Abstract
In the past decade, metagenomics studies have become widespread due to the arrival of second-generation sequencing platforms characterized by low costs, high throughput and short read lengths. Today, although benchtop sequencers are considered to be accurate platforms to deliver data for targeted metagenomics studies, the limiting factor has become the analysis of these data. In a previous paper, we performed an Ion Torrent PGM 16S rDNA gene sequencing of faecal DNAs from 48 Blastocystis-colonized patients and 48 Blastocystis-negative subjects, in order to decipher the impact of this widespread protist on gut microbiota composition and diversity. We report here on the Ion Torrent targeted metagenomic sequencing and analysis of these 96 human faecal samples, and the complete datasets from raw to analysed data. We also provide the key steps of the bioinformatic analyses, from library preparation to data filtering and OTUs tables generation. This data represents a valuable resource for the scientific community, enabling re-processing of these targeted metagenomic datasets through various pipelines and a comparative evaluation of microbiota analysis methods.
Design Type(s) | parallel group design • observation design • metagenomics analysis |
Measurement Type(s) | rRNA_16S |
Technology Type(s) | DNA sequencing |
Factor Type(s) | Clinical Diagnosis • Risk Factor Assessment |
Sample Characteristic(s) | Homo sapiens • feces |
Machine-accessible metadata file describing the reported data (ISA-Tab format)
Similar content being viewed by others
Background & Summary
In recent years, the advent of high-throughput sequencing (HTS)-based metagenomics and the development of related bioinformatics approaches have greatly facilitated the study of microbial communities in humans. For example, it has allowed deeper insight into the critical role played by the community of microorganisms inhabiting our gastrointestinal tract in determining human health and disease. From a technological viewpoint, shotgun metagenomics still remains very expensive, and data analysis remains a challenging issue, due to both the size and complex structure of the data1. Thanks to the recent benchtop sequencer releases (Ion Torrent and MiSeq technologies), targeted metagenomics that focuses only on an informative genomic marker like 16S rDNA, has become widespread in microbiology studies, and is commonly used today to assess the gut microbiota.
Besides bacteria, which are the most numerous organisms, diverse eukaryotes, such as protists, also inhabit the human gut. Among all the human intestinal protists, some are difficult to define unequivocally as pathogens or commensals2. Indeed, pathogenicity is undoubtedly the rule for Cryptosporidium spp. and Entamoeba histolytica, but the clinical significance of other intestinal protists like Blastocystis spp. remains undetermined2,3.
Since their discovery, Blastocystis spp., reported to be the most common unicellular eukaryotes detected in human faecal samples, have generally been investigated under conditions in which they were assumed to cause disease. However, they are also detected with high prevalence in healthy people, and it has been shown that they can colonize the healthy human gut for long periods of time without resulting in symptomatic carrier status4. In order to address the issue of the clinical significance of these intestinal protists, studying the impact of their colonization on gut microbiota began to attract interest from the scientific community3, 5,6,7.
In a previous paper, we investigated the impact of Blastocystis colonization on the composition and diversity of the human gut microbiota by comparing the microbiota of Blastocystis-colonized patients and Blastocystis-negative individuals3. This study suggested that colonization by this protist is usually associated with a healthy gut microbiota, rather than with the gut dysbiosis generally observed in metabolic or infectious inflammatory diseases of the lower gastrointestinal tract3. This result was obtained from a cross-sectional study including two groups of 48 subjects, colonized or not by Blastocystis, and comparable in terms of clinical and environmental variables. Ion Torrent 16S rDNA gene sequencing of faecal DNAs was performed, followed by bioinformatics analyses to profile and compare the gut bacterial communities of these two groups.
