing of the global coordinates of each tracked cell in every frame, obtained from our tracking algorithm, allowed us to generate the necessary MSD-time curves. Subsequent curve fitting was used to extract/measure the value of D. In this work, we present mean displacement curves since they show the true cell movement, rather than MSD curves which are used for computation of D. These mean displacement curves are plotted both on a per-cell basis and as an averaged result among all the cells for a given set of conditions (see Figure 3).
Cell motility as a function of stiffness and surface protein.
(a) Representative ‘Displacement vs. Time’ plots for each experimental condition (0.4 kPa, 4 kPa, 40 kPa PA hydrogels coated with either fibronectin (FN) or collagen type IV (Col-IV)). Single-cell displacement from the initial position of the cell is shown over the course of 20 hrs. These plots reflect only one of the three experiments done for each set of conditions. (b) ‘Displacement vs. Time’ plots averaged across all three trials for each condition. Cells cultured on FN-coated gels were generally less motile than those cultured on gels coated with Col-IV, except in the 0.4 kPa case (most flexible). Thick lines represent the mean and thin lines represent the standard error of the mean (SEM). (c) Average diffusion coefficients calculated for each experimental condition. Average diffusivity was positively correlated with gel stiffness for both proteins. The error bars represent SEM.
Effects of Young's modulus and ECM on cell motility
Calculated diffusion coefficients for all trials are shown in Figure 3(c). The degree of cell motility is correlated with substrate stiffness. Regardless of whether the ECM used for coating was FN or Col-IV, the highest degree of cell movement occurred on gels with 40 kPa stiffness with mean calculated diffusion coefficients (D) of 29.2 μm2/min (the standard error of the mean, SEM = 6.1) and 117.2 μm2/min (SEM = 7.0) for FN and Col-IV, respectively. Gels with 4 kPa stiffness yielded the next highest degree of cell motility in each case with D values of 19.2 μm2/min (SEM = 1.9) and 44.5 μm2/min (SEM = 6.2) for FN and Col-IV, respectively. The softest gels we tested, 0.4 kPa gels, yielded mean D values of 5.1 μm2/min (SEM = 0.4) and 0.5 μm2/min (SEM = 0.1) for FN and Col-IV, respectively.
One-tailed t-tests were used to compare the measured diffusivities. The result from each condition was tested against at least three other conditions, i.e., the same ECM with the two other stiffnesses and the other ECM on the same stiffness. Additionally, the 4 kPa Col-IV result was compared to the 40 kPa FN result, totaling four comparisons for both of these conditions. Bonferroni corrections were applied accordingly23. The t-tests confirmed that 40 kPa Col-IV experiments yielded significantly higher motilities than 4 kPa Col-IV trials (p < 0.001), which in turn were much higher than motilities from 0.4 kPa Col-IV trials (p < 0.0125, corrected for four comparisons). All 40 kPa FN experiments yielded higher D values than 4 kPa FN experiments but failed to achieve statistical significance with a p-value of 0.066 (larger than the critical p-value = 0.016 corrected for three comparisons). D values from the 4 kPa FN trials were decisively greater than those from 0.4 kPa FN trials (p < 0.001).
Interestingly, these measured D values varied greatly between FN and Col-IV experiments. Col-IV coated 40 kPa gels induced a mean D value (117.2 μm2/min) nearly 4 times that induced by FN-coated gels (29.2 μm2/min) (p < 0.016, corrected for three comparisons). This disparity, though not as pronounced, also held for 4 kPa gels on which Col-IV coatings induced a larger mean D (44.5 μm2/min) than FN coatings (19.2 μm2/min) (p < 0.0125, corrected for four comparisons). In fact, all 4 kPA Col-IV gels had larger D values than any 40 kPa FN gels, achieving a p value of 0.056. This pattern of Col-IV coated gels inducing greater movement than FN coated gels of the same stiffness was broken for the 0.4 kPa stiffness. For that stiffness, FN coated gels induced a mean D value (5.1 μm2/min) that is 10 times greater than the mean D value recorded for Col-IV coated gels (0.5 μm2/min) (p < 0.001).
