Abstract
Next-generation sequencing (NGS) has been applied to various kinds of omics studies, resulting in many biological and medical discoveries. However, high-throughput protein-protein interactome datasets derived from detection by sequencing are scarce, because protein-protein interaction analysis requires many cell manipulations to examine the interactions. The low reliability of the high-throughput data is also a problem. Here, we describe a cell-free display technology combined with NGS that can improve both the coverage and reliability of interactome datasets. The completely cell-free method gives a high-throughput and a large detection space, testing the interactions without using clones. The quantitative information provided by NGS reduces the number of false positives. The method is suitable for the in vitro detection of proteins that interact not only with the bait protein, but also with DNA, RNA and chemical compounds. Thus, it could become a universal approach for exploring the large space of protein sequences and interactome networks.
Similar content being viewed by others
Introduction
Since its inception, next-generation sequencing (NGS) has been employed to collect various types of “omics” data1, resulting in many scientific findings in biology and medicine. However, current high-throughput protein-protein interactome datasets have low coverage2,3. For example, the fraction of the human interactome that has been identified to date is estimated to be less than 10%2. Recently, Yu et al.4 reported that a massively parallel interactome-mapping pipeline for the yeast two-hybrid system coupled with NGS produced an interactome dataset more efficiently than the conventional method5 using Sanger sequencing. However, the data coverage is limited by numbers of colonies, even if using NGS. Moreover, a large amount of low reliability data could be produced at the same time, resulting in the as yet unsolved problem of the high false-positive rate in high-throughput production of interactome data6.
To simultaneously address coverage and reliability problems in the interactome, we have developed the IVV-HiTSeq (IVV high-throughput sequencing) method, which is a combination of NGS and the in vitro virus (IVV) method7,8, an mRNA display method chosen from among cell-free display technologies. In the present study, the Roche 454 Genome Sequencer FLX System (454 sequencer) was used as the next-generation sequencer. In the IVV method, proteins are covalently linked to corresponding mRNAs encoding them and can be detected by reverse transcription-PCRs (RT-PCRs) and sequencing of the mRNA moieties. The IVV method employs a complete in vitro treatment with cDNA libraries (extracted from cells and tissues) and has >1012 different molecules more than the capacity of Sanger sequencing and other high-throughput protein selection methods9,10. Thus, NGS is expected to permit the analysis of the abandoned fraction of the interactome. Selections using the IVV method are conducted under cell-free conditions and subsequent sequencing by NGS is not limited by cloning steps using any kind of cells. Thus, our method consists of completely cell-free procedures and detection of the interactions is highly efficient. Current high-throughput interactome datasets usually require post-screening assays to reduce the number of false positives and increase the reliability of the dataset. Similarly, the conventional IVV method requires post-screening assays, such as quantitative real-time PCR7,8. In the IVV-HiTSeq method, an in silico analysis of the quantitative data obtained by counting library-specific barcode tags is conducted instead of the verification assay. The results of a comparison with real-time PCR assays are also described to demonstrate the ability of the in silico analysis.
Results
An overview of IVV-HiTSeq and its two major parts are shown in Fig. 1. The first part is the in vitro selection, which follows the procedure of the previously reported mRNA display method using IVV7. The second part includes the NGS procedure and the subsequent in silico analysis. RT-PCR amplifications with 4-base barcoded primers specific for the selection libraries were employed to deal with the large amount of sequenced reads derived from the mixture of selection libraries. The barcoded RT-PCR products allowed an in silico quantitative analysis of interaction sequence tags in each round of selection. For the negative control, the same procedure was conducted in the absence of bait protein [bait(−)]. Finally, the bait(+), bait(−) and pre-selection samples (initial library) were separately sequenced by the 454 sequencer.
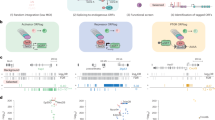
Overview of IVV-HiTSeq as a completely cell-free system for detecting interactors of a target bait protein.