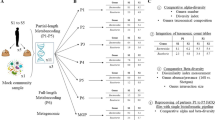
We here provide detailed descriptions of DNA sample processing, including DNA extraction, quantification and monitoring, sequencing library creation, and sequencing procedures. Moreover, we report the Ion Torrent raw sequencing data (Data Citation 1), and provide a full description of the bioinformatics pipeline we used for the analysis (Fig. 1). The datasets provided here represent a valuable resource for the scientific community, enabling the re-processing of the data and a comparative evaluation of other targeted metagenomic analysis pipelines. They include: (i) the raw sequencing data (Data Citation 1) (ii) the intermediate files from our home-made 16S targeted metagenomic analysis pipeline (i.e., the OTU count tables (OTU_count_tables tsv files, Data Citation 2) and the BIOM file before data normalization (raw_global BIOM file, Data Citation 2), and (iii) the final BIOM file (normalized_global BIOM file, Data Citation 2). The intermediate files allow the scientific community to monitor, evaluate and/or optimize certain analytical steps, whereas the final BIOM file (normalized_global BIOM file, Data Citation 2) allows the confirmation of our biological results, particularly the higher bacterial diversity in faecal microbiota of Blastocystis colonized patients3, as well as a comparison of the available tools for secondary analysis of targeted metagenomic data. Finally, the release of all these datasets will be useful for the scientific community to compare microbial patterns for a variety of case-control studies.
Schematic representation of the home-made bioinformatics pipeline for the analysis of targeted metagenomic Ion Torrent sequencing data, that included several publicly available tools (e.g. Mothur10, EspritTree12, QIIME17 or DESeq216), databases (the Silva small subunit RNA database11 and Ribosomal Database Project (RDP)13) and home-made Perl/Python scripts.
Methods
The patient sampling, Blastocystis diagnosis by real-time PCR and additional methods presented here are expanded from the descriptions in our previous paper3.
Briefly, in order to avoid selection bias in the choice of samples, statistical analyses were performed on patients’ clinical and environmental variables to obtain a ranked Blastocystis colonization risk score for each patient. The risk scores were then used to select the 24 highest score values and the 24 lowest scores in each population of Blastocystis-positive and Blastocystis-negative patients, thus determining 4 groups of 24 patients. Groups 1 and 2 were composed of Blastocystis-colonized patients and groups 3 and 4 included Blastocystis-negative patients. Groups 1 and 3 were linked in terms of low environmental and clinical risk factors, and groups 2 and 4 shared high risk factors (see Supplementary File 1).
DNA extraction
For each subject, approx. 100 mg of fresh stools was collected and homogenized by shaking in 1.5 ml of Stool Transport and Recovery (S.T.A.R.) buffer (Roche Diagnostics, Indianapolis, IN) (ratio 1:3 according to the manufacturer’s recommendations). The stool samples stored in the S.T.A.R. buffer were centrifuged for 1 min at 1,000×g. Total genomic DNA was extracted from 200 μl of the supernatant using the QIAamp DNA Stool Mini Kit (Qiagen, Hilden, Germany) according to the manufacturer’s recommended procedure. The DNA was eluted in 100 μl of AE elution buffer (Qiagen) and stored at −20 °C until use.
DNA extraction monitoring
Faecal DNA samples corresponding to 48 Blastocystis-positive and 48 Blastocystis-negative subjects were randomly distributed in a 96-well microplate. The total DNA concentration of each DNA extract was monitored using the Quant-iT PicoGreen dsDNA assay (Invitrogen). The 16S gene DNA copy number was assessed by a SybrGreen quantitative PCR method adapted from Maeda et al.,8 allowing both inhibition effect estimation and DNA concentration adjustment. The reaction mixture (15 μl) for the SybrGreen assay performed in RotorGene (Corbett Life Science) contained 2X Brilliant III SybrGreen qPCR Mastermix (Stratagene), primers ( GTGSTGCAYGGYTGTCGTCA, Univ16S_1048-1067 as the forward primer and ACGTCRTCCMCACCTTCCTC, Univ16S_1175_1194 as the reverse primer) with a final concentration of 560 nM and 2 μl of DNA extract as the template. The amplification conditions were 3 min at 94 °C, 45 cycles of 15 s at 94 °C for denaturation, 22 s at 60 °C for annealing and extension, followed by a melting curve from 54 °C to 95 °C with 0.5 °C increments (10 s).
HTS library preparation
The sequence regions of the 16S rRNA gene-spanning variable regions V3-V5 were then amplified with a fusion PCR using the broad-range forward primer For16S_519, CAGCMGCCGCGGTAATAC and the reverse primer Rev16S_926, CCGTCAATTCMTTTGAGTTT (see Supplementary File 2).