Traditional measurements of cell motility on two-dimensional substrates
Mean cell velocity, mean track length and directional persistence were also determined for each condition (Figure 4). These measurements were made using both 10 minute and 30 minute sampling intervals to allow for easier comparisons to previous work on cell motility. Although these metrics were highly dependent on sampling rate, the trends seen in our diffusivity measurements were generally preserved. Specifically, cell velocity (directly indicative of the degree of cell motility) correlated positively with increasing stiffness. Likewise, Col-IV promoted greater cell motility than FN on stiffer gels but much less on the softest gels. One-tailed t-tests confirmed that velocity was greater for 40 kPa Col-IV gels than 40 kPa FN gels (p < 0.0125, corrected for 4 comparisons) and for 0.4 kPa FN gels than for 0.4 Col-IV FN gels (p < 0.016, corrected for 3 comparisons) for both sampling intervals. Velocity on 4 kPa Col-IV gels was only marginally greater than on 4 kPa FN gels (p = 0.069 for 10 min intervals, p = 0.067 for 30 minute intervals). Directional persistence was also greater for cells on the stiffer gels coated with either ECM than for the 0.4 kPa gels (p < 0.001 in all cases). Comparisons between the two ECMs showed that directional persistence was higher on 40 kPa Col-IV gels than on 40 kPa FN gels (p < 0.0125 for both sampling rates) and on 0.4 kPa FN gels than on 0.4 kPa (p < 0.0125 for both sampling rates). Comparisons of the ECMs on 4 kPa gels showed no significant differences.
Standard measurements of cell motility using 10 minute and 30 minute intervals.
(a) Mean cell velocity and mean track length over 20 hrs. Mean cell velocity in μm/min, shown on the left y-axis, was calculated by dividing the mean displacement per interval by the interval time. Mean track length in mm, shown on the right y-axis, was calculated by averaging the sum of the displacements from each interval. Thus, the two values are related by a constant equal to the total time of tracking, i.e., 20 hrs. (b) ‘Directional Persistence’, defined as net displacement divided by corresponding track length, over the full 20 hrs. The differences between 10 minute interval and 30 minute interval measurements reflect the loss of information associated with lower frequency sampling. The error bars indicate the standard error of the mean (SEM). Specific values for each measurement and SEM are reported in Supplementary Table 1.
Discussion
Our results serve as a proof-of-concept that lensfree on-chip imaging capably performs automatic cell tracking over a large FOV within a high temperature and humidity culture environment. In our experiments, we set out to characterize our system's performance by applying it to study the effects of specific mechanical cues on cells. The success of these initial cell tracking experiments suggests that this computational on-chip imaging platform could be used for future studies such as rapid screening of drugs that promote or inhibit cell motility. Additionally, the same on-chip imaging platform can be expanded to study cell attributes such as morphology, polarizability or other natural or inducible behaviors that can be tracked using lensfree holographic images.
As demonstrated in our Results section, our on-chip imaging system tracks cells individually and records specific parameters for each cell instead of giving a bulk measurement for a given population of cells. A high-throughput tool allowing such single-cell analysis becomes increasingly useful when rare behaviors are present in a sample as their contributions to a bulk measurement of motility such as obtained with a trans-well migration assay will be averaged and, hence, rare outliers can be overlooked/unnoticed when looking at a large population. However, if needed, our single-cell level data set still allows for a bulk evaluation of a sample of cells, since individual trajectories can be averaged or individual displacement vectors can be super-imposed. In its current state, our on-chip imaging platform and tracking algorithms are best suited for handling cells seeded in low to medium confluency. Higher spatial densities cause greater cell-cell contact and may lead to difficulties in monitoring the same cell for the duration of the imaging experiment. Our chosen metric, the diffusion coefficient, is most applicable when the assumption of cell isolation holds and since this would no longer be the case at higher seeding densities, a different metric would need to be created to quantify cell motility.
