Abstract
Metagenome-based resources have revealed the diversity and function of the human gut microbiome, but further understanding is limited by insufficient genome quality and a lack of samples from typically understudied populations. Here we used hybrid long-read PromethION and short-read HiSeq sequencing to characterize the faecal microbiota of 60 Inner Mongolian individuals (n = 180 samples over three time points) who were part of a probiotic yogurt intervention trial. We present the Inner Mongolian Gut Genome catalogue, comprising 802 closed and 5,927 high-quality metagenome-assembled genomes. This approach achieved high genome continuity and substantially increased the resolution of genomic elements, including ribosomal RNA operons, metabolic gene clusters, prophages and insertion sequences. Particularly, we report the ribosomal RNA operon copy numbers for uncultured species, over 12,000 previously undescribed gut prophages and the distribution of insertion sequence elements across gut bacteria. Overall, these data provide a high-quality, large-scale resource for studying the human gut microbiota.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data Availability
All sequencing data (Illumina and Nanopore) generated in this study and the high-quality genomes in the IMGG dataset can be found under NCBI BioProject PRJNA763692. The 6,729 high-quality IMGGs are available at https://doi.org/10.6084/m9.figshare.19661523. Source data are provided with this paper.
Code availability
The in-house scripts for performing bioinformatics analyses in this work can be found in GitHub at https://github.com/jinhao94/nanopore_script.git and https://github.com/jinhao94/binning_script.git.
References
Fan, Y. & Pedersen, O. Gut microbiota in human metabolic health and disease. Nat. Rev. Microbiol. 19, 55–71 (2021).
Byrd, A. L., Belkaid, Y. & Segre, J. A. The human skin microbiome. Nat. Rev. Microbiol. 16, 143–155 (2018).
Zheng, D. P., Liwinski, T. & Elinav, E. Interaction between microbiota and immunity in health and disease. Cell Res. 30, 492–506 (2020).
Zhang, Z., Wang, J., Wang, J., Wang, J. & Li, Y. Estimate of the sequenced proportion of the global prokaryotic genome. Microbiome 8, 134 (2020).
Almeida, A. et al. A unified catalog of 204,938 reference genomes from the human gut microbiome. Nat. Biotechnol. 39, 105–114 (2021).
Pasolli, E. et al. Extensive unexplored human microbiome diversity revealed by over 150,000 genomes from metagenomes spanning age, geography, and lifestyle. Cell 176, 649–662.e20 (2019).
Nayfach, S., Shi, Z. J., Seshadri, R., Pollard, K. S. & Kyrpides, N. C. New insights from uncultivated genomes of the global human gut microbiome. Nature 568, 505–510 (2019).
Almeida, A. et al. A new genomic blueprint of the human gut microbiota. Nature 568, 499–504 (2019).
Shaiber, A. & Eren, A. M. Composite metagenome-assembled genomes reduce the quality of public genome repositories. mBio 10, e00725-19 (2019).
Bowers, R. M. et al. Minimum information about a single amplified genome (MISAG) and a metagenome-assembled genome (MIMAG) of bacteria and archaea. Nat. Biotechnol. 35, 725–731 (2017).
Nissen, J. N. et al. Improved metagenome binning and assembly using deep variational autoencoders. Nat. Biotechnol. 39, 555–560 (2021).
Moss, E. L., Maghini, D. G. & Bhatt, A. S. Complete, closed bacterial genomes from microbiomes using nanopore sequencing. Nat. Biotechnol. 38, 701–707 (2020).
Bertrand, D. et al. Hybrid metagenomic assembly enables high-resolution analysis of resistance determinants and mobile elements in human microbiomes. Nat. Biotechnol. 37, 937–944 (2019).
Driscoll, C. B., Otten, T. G., Brown, N. M. & Dreher, T. W. Towards long-read metagenomics: complete assembly of three novel genomes from bacteria dependent on a diazotrophic cyanobacterium in a freshwater lake co-culture. Stand. Genom. Sci. 12, 9 (2017).
Chng, K. R. et al. Cartography of opportunistic pathogens and antibiotic resistance genes in a tertiary hospital environment. Nat. Med. 26, 941–951 (2020).
Kolmogorov, M. et al. metaFlye: scalable long-read metagenome assembly using repeat graphs. Nat. Methods 17, 1103–1110 (2020).
Waschulin, V. et al. Biosynthetic potential of uncultured Antarctic soil bacteria revealed through long-read metagenomic sequencing. ISME J. 16, 101–111 (2022).