Library preparations for amplicon sequencing were performed in a final volume of 100 μl containing 1X PCR buffer, 2 mM MgSO4, 1 U of DNA High-Fidelity Taq Polymerase (Invitrogen), 625 nM of each barcoded primer (IDT), 250 μM of each dNTP (Invitrogen), and a concentration-adjusted DNA sample. Each sample was taken following two PCRs, one with the sequencing adapter linked to the forward primer, and the other with the sequencing adapter linked to the reverse primer. The two resulting PCR products were equimolarly pooled after DNA purification with NucleoFast 96 PCR (Macherey Nagel), followed by a Quant-iT PicoGreen ds DNA quantification (Invitrogen).
HTS template preparation and sequencing
Before emulsion PCR (emPCR), the library concentration was adjusted to 26 pM. Template preparation resulted from an emPCR in Ion 400 Template OT2 following a purification with Ion Torrent OneTouch ES according to the manufacturer's protocol.
The 96-barcoded bidirectional library was sequenced through PGM, Ion Torrent (Life Technologies) with the Ion 318 Chip and Ion PGM 400 Sequencing Kit (Life Technologies), according to the recommended protocol.
Sequence-based microbiota analysis
The analysis of all samples was performed using a home-made Galaxy (v1.0.0)9 pipeline (http://www.pegase-biosciences.com/pub_2014/#ECCB), as shown in Fig. 1, linking three main analytical steps: raw data preprocessing, clustering analysis and OTU classification, and read count normalization. Unless otherwise mentioned in Fig. 1, default parameters were used for all software programs. The home-made scripts are available in Figshare (scripts.zip file, Data Citation 2).
The preprocessing step (Fig. 1), using Mothur (v1.27.0)10 and home-made scripts (scripts.zip file, Data Citation 2), filtered the raw data (Data Citation 1) to minimize erroneous reads generated by the Ion Torrent PGM sequencer. Reads shorter than 150 bases and/or containing large homopolymers were removed. The reads were then aligned against the SILVA 102 bacterial database11, and those reads with alignments of fewer than 100 bases were filtered out. Finally, the filtered reads were deduplicated to reduce the datasets. Three of the 96 samples, considered to be outliers (see the Technical validation section) were discarded in this step, before proceeding with the subsequent analyses. The number of reads remaining after this preprocessing step was 2,742,108.
In the second analytical step, OTU clustering was performed using ESPRIT-Tree12 version 11152011, which allows the same OTU definition precision as standard hierarchical clustering procedures, but requires less execution time. The OTUs were classified using classify.seqs in Mothur (v1.27.0)10 with the SILVA 102 database11 and the RDP taxonomy13 (Fig. 1).
Intra-sample rarefaction curves were generated using the home-made rarefaction curve plotting tool (rarefaction.R in scripts.zip file, Data Citation 2). It provides a way of comparing the richness observed in the samples. Graphically it presents the number of OTUs theoretically observed for a range number of sequences into the sample at variable distances (91%, 93%, 95%, 97%).
For each sample, the output of this second analytical step is an OTU table file (OTU_count_tables tsv file, Data Citation 2) containing four columns: the first column is the consensus read name associated to the OTU, the second column is the OTU raw counts, the third column is the consensus read name (same as the first column) and the fourth column is the associated taxon. The consensus sequence for a given OTU is the most abundant sequence in this OTU. The characteristics of these OTU_count_tables files are summarized in Table 1.
In the third analytical step of the home-made pipeline (Fig. 1), all the annotated OTU tables were merged into a global OTU table using a home-made python script (v2.7.3) (OTU_tables_merge.py script from scripts.zip file, Data Citation 2), based on each OTU's taxonomic annotation.
The Global OTU table is a tabulation-formatted file (TSV) in which each column of this table represents one sample, and each line represents one taxon (identified by its OTU identifier in the first column and by the taxonomic annotation in the last column). This merged OTU table describes 474 OTUs, their annotation, and the number of reads belonging to each OTU per sample. Finally, this annotated OTU table was converted into a global BIOM file14 by the biom (v2.1.4) convert command. HDF5 format was chosen to optimize the storage. The characteristics of the BIOM file (Data Citation 2) are summarized in Table 2.