In our cell tracking experiments, typically, more than 120 cells are detected in the initial frame covering 24 mm2 FOV and of the lost cell tracks, the majority are the result of cell division or death events. When a cell first divides, the resulting daughter cells can overlap causing the center of the best correlation to be assigned (in some cases) to the intersecting region of the cells rather than to either daughter cell's center. This loss of true reference might propagate to future frames and a cell's actual center may not be recovered after a cell division event causing failure in tracking the divided cells in some cases. If a further increase in N is desired in certain applications, more advanced automated decision-making processes may be implemented to better handle such cell division events.
Cell death events are also partially responsible for losing some of the cell tracks. While this is unavoidable, the holograms for dead cells have unique features such as a low light intensity at the central region and lower overall intensity making them easy to detect during post-processing. Similar observations were made by S. Kesavan et al. about the lensfree holograms of adhered cells as compared to floating cells obtained using on-chip imaging24. Note that the lost cells due to cell division were taken into account in evaluating our tracking success while dead cells were not. Also note that the relatively stagnant cell motion on Col-IV coated 0.4 kPa gels corresponded to minimal morphological diversity per cell over time and naturally allowed greater tracking success for our hologram intensity based correlation calculations.
Our lensfree on-chip imaging platform (Figure 1), as detailed in the Methods section, is rather compact, light-weight and cost-effective. Its crucial components are an LED based light source and a CMOS image sensor chip, both of which are robust in a variety of environments, allowing the on-chip imaging device to be placed directly inside of a standard laboratory incubator instead of requiring that an incubator-like customized environment be created around the imaging platform, as is necessary with traditional bench-top optical microscopes. Doing this is much less expensive and ensures that the cells are studied in the very same environment as they are cultured in. In our experiments, a thermocouple was also inserted into our imaging platform to confirm that, even though the cells are subjected to periodically pulsing LED light in a small, enclosed space, the internal temperature did not exceed 37°C for the duration of the experiments. The relay used to trigger the image capture and the laptop computer running the software connect to the lensfree microscope through a USB cable and therefore are left outside of the incubator. While this study was done using relatively large mammalian cells (spread areas of up to 2000 μm2 on FN- or Col-coated PA gels25), lensfree on-chip imaging can in general be used to detect micro-objects even smaller than e.g., 0.5 μm as demonstrated in our earlier work (see e.g., Ref. 9). Therefore we believe that this platform should be suitable for tracking the motion of any mammalian cell.
Past studies on the effects of substrate stiffness on cell motility have had a diverse set of results. For example, a study on wound healing showed that the cell migration speed toward the wound increased with increasing substrate stiffness. Another study on durotaxis showed that upon crossing the boundary from a soft substrate to a rigid one, cell migration speeds increased26. A separate study found that cell migration speed, instead, monotonically decreased with increasing cell stiffness27. This previous work, however, tracked only ~15 cells per experiment and followed cells at 2 min intervals for 10 mins in total. Our experiments were longer and higher in throughput as we sampled 30–70 cells each minute for a duration of 20 hrs. Most previous studies agree that on substrates of uniform stiffness, cell migration speeds vary with stiffness in a biphasic manner26,18. For example, Peyton et al. found that across a stiffness range of 1–308 kPa, migration speeds reached local maximums near 20 and 50 kPa. This previous study took measurements at 1 kPa, 21 kPa and 45 kPa stiffnesses (along with higher stiffnesses) and the shape of their stiffness-migration speed curve indicates that motilities at 40 kPa are in fact greater than those at 4 kPa.