First, the initial cDNA library from poly(A)+ RNAs is created by random priming. cDNA libraries are transcribed into mRNA and PEG+Puromycin spacers are ligated to their 3′ ends. mRNA-protein molecules, linked via puromycin, are formed during in vitro translation. Prey molecules that interact with tagged bait proteins are then captured by affinity beads and purified. The mRNA moieties of selected prey molecules are amplified by RT-PCR using two types of primers; one for the next selection round and another for high-throughput sequencing. The second type of primer contains a barcoded region (indicated in grey, green, blue, yellow and red), with four selection-round-specific bases. After four rounds of selection, the RT-PCR products that were amplified using the barcoded primers are mixed and analyzed together by high-throughput sequencing. The same procedure without bait protein was performed as the negative control. The reads generated by high-throughput sequencing are sorted by their barcoded parts and mapped to known genomic sequences. Read frequencies for each genomic position are calculated for each selection round and used to determine the enriched regions. Statistical significance was calculated by comparing the read frequencies with the frequencies of the initial library and the negative control.
To demonstrate the IVV-HiTSeq method, the above procedure was iterated for four rounds to enrich prey proteins that interacted with mouse interferon regulatory factor 7 (Irf7) from a randomly fragmented cDNA library created from mouse spleen. The primary sequence data included 206,322 reads for the bait(+), 304,504 reads for the bait(−) and 277,833 reads for initial library samples (see Supplementary Table S1). After eliminating erroneous reads, selection-round information was assigned to each read based on its round-specific barcoded sequence. This process yielded 177,935, 278,816 and 238,683 reads for the bait(+), bait(−) and initial libraries (see Supplementary Table S1). Finally, 47,849, 63,306 and 102,092 post-mapping reads were obtained for the bait(+), bait(−) and initial libraries, respectively. These sets of reads were then mapped to the genomic sequences (see Supplementary Table S1) and formed the datasets that were used in the subsequent in silico analysis to identify true positives, without the need for real-time PCR verification assays.
To validate the accuracy of the in silico analysis of IVV-HiTSeq, we compared the results of in silico analysis with real-time PCR assays for 21 interacting regions (IRs) that were randomly selected from all the IRs (including false-positive candidates) that overlapped with sequences in the NCBI RefSeq database. Full details of the results of this comparison can be found in Supplementary Table S2 and Supplementary Fig. S1. First, read frequencies in each IR per selection round were calculated. These frequencies were based on the number of aligned reads in each IR. Two examples of the results of the comparison between numbers of reads obtained by NGS and numbers of molecules quantified by real-time PCR assays are shown in Fig. 2a,b. The correlation coefficients between the NGS and real-time PCR datasets were calculated (Fig. 2c) and this confirmed a highly positive correlation (Pearson's correlation coefficient = 0.92) between the two. Using this ability of IVV-HiTSeq for quantification, we determined whether or not each of the 21 IRs was a true IR with statistical significance (P < 0.001). P values were calculated using Fisher's exact probability test for 2 × 2 contingency tables. Each contingency table consists of the number of read at a given region for a given round of the bait(+) and bait(−) experiments and the total numbers of reads for the corresponding experiments in the selection round being compared. Differences between the initial and given rounds were compared in the same manner.
Comparison between real-time PCR data and the read frequency of 454 sequencing.
(a, b) Upper panels show the results for reads mapped to genes by selection rounds with the bait Irf7 [bait(+)] and in the absence of bait [bait(−)]. Lower panels show the corresponding results of quantification by real-time PCR. (a) Gtpbp4, positive example; and (b) Mpeg1, negative example. (c) Scatter plots of real-time PCR versus 454 sequencing. Twenty-one regions were targeted and 105 pairs of data were obtained in each round of selection in the presence or absence of bait.
Real-time PCR assays showed that 88% (7/8) of the true positives identified by the statistical test during in silico analysis were also recognized as positives in the real-time PCR assays, indicating that IVV-HiTSeq is highly reliable. Furthermore, 89% (8/9) of the positives recognized by the RT-PCR assays were correctly recognized as positives in the in silico analysis, indicating that IVV-HiTSeq also had high coverage. When the in silico procedure was applied to all the data in the datasets, 110 enriched IRs were identified that overlapped with protein-coding regions in 106 RefSeq genes (the equivalent of 106 protein-protein interactions; see Supplementary Table S3 online).
IVV-HiTSeq was compared with conventional method using Sanger sequencing for the same prey library and bait and 640 sequences (87%) determined by Sanger sequencing were also obtained by IVV-HiTSeq; however, most of the sequences (99.7%) obtained by IVV-HiTSeq were new and not found by Sanger sequencing (Fig. 3). Moreover, 88% (7/8) of the real-time PCR assays that were followed by IVV-HiTSeq, including in silico analysis, were positive, while only 43% (9/21) of the randomly chosen samples from the obtained raw reads were positive.