As advised by previous recommendations15, the DESeq2 package16 integrated into QIIME (v1.9.0)17, was used to normalize the total read counts and avoid rarefaction of the read count data. The normalization_table.py python script from QIIME (v1.9.0) was configured with the following options: the algorithm chosen was DESeq2, replacing negative numbers produced by the DESeq normalization technique with zeros. A normalized Global BIOM file was then produced (Table 3). Note that the taxonomic information disappears after this normalization step. The biom (v2.1.4) 'add-metadata' option was used to add the taxonomic information back and obtain a fully annotated and normalized BIOM file (Data Citation 2).
Ethical approval
Ethical approval surrounding the procurement of samples used in this study was obtained from the Research Ethics Committee ‘Comité de Protection des Personnes Sud-Est 6, France’ (reference number 2015/CE82), which waived the requirement for informed consent because the experiments did not result in additional constraints for the patients. All the methods were carried out in accordance with the approved guidelines (World Medical Association’s (WMA) Declaration of Helsinki-Ethical Principles for Medical Research Involving Human Subjects).
Data Records
The raw data from each of the 96 sequenced samples were submitted to the Sequence Read Archive (SRA) of the NCBI, under Project ID PRJNA342805 (Data Citation 1). The raw data were deposited under each barcode index. The association between the sample name, sample ID (index), and Blastocystis colonization status and group is available in Supplementary File 1.
The data analyzed using the home-made metagenomics pipeline (Fig. 1) are available on the PEGASE-biosciences website: http://www.pegase-biosciences.com/collaborations/blastocystis-gut-microbiota/.
The taxonomic assignment before data normalization is available per sample in the OTU_count_tables tsv files (Data Citation 2) and the raw_global.biom file (Data Citation 2); and after data normalization (process described in the Methods section) in the normalized_global.biom file (Data Citation 2).
Finally, the four home-made scripts used in the bioinformatics pipeline (Fig. 1) are available in the scripts.zip file (Data Citation 2).
Technical Validation
Data quality check
As shown in Fig. 2, the quality scores of the raw Ion Torrent reads (average Q27) are quite stable across the reads, above the Q20 reported accuracy for Ion Torrent PGM sequencing. Even the ends of the reads, which are known to be of lesser quality, are still within the Q20 range. This allowed the use of the complete reads for downstream analyses, without requiring a quality read trimming step.
Raw data output
A total of 3,962,103 reads were obtained from the Ion 318 Chip sequencing run of the 96 indexed samples. As expected, a length mode of approximately 400 bases (403 bases) was observed, with an average read number per index of 41,562 and an average read length of 272 bases. These metrics assessed the good quality/quantity of the generated raw data (Table 4).
Three of the 96 samples were discarded after sequencing (see Supplementary File 1). Indexes 50 and 63 were considered to be outliers due to the low number of reads (<10,000) for these samples. A third outlier (index 18) was due to an accidental handling error and was also discarded from the data set.
ANOVA tests were conducted to compare the read number per index and the read lengths from the output sequence data in the four groups of patients. There were no significant differences between the four groups for any variable, indicating that there was no technical bias in sequencing for the four groups of patients (mean read length per group (P-value=0.47) and mean read number per group (P-value=0.78)) (Table 5).
Additional Information
How to cite this article: Siegwald, L. et al. Targeted metagenomic sequencing data of human gut microbiota associated with Blastocystis colonization. Sci. Data 4:170081 doi: 10.1038/sdata.2017.81 (2017).
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
References
References
Lindgreen, S., Adair, K. L. & Gardner, P. An evaluation of the accuracy and speed of metagenome analysis tools. Sci. Rep 6, 19233 (2015).
Lukeš, J., Stensvold, C. R., Jirků-Pomajbíková, K. & Wegener Parfrey, L. Are human intestinal eukaryotes beneficial or commensals? PLoS Pathog. 11, e1005039 (2015).
Audebert, C. et al. Colonization with the enteric protozoa Blastocystis is associated with increased diversity of human gut bacterial microbiota. Sci. Rep. 6, 25255 (2016).
Scanlan, P. D. et al. The microbial eukaryote Blastocystis is a prevalent and diverse member of the healthy human gut microbiota. FEMS Microbiol. Ecol. 90, 326–330 (2014).