In addition to the expected differences in cell motility caused by substrate stiffness, significant differences were also seen in our experiments between the two tested ECMs. In general, cells were much more motile on Col-IV with mean diffusion coefficients 4 times larger than FN on 40 kPa gels and more than twice as large as FN on 4 kPa gels. This result supports a past study on the effects of various ECMs on cell behaviors such as proliferation, spreading and migration which showed that collagen type IV (compared with laminin, fibronectin, collagen type I and Matrigel) enhanced migration the most for mouse embryonic stem cells cultured in well-plates31. In our experiments done on 0.4 kPa gels, cells grown on Col-IV were actually less motile than cells grown on FN, although the calculated diffusion coefficients for both cases were substantially smaller compared to the other experimental conditions (see Figure 3). This suggests that the substrate stiffness may have been a greater contributor to cell motility than the ECM in that lower stiffness range. Such quantitative data on ECM- and stiffness-specific effects become important when designing scaffolds for tissue engineering applications and may be useful in, for example, improving existing transplantation strategies.
Although we demonstrate a proof-of-concept, several additional steps must be taken before this platform can provide practical utility in the field of drug and gene screening. First, as-is, the device can image only one sample at a time, prohibiting simultaneous testing of multiple environmental conditions and/or drugs or genes. The loading tray would need to be modified to accommodate many segregated chambers and a mechanism enabling movement of either the loading tray or CMOS sensor would need to be developed to realize this capability. Since the device architecture is 3-D printed, modification of the loading tray would be inexpensive. Furthermore, as a possible solution, LabVIEW, (already used to automate image capture) may be used to control miniature servo motors for driving the modified loading tray along a single track in the image plane. Additionally, the current coverslip sandwich method of sealing the samples (described in the Methods Section), could be further simplified to streamline sample preparation and imaging through, for example, replacing the individual top coverslips with a single lid spanning the length of the multi–compartment tray.
While some previous experiments studying cell migration provided valuable insights with high statistical significance20, most could provide only limited statistics since N for the tracked cells was less than optimal18,19. For this reason, many of the observed cell migratory behaviors that have been reported cannot be assessed quantitatively with the degree of confidence needed to successfully apply this knowledge to developing bioengineering applications. Furthermore, while the studies do offer preliminary indications of various behaviors, a more comprehensive platform enabling high N will be needed to confirm them. Moving forward, a lensfree on-chip cell-tracking platform would be one such platform enabling investigators to make these cell migration measurements with the higher sampling-rates (e.g., every minute or even more frequently) over longer periods of time (>20 hrs) and with much higher N per experiment, necessary for obtaining sufficient statistical significance to make accurate claims. Importantly, our platform will allow these studies to be conducted without a bench-top microscope and its associated costs and securely within an incubator environment.
Methods
Lensfree on-chip imaging set-up
Our lensfree on-chip imaging platform operates by recording the interference of the light that is scattered from the target objects with the unscattered background light, which is digitally sampled using a complementary metal-oxide semiconductor (CMOS) imager chip (Aptina MT9P031STC, 5 megapixels, 2.2 μm pixel size, monochrome). Briefly, a glass cover slip containing the cells of interest, located on a sliding tray approximately 1 mm above the CMOS chip, is illuminated with a single green LED that is spatially masked by a 0.1 mm pinhole, which is used to fine tune the spatial coherence diameter at the cell and detector planes so that lensfree holograms can be sampled at the CMOS imager chip. The overall fringe magnification of this system is ~1 and therefore the entire active area of the CMOS chip becomes the imaging FOV, where the cells can be imaged across e.g., ~24 mm2, which is around 10-fold larger than the FOV of a typical 10× objective-lens. The complete mode of operation and the physics of this on-chip imaging platform has been thoroughly described previously13,28.
All of the mentioned components were housed in a 3D printed enclosure, specifically designed for use with our samples that features a 1 cm tall slot through which the tray containing the samples was loaded. A USB port on the side of the device was used both for power and data communication. The device was secured to a shelf inside a standard incubator using autoclave tape. A relay, used to trigger the image capture, was placed atop the incubator and a cable connecting it to the device was fed into the incubator and plugged into the port on the device. Custom LabVIEW software on a nearby laptop was used to control the relay as well as to set experimental parameters such as sampling rate (every 1 min in these experiments) and exposure time (80 ms was used in all videos). The entire imaging platform was not found to be affected by the incubator environment even after >500 hrs of use.