Li, Y. et al. Recovery of human gut microbiota genomes with third-generation sequencing. Cell Death Dis. 12, 569 (2021).
Bishara, A. et al. High-quality genome sequences of uncultured microbes by assembly of read clouds. Nat. Biotechnol. 36, 1067–1075 (2018).
Bickhart, D. M. et al. Generating lineage-resolved, complete metagenome-assembled genomes from complex microbial communities. Nat. Biotechnol. 40, 711–719 (2022).
Beaulaurier, J. et al. Metagenomic binning and association of plasmids with bacterial host genomes using DNA methylation. Nat. Biotechnol. 36, 61–69 (2018).
Watson, M. & Warr, A. Errors in long-read assemblies can critically affect protein prediction. Nat. Biotechnol. 37, 124–126 (2019).
Kang, D. D. et al. MetaBAT 2: an adaptive binning algorithm for robust and efficient genome reconstruction from metagenome assemblies. PeerJ 7, e7359 (2019).
Louca, S., Doebeli, M. & Parfrey, L. W. Correcting for 16S rRNA gene copy numbers in microbiome surveys remains an unsolved problem. Microbiome 6, 41 (2018).
Roux, S. et al. Minimum Information about an Uncultivated Virus Genome (MIUViG). Nat. Biotechnol. 37, 29–37 (2019).
Nayfach, S. et al. Metagenomic compendium of 189,680 DNA viruses from the human gut microbiome. Nat. Microbiol. 6, 960–970 (2021).
Debroas, D. & Siguret, C. Viruses as key reservoirs of antibiotic resistance genes in the environment. ISME J. 13, 2856–2867 (2019).
Siguier, P., Gourbeyre, E. & Chandler, M. Bacterial insertion sequences: their genomic impact and diversity. FEMS Microbiol. Rev. 38, 865–891 (2014).
Consuegra, J. et al. Insertion-sequence-mediated mutations both promote and constrain evolvability during a long-term experiment with bacteria. Nat. Commun. 12, 980 (2021).
Stoddard, S. F., Smith, B. J., Hein, R., Roller, B. R. & Schmidt, T. M. rrn DB: improved tools for interpreting rRNA gene abundance in bacteria and archaea and a new foundation for future development. Nucleic Acids Res. 43, D593–D598 (2015).
Degnan, P. H., Taga, M. E. & Goodman, A. L. Vitamin B12 as a modulator of gut microbial ecology. Cell Metab. 20, 769–778 (2014).
Bhattacharya, T., Ghosh, T. S. & Mande, S. S. Global profiling of carbohydrate active enzymes in human gut microbiome. PLoS ONE 10, e0142038 (2015).
Martínez, J. L., Coque, T. M. & Baquero, F. What is a resistance gene? Ranking risk in resistomes. Nat. Rev. Microbiol. 13, 116–123 (2015).
Carr, V. R. et al. Abundance and diversity of resistomes differ between healthy human oral cavities and gut. Nat. Commun. 11, 693 (2020).
Durrant, M. G., Li, M. M., Siranosian, B. A., Montgomery, S. B. & Bhatt, A. S. A bioinformatic analysis of integrative mobile genetic elements highlights their role in bacterial adaptation. Cell Host Microbe 27, 140–153.e9 (2020).
Feng, X. W., Cheng, H. Y., Portik, D. & Li, H. Metagenome assembly of high-fidelity long reads with hifiasm-meta. Nat. Methods 19, 671 (2022).
Sereika, M. et al. Oxford Nanopore R10.4 long-read sequencing enables the generation of near-finished bacterial genomes from pure cultures and metagenomes without short-read or reference polishing. Nat. Methods 19, 823 (2022).
De Coster, W., D’Hert, S., Schultz, D. T., Cruts, M. & Van Broeckhoven, C. NanoPack: visualizing and processing long-read sequencing data. Bioinformatics 34, 2666–2669 (2018).
Hu, J., Fan, J., Sun, Z. & Liu, S. NextPolish: a fast and efficient genome polishing tool for long-read assembly. Bioinformatics 36, 2253–2255 (2020).
Parks, D. H., Imelfort, M., Skennerton, C. T., Hugenholtz, P. & Tyson, G. W. CheckM: assessing the quality of microbial genomes recovered from isolates, single cells, and metagenomes. Genome Res. 25, 1043–1055 (2015).