Andersen, L. O., Bonde, I., Nielsen, H. B. & Stensvold, C. R. A retrospective metagenomics approach to studying Blastocystis. FEMS Microbiol. Ecol. 91 https://doi.org/10.1093/femsec/fiv072 (2015).
Iebba, V. et al. Gut microbiota related to Giardia duodenalis, Entamoeba spp. and Blastocystis hominis infections in humans from Côte d'Ivoire. J. Infect. Dev. Ctries 10, 1035–1041 (2016).
Nagel, R., Traub, R. J., Allcock, R. J., Kwan, M. M. & Bielefeldt-Ohmann, H. Comparison of faecal microbiota in Blastocystis-positive and Blastocystis-negative irritable bowel syndrome patients. Microbiome 4, 47 (2016).
Maeda, H. et al. Quantitative real-time PCR using TaqMan and SYBR Green for Actinobacillus actinomycetemcomitans, Porphyromonas gingivalis, Prevotella intermedia, tetQ gene and total bacteria. FEMS Immunol. Med. Microbiol. 39, 81–86 (2003).
Goecks, J., Nekrutenko, A. & Taylor, J. Galaxy: a comprehensive approach for supporting accessible, reproducible, and transparent computational research in the life sciences. Genome Biol. 11, R86 (2010).
Schloss, P. D. et al. Introducing mothur: Open-source, platform-independent, community-supported software for describing and comparing microbial communities. Appl. Environ. Microbiol. 75, 7537–7541 (2009).
Quast, C. et al. The SILVA ribosomal RNA gene database project: improved data processing and web-based tools. Nucleic Acids Res. 41, D590–D596 (2013).
Cai, Y. & Sun, Y. ESPRIT-Tree: Hierarchical clustering analysis of millions of 16S rRNA pyrosequences in quasilinear computational time. Nucleic Acids Res. 39, e95 (2011).
Wang, Q., Garrity, G. M., Tiedje, J. M. & Cole, J. R. Naïve Bayesian classifier for rapid assignment of rRNA sequences into the new bacterial taxonomy. Appl. Environ. Microbiol. 73, 5261–5267 (2007).
McDonald, D. et al. The Biological Observation Matrix (BIOM) format or: how I learned to stop worrying and love the ome-ome. Gigascience 1, 7 (2012).
McMurdie, P. J. & Holmes, S. Waste not, want not: why rarefying microbiome data is inadmissible. PLoS Comput. Biol. 10, e1003531 (2014).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-Seq data with DESeq2. Genome Biol. 15, 550 (2014).
Caporaso, J. G. et al. QIIME allows analysis of high-throughput community sequencing data. Nat. methods 7, 335–336 (2010).
Data Citations
NCBI Sequence Read Archive PRJNA342805 https://www.ncbi.nlm.nih.gov/bioproject/?term=PRJNA342805 (2016)
Siegwald, L. Figshare https://doi.org/10.6084/m9.figshare.c.3751037 (2017)
Acknowledgements
This work was supported by grants from the Programmes Orientations Stratégiques of the University de Lille 2 and the Centre National de la Recherche Scientifique and the Institut Pasteur de Lille. This work was also supported by CIFRE grant no. 2013/0920 from the Association Nationale de la Recherche et de la Technologie to L.S.
Author information
Authors and Affiliations
Contributions
All the authors collaborated to conceptualize the study. L.S. and G.E. conducted the bioinformatic analyses. S.C. and C.A. deposited the sequencing data in the databases. M.C. and C.A. wrote the paper. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
ISA-Tab metadata
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/ The Creative Commons Public Domain Dedication waiver http://creativecommons.org/publicdomain/zero/1.0/ applies to the metadata files made available in this article.
About this article
Cite this article
Siegwald, L., Audebert, C., Even, G. et al. Targeted metagenomic sequencing data of human gut microbiota associated with Blastocystis colonization. Sci Data 4, 170081 (2017). https://doi.org/10.1038/sdata.2017.81
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/sdata.2017.81
This article is cited by
-
Comparative shotgun metagenomic data of the silkworm Bombyx mori gut microbiome
Scientific Data (2018)
-
Progress of analytical tools and techniques for human gut microbiome research
Journal of Microbiology (2018)