Preparation and coating of polyacrylamide substrates
The general procedure for creating polyacrylamide hydrogels with various stiffness by mixing appropriate concentrations of acrylamide (Acr) and bis-acrylamide (bis-Acr) and subsequently inducing free radical polymerization has been described in detail by Tse et al.29. Referencing their concentration-stiffness tables, we determined appropriate amounts of these reagents needed to create hydrogels with Young's moduli of 0.4 kPa (3% w/v Acr and 0.06% w/v bis-Acr), 4 kPa (5 w/v Acr and 0.15% w/v bis-Acr) and 40 kpa (8% w/v Acr and 0.5% w/v bis-Acr). The final PA solutions consisted of the appropriate amounts of acrylamide and bisacrylamide diluted in deionized H20 and sterile-filtered 0.5% (w/v) Irgacure 2959 photoinitiator in PBS.
First, a continuous square gasket small enough to be contained on a square 22 mm × 22 mm glass coverslip was cut from a 1 cm tall slab of PDMS and bonded to the coverslip using plasma-based glass-PDMS bonding, which has been extensively described elsewhere30. The square cover slip and a smaller circular coverslip (12 mm diameter) were then treated with O2 plasma for ~1 min. The PDMS-containing surface of the square coverslip was made hydrophilic with treatment with 3-aminopropyl-triethoxy silane (APTS) for 40 minutes. Similarly, the circular coverslip was made hydrophobic with treatment with triethoxy(octyl)silane (OTES). Each treatment consisted of 1% (v/v) appropriate silane solution in a 5% (v/v) H2O in ethanol (EtOH) solvent, with 10% (v/v) acetic acid as an acid catalyst. Following silane treatment, both coverslips were washed with 95% EtOH. Additionally, the square coverslip was treated with 50% (v/v) grade 1 glutaraldehyde in dd-H20 for 30 mins. Both coverslips were then allowed to incubate in dd-H20 overnight. Finally, both coverslips were cleaned from smudges and particles with sonication in dd-H20.
After proper cleaning, the round coverslip was elevated at least to the height of the PDMS gasket on a narrow glass rod and secured with tape from underneath, with the silane-treated side facing up. For this we used a shortened glass Pasteur pipette with a rubber stopper as a base (see Figure S1(c)). Small strips of spun-on PDMS ~100 μm in height were placed at three positions on the round coverslip to act as spacers. 9 μL of the appropriate acrylamide-bisacrylamide solution was then pipetted onto the center of the circular coverslip such that it did not contact the spacers and sandwiched between the two coverslips. The set-up was cured from above with UV light for 3.5 mins. After curing, the circular coverslip was carefully removed from the square coverslip to which the cured gel was left securely attached.
In order to attach protein to the gels, a 0.2 mg/ml sufosuccinimidyl-6-(4-azido-2-nitrophenylamino)-hexanoate (sulfo-SANPAH; Pierce Biotechnology) solution was prepared in 50 mM HEPES solution, pH 8.5. The sulfo-SANPAH solution was pipetted onto the cured hydrogels using enough volume to fully cover them and the gels were placed under UV light for 1–2 mins. The gels were then washed 3 times with HEPES solution to remove unreacted sulfo-SANPAH. 30 μl of 50 μg/mL solutions of fibronectin (FN) in PBS or collagen type IV (Col-IV) in HEPES were pipetted onto the gel. The FN coated gel was allowed to sit at room temperature for 2 hrs while Col-IV was incubated with the gel at 37°C overnight. After successful ECM conjugation, the gel was washed 3 times with and stored in PBS until cells were ready to be seeded. Uniform ECM conjugation across the surface was confirmed with fluorescent intensity measurements of FITC-conjugated ECM compared with unconjugated ECM, FITC-conjugated ECM without using sulfa-SANPAH, no ECM and FITC-conjugated ECM intentionally denatured with addition of HCl (data not shown).