Discussion
This study demonstrated that the IVV-HiTSeq method could produce a large amount of accurate protein interaction data. One of the main reasons for this highly efficient production of interactions data is the completely cell-free experimental procedure. The combination of IVV and high-throughput sequencing technologies does not require any host cells, including bacterial ones, for DNA cloning, which previously limited the efficiency of screening and numbers of interactions that could be examined. The IVV method can conduct a selection with a cDNA library consisting of >1012 molecules, which is beyond the capacity of conventional high-throughput protein selection methods9,10. With the further increase in throughput expected of NGS in the future, the coverage of the interactome is expected to increase. Notably, our completely cell-free procedure will also be effective for analyzing cytotoxic proteins, leading to a more comprehensive interactome analysis.
With respect to the accuracy of the IVV-HiTSeq data, the use of library-specific barcoded primers and in silico analysis reduced the number of false positive interactions contained in the initial raw data. Considering the comparison with the real-time PCR assays, IVV-HiTSeq could provide verification data comparable to real-time PCR assays. Thus, IVV-HiTSeq generates data equivalent to several thousands of real-time PCR confirmations, resulting in reduced cost and time. IVV-HiTSeq also has the ability to reproduce data and to reduce false negatives compared with the conventional method using Sanger sequencing. The most important point, however, is that the method dramatically reduces false positives. Researchers in the fields of cellular biology and physiology will have more confidence in the interaction data than they have for data obtained using conventional methods. Nonetheless, further in vivo validations are required, because the results presented here were obtained under cell-free conditions.
In the present study, IVV-HiTSeq used the 454 sequencer; thus, the maximum read lengths of the sequences generated by NGSs1 are long enough for the IVV-HiTSeq. Alternatively, the length problem might be solved by pair-end tag sequencing11. IVV-HiTSeq is potentially applicable to many cell-free display technologies, for example, mRNA display, DNA display and ribosome display. Moreover, IVV can be applied not only to the in vitro selection of protein-protein interactions, but also to the detection of protein-DNA, protein-RNA and protein-chemical compound interactions12, suggesting that IVV-HiTSeq could become a universal tool for exploring the large space of protein sequences and interaction networks.
Methods
Preparation of the prey mRNA library
An RNA library for mRNA display was created from the poly(A)+ mRNAs extracted from the spleens of 6- to 8-week-old C57BL/6 mice, according to a previously described method7,13. First, randomly primed reverse transcription (RT) of poly(A)+ mRNAs were subjected to ligation-mediated amplification14 and multi-step PCRs to create cDNA constructs for in vitro expression. The resulting PCR products (SP6-Ω-T7-Flagment-Kpn1-FLAG-A{8}) were purified with a QIAquick PCR Purification Kit (Qiagen, Germany) and transcribed into mRNA with a RiboMAX Large Scale RNA Production System-SP6 (Promega, WI, USA) and an m7G(5′)ppp(5′)G RNA Cap Structure Analog (Ambion, Life Technologies, CA, USA). After purification of the transcribed mRNAs using an RNeasy 96 BioRobot 8000 Kit (Qiagen), PEG Puro spacer was ligated to the 3′ends of mRNAs using T4 RNA ligase (Promega) and the RNA was purified again.
Preparation of bait
A cDNA for the bait (Irf7) was prepared according to the previously reported method7,13. The structure of the cDNA construct, SP6-(O')-T7-Irf7-CBP-zz-His, created through multi-step PCR, includes the full-length coding region of the mouse interferon regulatory factor 7 (Irf7; NM_016850.2), which was used as the bait. The PCR products, purified by a QIAquick PCR Purification Kit (Qiagen), were transcribed into mRNA with a RiboMAX Large Scale RNA Production System-SP6 (Promega) and m7G(5′)ppp(5′)G RNA Cap Structure Analog (Ambion, Life technologies) and then purified with the RNeasy 96 BioRobot 8000 Kit (Qiagen).