Gurevich, A., Saveliev, V., Vyahhi, N. & Tesler, G. QUAST: quality assessment tool for genome assemblies. Bioinformatics 29, 1072–1075 (2013).
Cantalapiedra, C. P. et al. eggNOG-mapper v2: functional annotation, orthology assignments, and domain prediction at the metagenomic scale. Mol. Biol. Evol. 38, 5825–5829 (2021).
Sieber, C. M. K. et al. Recovery of genomes from metagenomes via a dereplication, aggregation and scoring strategy. Nat. Microbiol. 3, 836–843 (2018).
Olm, M. R., Brown, C. T., Brooks, B. & Banfield, J. F. dRep: a tool for fast and accurate genomic comparisons that enables improved genome recovery from metagenomes through de-replication. ISME J. 11, 2864–2868 (2017).
Stewart, R. D. et al. Compendium of 4,941 rumen metagenome-assembled genomes for rumen microbiome biology and enzyme discovery. Nat. Biotechnol. 37, 953–961 (2019).
Chan, P. P. & Lowe, T. M. tRNAscan-SE: searching for tRNA genes in genomic sequences. Methods Mol. Biol. 1962, 1–14 (2019).
Hyatt, D. et al. Prodigal: prokaryotic gene recognition and translation initiation site identification. BMC Bioinformatics 11, 119 (2010).
Asnicar, F. et al. Precise phylogenetic analysis of microbial isolates and genomes from metagenomes using PhyloPhlAn 3.0. Nat. Commun. 11, 2500 (2020).
Letunic, I. & Bork, P. Interactive Tree Of Life (iTOL) v5: an online tool for phylogenetic tree display and annotation. Nucleic Acids Res. 49, W293–W296 (2021).
Pascal Andreu, V., Roel-Touris, J., Dodd, D., Fischbach, M. A. & Medema, M. H. The gutSMASH web server: automated identification of primary metabolic gene clusters from the gut microbiota. Nucleic Acids Res. 49, W263–W270 (2021).
Navarro-Muñoz, J. C. et al. A computational framework to explore large-scale biosynthetic diversity. Nat. Chem. Biol. 16, 60–68 (2020).
Akhter, S., Aziz, R. K. & Edwards, R. A. PhiSpy: a novel algorithm for finding prophages in bacterial genomes that combines similarity- and composition-based strategies. Nucleic Acids Res. 40, e126 (2012).
Roach, M. J. et al. Philympics 2021: prophage predictions perplex programs [version 2; peer review: 1 approved, 1 approved with reservations]. F1000Research 10, 758 (2022).
Kanehisa, M. & Goto, S. KEGG: Kyoto encyclopedia of genes and genomes. Nucleic Acids Res. 28, 27–30 (2000).
Aramaki, T. et al. KofamKOALA: KEGG ortholog assignment based on profile HMM and adaptive score threshold. Bioinformatics 36, 2251–2252 (2020).
Cantarel, B. L. et al. The Carbohydrate-Active EnZymes database (CAZy): an expert resource for glycogenomics. Nucleic Acids Res. 37, D233–D238 (2009).
Huang, L. et al. dbCAN-seq: a database of carbohydrate-active enzyme (CAZyme) sequence and annotation. Nucleic Acids Res. 46, D516–D521 (2018).
Bateman, A. et al. The Pfam protein families database. Nucleic Acids Res. 32, D138–D141 (2004).
Potter, S. C. et al. HMMER web server: 2018 update. Nucleic Acids Res. 46, W200–W204 (2018).
Xie, Z. & Tang, H. ISEScan: automated identification of insertion sequence elements in prokaryotic genomes. Bioinformatics 33, 3340–3347 (2017).
Acknowledgements
This study was supported by the National Natural Science Foundation of China (grant numbers 31622043 (Z.S.), 31720103911 (H.Z.), 31972083 (L.-Y.K.), 32001711 (Q.H.)), the earmarked fund for China Agriculture Research System (CARS-36, H.Z.), the Inner Mongolia Science and Technology Major Projects (2021ZD0014, Z.S.), and the Natural Science Foundation of Inner Mongolia Autonomous Region (2020ZD12, Z.S.). We thank Jiachao Zhang (Hainan University) and Shenghui Li for their suggestions; all volunteers for their participation; and the Inner Mongolia Tongfang Discovery Tech. Co., Ltd. for providing storage space and computing resources.