Cell culture and seeding
NIH 3T3 cells were cultured in high glucose Dulbecco's Modified Eagle Medium with 10% fetal bovine serum and 1% penicillin-streptomycin in T75 polystyrene flasks at 37°C with 5% C02 content. Upon reaching 100% confluency, the cells were suspended in DMEM and diluted to ~25,000 cells/mL. PBS was aspirated from the gels and ~15 μL of cell solution was carefully pipetted onto the gel. Due to the hydrophobic properties of the ECM, the pipetted solution had a high contact angle and had to be spread across the entirety of the gel with a pipet tip for a uniform surface cell density. Care was taken to avoid deforming the gel surface as this may have impacted the cells' paths. Care was also taken not to allow the cell solution to spill off of the gels as very few cells would be left to adhere to the ECM. Once this solution was spread, an additional square coverslip was placed over the top of the gasket and the gels were incubated at 37°C for 1.5 h. After incubation, the gels were gently washed with DMEM to remove non-adhered cells. The volume enclosed by the gasket was then fully filled with DMEM. A coverslip was again placed on top of the gasket such that there was no air pocket between the media and the coverslip (Figure S1(e)). This slowed evaporation of the media allowing for longer on-chip imaging. Since the gasket was cut from PDMS, a gas permeable elastomer, CO2 gas could freely perfuse into the cell chamber.
Tracking algorithm and cell data extraction
Adherent cells such as NIH 3T3 cells used in our work assume a basic circular morphology upon initial adherence to a substrate, independent of its stiffness. This characteristic guarantees that all the cells seeded together will have very similar two dimensional lensfree hologram signatures after seeding. Therefore, at the time of initial adherence, all the cells will exhibit strong similarity in their shape and spatial holographic patterns enabling the use of a single template for their initial detection. The user selects and crops a cell from the first frame to serve as this template.
Next, depending on the mechanical properties of the substrates, the cells will elongate in various orientations as they begin their random walk and a single template will no longer be useful in detecting all the cells within a given frame. Instead, an automatically cropped cell from a given frame is used as its own unique template in the following frame and this template is subsequently updated for each cell that is being tracked. In this manner, all the cells serve as their own templates beginning in frame 2. Global coordinates and a cropped image are saved for each detected cell in every lensfree frame.
MATLAB was used to implement the following 4-step algorithm using the above outlined 2D intensity correlation technique:
-
Step 1: Use single template (T0) to detect all the cells (all Ck) in frame 1 (F1).
-
Step 2: For each detected cell Ck, obtain the global coordinates (xCk_fi, yCk_fi) and crop a box of a fixed size centered at the cell's coordinates to serve as a new cell template (Ti_Ck) in the next frame.
-
Step 3: For each previously detected cell, use the unique template (Ti_Ck) and coordinates obtained in Fi-1 to scan a small area near the cell's previous location (xCk_fi-1 ± δ, yCk_fi-1 ± δ) in Fi to detect the cell in its new location on the substrate.
-
Step 4: Repeat steps 2–3 for all the subsequent lensfree frames.
Prior to running the algorithm, all frames for a given video were registered to ensure reliable position comparisons and to eliminate artificial movements and FOV changes that may be caused by the slight dimensional mismatch between the loading tray and device itself due to the inherent tolerances (e.g., <50 μm) of the 3-D printer used to print the components. ImageJ plugin StackReg, an intensity based sub-pixel image registration algorithm, was used to register all the frames using the marked corners on the substrates as coordinate references31.
The saved cropped images of each detected cell in every frame were analyzed to determine which cells were successfully tracked. Lost cells, identified when the positional crops in any iteration of the algorithm contained no cell and dead cells, identified by their unique low-contrast and perpetually circular holograms, were removed from our analysis. MATLAB was used for post-processing of the saved coordinates of each cell. These spatial coordinates provided all the information needed to determine the total path traveled and the displacement at every frame. Although the samples were imaged for 24 hrs (1440 frames), the first 4 hours were not evaluated to allow the cells seeded from suspension sufficient time to reform connections and adhere to the substrate through focal adhesions.
Change history
05 September 2014
A correction has been published and is appended to both the HTML and PDF versions of this paper. The error has been fixed in the paper.