In vitro translation and selection using IVV
The prey library and Irf7 mRNA were co-translated using Wheat Germ Extract (Promega) as a cell-free translation system. At the same time, the IVV molecules were formed by covalently attaching the 3′end of mRNA for prey to the C-terminus of its coding protein via puromycin15.
One- or two-step purifications16 using tagged bait protein were performed as one round of selection with unpurified, cell-free co-translation products. After each round of selection, prey mRNA was amplified by RT-PCR, followed by the in vitro transcription and translation reactions that prepared the library for the next round of selection. As a negative control, the experiment was performed under the same conditions except that the bait protein was absent.
Sample preparation for 454 sequencing
To distinguish derivations of prey sequences after sequencing on the 454 GS FLX system (Roche, Switzerland)17, mRNAs libraries at the pre-1st and post- 1st, 2nd, 3rd and 4th selection rounds were amplified with barcoded primers that had the following round-specific 4-base regions:
ACTA+(T7_5′end_forward: TGCGGCCGCGAATTCC) and TAGT+(FLAG_3′end_reverse: GTCGTCATCGTCCTTGTAGTC); AGTG+(T7_5′end_forward: TGCGGCCGCGAATTCC) and CACT+(FLAG_3′end_reverse: GTCGTCATCGTCCTTGTAGTC); CAGC+(T7_5′end_forward: TGCGGCCGCGAATTCC) and GCTG+(FLAG_3′end_reverse: GTCGTCATCGTCCTTGTAGTCA); CGCA+(T7_5′end_forward: TGCGGCCGCGAATTCC) and TGCG+(FLAG_3′end_reverse: GTCGTCATCGTCCTTGTAGTCA); and CTCG+(T7_5′end_forward: TGCGGCCGCGAATTCC) and CGAG+(FLAG_3′end_reverse: GTCGTCATCGTCCTTGTAGTCA), for samples from the initial and post 1st to 4th selection rounds, respectively.
DNA samples amplified by barcoded primers were purified using a QIAquick PCR Purification Kit (Qiagen). After purification, the DNA concentrations for each sample were measured by NanoDrop. Finally, the 1st to 4th selection round samples were mixed in a weight ratio of 10:7:2:1. The negative control samples were produced using the same method. Finally, the initial library sample and the two combined samples were applied to the GS FLX 454 sequencer (Roche).
Real-time PCR assays
Real-time PCR assays were performed using a 7300 Real-Time PCR System (Applied Biosystems), according to the manufacturer's instruction. A total of 25 μl of the reaction mixture, consisting of 5 ng of DNA template from the prey library, 0.5 μl of 10 μM primers for each forward and reverse strand and 12.5 μl of Power SYBR Green PCR Master Mix (Applied Biosystems), was used for each round of selection. Synthetic primers that specifically amplify the target sequences are listed in Supplementary Table S2. Measurements for each sample were made twice and the values were averaged.
In silico analysis
The round specific 4-base barcoded ends of the cDNA sequences were first decoded to identify which round each read was derived from. The constant regions that were used for amplification were then masked before mapping to the masked mouse genome (mm8 http://hgdownload.cse.ucsc.edu/downloads.html#mouse) using BLAT18 with the following cut-off conditions: match length ≥ 30 bp and identity ≥ 95%. Frequencies of mapped reads for each round of selections were calculated for each nucleotide position in the genome and compared with frequencies of reads from the initial library and the negative control of the corresponding round. Positions that had frequencies higher than those for the same positions in both the initial library and negative control samples were subjected to statistical tests. To calculate the statistical significance of the differences in frequencies, Fisher's exact tests were conducted using R software (R: A Language and Environment for Statistical Computing; http://www.R-project.org/). When statistical significance (P < 0.001) was confirmed in the comparisons with both the initial library and the corresponding negative control, these positions were determined as enriched positions. Consecutive positions that showed statistically significant enrichment were merged and regarded as an interaction region.
References
Zhou, X. et al. The next-generation sequencing technology and application. Protein Cell 1, 520–536 (2010).
Venkatesan, K. et al. An empirical framework for binary interactome mapping. Nat Methods 6, 83–90 (2009).
Yu, H. et al. High-quality binary protein interaction map of the yeast interactome network. Science 322, 104–110 (2008).
Yu, H. et al. Next-generation sequencing to generate interactome datasets. Nature methods 8, 478–480 (2011).