Author information
Authors and Affiliations
Contributions
Z.S. and H.Z. conceived and designed the study. Q.H., H.J., F.Z. and Y.L. performed the probiotic intervention trial and experimental work. H.J. and K.Q. performed bioinformatic analyses. H.J., K.Q., T.M. and L.Y. performed statistical analyses. Z.S. and H.Z. supervised all data analysis. H.J. drafted the manuscript. L.-Y.K. reviewed and revised the paper critically. All authors contributed to data interpretation, read and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Microbiology thanks Ami Bhatt and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
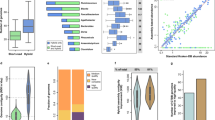
Extended Data Fig. 1 Enhanced accuracy of 16S rRNA gene copy number in Inner Mongolian Gut Genomes.
Comparison of ribosomal RNA operon (rrn) gene copy number between Inner Mongolian Gut Genomes (IMGGs) and their counterpart complete genomes identified in the National Center for Biotechnology Information (NCBI) database.
Extended Data Fig. 2 Functional distribution of complete metabolic gene clusters across the Inner Mongolian Gut Genomes dataset.
Functional distribution of complete metabolic gene clusters across the Inner Mongolian Gut Genomes dataset. The prediction was performed by gutSMASH, which categorized metabolic gene clusters (MGCs) into different gene cluster classes based on their products: [npAA] non-proteinogenic amino acids; [Aromatic] derivatives of benzene; [SCFA-other] a SCFA is produced in combination with another molecule; [Putative] gene clusters of unknown function; [SCFA] fatty acids with 5 carbon atoms maximum; [Other] unclassified pathways; [Aliphatic_amine] ammonia derivatives where at least one H has been replaced by alkyl substituents; [E-MGC] related to energy-capturing mechanisms.
Extended Data Fig. 3 Principal coordinates analysis showing phylum-based clustering trends of metabolic gene clusters.
Permutational analysis of variance (Adonis test; R = 0.38, P < 0.001; n = 15,476) was performed using the adonis function in the vegan package based on the Bray-Curtis distance with 9999 permutations.
Extended Data Fig. 4 Size and frequency of hybrid metabolic gene clusters.
(a) Comparison between the length of hybrid (containing multiple functional domains; n = 11,693) and single-functional-domain (n = 85,654) metabolic gene clusters (MGCs). The boxes represent the interquartile range, the lines inside the boxes represent the medians, and the whiskers denote the lowest and highest values within 1.5 times the interquartile range. (b) The most frequently observed hybrid MGC combination pair. Statistical difference was tested by Wilcoxon rank-sum test (two-sided).
Extended Data Fig. 5 The uneven intra-species distribution of insertion sequence elements.
Distribution of insertion sequence (IS) elements across the 15 most represented metagenome-assembled genomes (MAGs) in the dataset.
Extended Data Fig. 6 The most frequently involved Kyoto Encyclopedia of Genes and Genomes (KEGG) brites and pathways (3rd level) of neighboring genes of insertion sequence elements.
BR and PATH represent Kyoto Encyclopedia of Genes and Genomes (KEGG) brites and pathways, respectively, and codes are not given to components that are ‘not included in pathway or brite’ based on KEGG orthology (KO). The color key shows the 2nd level KEGG pathways, of which the brites and pathways (3rd level) in the horizontal bar chart belong to.
Supplementary information
Supplementary Information
Supplementary Figs. 1 and 2.
Supplementary Table 1
Supplementary Tables 1–15.
Source data
Source Data Fig. 2
Statistical source data.
Source Data Fig. 3
Statistical source data.
Source Data Fig. 4
Statistical source data.
Source Data Fig. 5
Statistical source data.
Source Data Extended Data Fig. 1
Statistical source data.
Source Data Extended Data Fig. 2
Statistical source data.
Source Data Extended Data Fig. 3
Statistical source data.
Source Data Extended Data Fig. 4
Statistical source data.
Source Data Extended Data Fig. 5
Statistical source data.
Source Data Extended Data Fig. 6
Statistical source data.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Jin, H., Quan, K., He, Q. et al. A high-quality genome compendium of the human gut microbiome of Inner Mongolians. Nat Microbiol 8, 150–161 (2023). https://doi.org/10.1038/s41564-022-01270-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41564-022-01270-1
This article is cited by
-
Differences in gut microbiota and its metabolic function among different fasting plasma glucose groups in Mongolian population of China
BMC Microbiology (2023)
-
The multi-kingdom microbiome of the goat gastrointestinal tract
Microbiome (2023)
-
A genome catalog of the early-life human skin microbiome
Genome Biology (2023)