References
Thurston, G. & Palcic, B. 3T3 cell motility in the temperature range 33°C to 39°C. Cell Motil. Cytoskeleton 7, 361–367 (1987).
Hartmann-Petersen, R., Walmod, P. S., Berezin, A., Berezin, V. & Bock, E. Individual cell motility studied by time-lapse video recording: Influence of experimental conditions. Cytometry 40, 260–270 (2000).
Li, S., Guan, J.-L. & Chien, S. Biochemistry and biomechanics of cell motility. Annu. Rev. Biomed. Eng. 7, 105–150 (2005).
Yoshida, K. & Soldati, T. Dissection of amoeboid movement into two mechanically distinct modes. J. Cell Sci. 119, 3833–3844 (2006).
Álamo, J. C. et al. Spatio-temporal analysis of eukaryotic cell motility by improved force cytometry. Proc. Natl. Acad. Sci. 104, 13343–13348 (2007).
Nogalski, M. T., Chan, G. C. T., Stevenson, E. V., Collins-McMillen, D. K. & Yurochko, A. D. A quantitative evaluation of cell migration by the phagokinetic track motility assay. J. Vis. Exp. JoVE e4165 (2012). 10.3791/4165.
Niinaka, Y., Haga, A. & Raz, A. Quantification of cell motility: gold colloidal phagokinetic track assay and wound healing assay. Methods Mol. Med. 58, 55–60 (2001).
Ozcan, A. & Demirci, U. Ultra wide-field lens-free monitoring of cells on-chip. Lab. Chip 8, 98–106 (2007).
Greenbaum, A. et al. Imaging without lenses: achievements and remaining challenges of wide-field on-chip microscopy. Nat. Methods 9, 889–895 (2012).
Seo, S. et al. High-Throughput Lens-Free Blood Analysis on a Chip. Anal. Chem. 82, 4621–4627 (2010).
Su, T.-W., Erlinger, A., Tseng, D. & Ozcan, A. Compact and Light-Weight Automated Semen Analysis Platform Using Lensfree on-Chip Microscopy. Anal. Chem. 82, 8307–8312 (2010).
Mudanyali, O., Oztoprak, C., Tseng, D., Erlinger, A. & Ozcan, A. Detection of waterborne parasites using field-portable and cost-effective lensfree microscopy. Lab. Chip 10, 2419–2423 (2010).
Zhu, H. et al. Cost-effective and rapid blood analysis on a cell-phone. Lab. Chip 13, 1282–1288 (2013).
Wei, Q. et al. On-Chip Cytometry using Plasmonic Nanoparticle Enhanced Lensfree Holography. Sci. Rep. 3, (2013).
Su, T.-W., Xue, L. & Ozcan, A. High-throughput lensfree 3D tracking of human sperms reveals rare statistics of helical trajectories. Proc. Natl. Acad. Sci. 109, 16018–16022 (2012).
Bishara, W., Isikman, S. O. & Ozcan, A. Lensfree Optofluidic Microscopy and Tomography. Ann. Biomed. Eng. 40, 251–262 (2012).
Greenbaum, A. et al. Increased space-bandwidth product in pixel super-resolved lensfree on-chip microscopy. Sci. Rep. 3, (2013).
Peyton, S. R. & Putnam, A. J. Extracellular matrix rigidity governs smooth muscle cell motility in a biphasic fashion. J. Cell. Physiol. 204, 198–209 (2005).
Wong, J. Y., Velasco, A., Rajagopalan, P. & Pham, Q. Directed Movement of Vascular Smooth Muscle Cells on Gradient-Compliant Hydrogels†. Langmuir 19, 1908–1913 (2003).
Nenasheva, T. A., Carter, T. & Mashanov, G. I. Automatic tracking of individual migrating cells using low-magnification dark-field microscopy. J. Microsc. 246, 83–88 (2012).
Berg, H. C. Random Walks in Biology. (Princeton University Press, 1993).