Rual, J. F. et al. Towards a proteome-scale map of the human protein-protein interaction network. Nature 437, 1173–1178 (2005).
Gentleman, R. & Huber, W. Making the most of high-throughput protein-interaction data. Genome Biol 8, 112 (2007).
Miyamoto-Sato, E. et al. Cell-free cotranslation and selection using in vitro virus for high-throughput analysis of protein-protein interactions and complexes. Genome Res 15, 710–717 (2005).
Miyamoto-Sato, E. et al. A comprehensive resource of interacting protein regions for refining human transcription factor networks. PLoS One 5, e9289 (2010).
Roberts, R. W. Totally in vitro protein selection using mRNA-protein fusions and ribosome display. Current opinion in chemical biology 3, 268–273 (1999).
Wang, H. & Liu, R. Advantages of mRNA display selections over other selection techniques for investigation of protein-protein interactions. Expert review of proteomics 8, 335–346 (2011).
Fullwood, M. J., Wei, C.-L., Liu, E. T. & Ruan, Y. Next-generation DNA sequencing of paired-end tags (PET) for transcriptome and genome analyses. Genome research 19, 521–532 (2009).
Takahashi, T. T., Austin, R. J. & Roberts, R. W. mRNA display: ligand discovery, interaction analysis and beyond. Trends Biochem Sci 28, 159–165 (2003).
Miyamoto-Sato, E. et al. Highly stable and efficient mRNA templates for mRNA–protein fusions and C-terminally labeled proteins. Nucleic Acids Research 31, e78 (2003).
Chenchik, A. et al. Full-length cDNA cloning and determination of mRNA 5′ and 3′ ends by amplification of adaptor-ligated cDNA. BioTechniques 21, 526–534 (1996).
Nemoto, N., Miyamoto-Sato, E., Husimi, Y. & Yanagawa, H. In vitro virus: bonding of mRNA bearing puromycin at the 3′-terminal end to the C-terminal end of its encoded protein on the ribosome in vitro. FEBS Lett 414, 405–408 (1997).
Rigaut, G. et al. A generic protein purification method for protein complex characterization and proteome exploration. Nature 17, 1030–1032 (1999).
Margulies, M. et al. Genome sequencing in microfabricated high-density picolitre reactors. Nature 437, 376–380 (2005).
Kent, W. J. BLAT--the BLAST-like alignment tool. Genome Res 12, 656–664 (2002).
Acknowledgements
We thank Dr. Masamichi Ishizaka and Dr. Hiroshi Yanagawa for helpful discussions and Katsuhito Hashizume and Akiyuki Sato of Takara Bio Inc. for their technical support in preparing samples for 454 sequencing. This research was partially supported by a Grant-in-Aid for Scientific Research on Innovative Areas “Integrative Systems Understanding of Cancer for Advanced Diagnosis, Therapy and Prevention (No. 4201)” (Grant No. 23134510) of The Ministry of Education, Culture, Sports, Science and Technology (MEXT), Japan (to S.F.); and a Female Researcher Science Grant from Shiseido Co., Ltd (to E.M.S.).
Author information
Authors and Affiliations
Contributions
S.F. and E.M.S. designed the experiments, developed the method and prepared the manuscript. N.H. and K.M. performed the experiments. S.F., T.O. and T.Y. performed the in silico analysis. A.N. and Y.F. provided the irf7 cDNA and total RNA library of mouse spleen. H.O. and T.W. gave technical advice on high-throughput sequencing and in silico analysis. E.M.S. supervised the project.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Supplementary Information
Next-generation sequencing coupled with a cell-free display technology for high-throughput production of reliable interactome data
Supplementary Information
Supplementary Table S3
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-ShareALike 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/3.0/
About this article
Cite this article
Fujimori, S., Hirai, N., Ohashi, H. et al. Next-generation sequencing coupled with a cell-free display technology for high-throughput production of reliable interactome data. Sci Rep 2, 691 (2012). https://doi.org/10.1038/srep00691
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep00691
This article is cited by
-
Multiplex single-molecule interaction profiling of DNA-barcoded proteins
Nature (2014)
-
Liposome display for in vitro selection and evolution of membrane proteins
Nature Protocols (2014)
-
Zellfreie Proteinexpression für Forschung und Produktion
BIOspektrum (2014)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.