Qian, H., Sheetz, M. P. & Elson, E. L. Single particle tracking. Analysis of diffusion and flow in two-dimensional systems. Biophys. J. 60, 910–921 (1991).
Glantz, S. A. Primer of biostatistics. (McGraw-Hill, Medical Pub. Div., 2002).
Vinjimore Kesavan, S. et al. Lensless imaging system to quantify cell proliferation. in (Farkas D. L., Nicolau D. V., & Leif R. C.) 858708–858708–11 (2013). 10.1117/12.2001826.
Yeung, T. et al. Effects of substrate stiffness on cell morphology, cytoskeletal structure and adhesion. Cell Motil. Cytoskeleton 60, 24–34 (2005).
Lo, C.-M., Wang, H.-B., Dembo, M. & Wang, Y. Cell Movement Is Guided by the Rigidity of the Substrate. Biophys. J. 79, 144–152 (2000).
Pelham, R. J. & Wang, Y. Cell locomotion and focal adhesions are regulated by substrate flexibility. Proc. Natl. Acad. Sci. U. S. A. 94, 13661–13665 (1997).
Tseng, D. et al. Lensfree microscopy on a cellphone. Lab. Chip 10, 1787–1792 (2010).
Tse, J. R. & Engler, A. J. in Curr. Protoc. Cell Biol. (John Wiley & Sons, Inc., 2001). at http://onlinelibrary.wiley.com/doi/10.1002/0471143030.cb1016s47/abstract.
Sia, S. K. & Whitesides, G. M. Microfluidic devices fabricated in Poly(dimethylsiloxane) for biological studies. ELECTROPHORESIS 24, 3563–3576 (2003).
Thevenaz, P., Ruttimann, U. E. & Unser, M. A pyramid approach to subpixel registration based on intensity. IEEE Trans. Image Process. 7, 27–41 (1998).
Acknowledgements
Di Carlo Research Group acknowledges the support of NSF Grant #0930501. Ozcan Research Group acknowledges the support of the Presidential Early Career Award for Scientists and Engineers (PECASE), Army Research Office (ARO) Life Sciences Division, ARO Young Investigator Award, National Science Foundation (NSF) CAREER Award, NSF CBET Biophotonics Program, NSF EFRI Award, Office of Naval Research (ONR) Young Investigator Award and National Institutes of Health (NIH) Director's New Innovator Award DP2OD006427 from the Office of the Director, National Institutes of Health.
Author information
Authors and Affiliations
Contributions
I.P. performed all the experiments, analyzed data and assisted in image processing. Y.L., T.S. and O.M. developed the lensfree imaging modules and assisted in data analysis. W.W. helped design experimental protocols. I.P., D.D. and A.O. wrote the manuscript. D.D. and A.O. supervised the project.
Ethics declarations
Competing interests
A.O. is the co-founder of a start-up company (Holomic LLC) which aims to commercialize computational imaging and sensing technologies licensed from UCLA.
Electronic supplementary material
Supplementary Information
Supplemental Material
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-ShareAlike 3.0 Unported License. The images in this article are included in the article's Creative Commons license, unless indicated otherwise in the image credit; if the image is not included under the Creative Commons license, users will need to obtain permission from the license holder in order to reproduce the image. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/3.0/
About this article
Cite this article
Pushkarsky, I., Liu, Y., Weaver, W. et al. Automated single-cell motility analysis on a chip using lensfree microscopy. Sci Rep 4, 4717 (2014). https://doi.org/10.1038/srep04717
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep04717
This article is cited by
-
On-chip Microscopy Using Random Phase Mask Scheme
Scientific Reports (2017)
-
Connecting individual to collective cell migration
Scientific Reports (2017)
-
Live-single-cell phenotypic cancer biomarkers-future role in precision oncology?
npj Precision Oncology (2017)
-
Compact, cost-effective and field-portable microscope prototype based on MISHELF microscopy
Scientific Reports (2017)
-
Flatbed epi relief-contrast cellular monitoring system for stable cell culture
Scientific Reports (2017)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.