Abstract
Nasopharyngeal swabs were taken from volunteers attending a general medical practice and a general hospital in Lancaster, UK, and at Lancaster University, in the winter of 2014–2015. 51 swabs were selected based on high RNA yield and allocated to deep sequencing pools as follows: patients with chronic obstructive pulmonary disease; asthmatics; adults with no respiratory symptoms; adults with feverish respiratory symptoms; adults with respiratory symptoms and presence of antibodies against influenza C; paediatric patients with respiratory symptoms (2 pools); adults with influenza C infection (2 pools), giving a total of 9 pools. Illumina sequencing was performed, with data yields per pool in the range of 345.6 megabases to 14 gigabases after removal of reads aligning to the human genome. The data were deposited in the Sequence Read Archive at NCBI, and constitute a resource for study of the viral, bacterial and fungal metagenome of the human nasopharynx in healthy and diseased states and comparison with other metagenomic studies on the human respiratory tract.
Design Type(s) | observation design • disease state design • parallel group design |
Measurement Type(s) | metagenomics analysis |
Technology Type(s) | RNA sequencing |
Factor Type(s) | diagnosis • life cycle stage |
Sample Characteristic(s) | nasopharyngeal metagenome |
Machine-accessible metadata file describing the reported data (ISA-Tab format)
Similar content being viewed by others
Background & Summary
Respiratory infectious disease is a major cause of morbidity and mortality in the elderly, infants, the immune-compromised and those with underlying serious pulmonary conditions such as asthma or cystic fibrosis. It is also a source of considerable economic damage through working days lost to illness in adults (ref. 1). Fungi, viruses and bacteria can all cause respiratory infections, but aside from a handful of high profile pathogens such as influenza A virus, Pneumocystis carinii, respiratory syncytial virus and Streptococcus, the problem of respiratory infectious disease diagnosis, treatment and prevention remains neglected.
The sheer diversity of microbes causing respiratory infections that are indistinguishable on clinical examination, means that the vast majority of patients receive only a generic diagnosis of coryza or pharyngitis. Active identification of a causative agent is usually only performed in the event of the illness progressing to severe bronchitis or pneumonia. Despite the highly selective nature of diagnostic results obtained in this way, they offer many important insights. For instance, the WHO tracks the progress of influenza across the world (ref. 2) using percentages of samples testing positive for influenza viruses A/H1N1, A/H3N2 and B. Cumulatively, such diagnostic samples may demonstrate trends in respiratory illness over several years. Nickbakhsh, et al. (ref. 3) tested for 11 virus groups in >60,000 samples collected from >40,000 patients in the west of Scotland from 2005 to 2013, revealing extensive co-infections and a general increase in the respiratory viral load in the population, especially in infants, which remains unexplained.
Deep sequencing, which produces a metagenome reflecting the total microbial genetic content in a sample, offers an alternative to such highly targeted approaches, but is only just beginning to enter the routine diagnostic pipeline (ref. 4–6). Nevertheless, accumulation of human respiratory tract metagenomes from research projects is well underway, providing a resource for the generation of a comprehensive profile of the respiratory microbiome in both health and disease. Within the Sequence Read Archive (SRA), the tag ‘human nasopharyngeal metagenome’ identifies 41 BioProject entries as of 30th August 2017, including that reported here. Within this group of datasets, there are upper respiratory tract metagenomes associated with influenza, respiratory syncytial virus, allergy, intubation, asthma, macular degeneration, oesophageal reflux, rhinosinusitis and Parkinsonism as well as samples taken from healthy volunteers. Our project (ref. 7) initially focussed on the prevalence and severity of influenza C infection in Lancaster, UK, during the winter of 2014–2015. We performed Ilumina deep sequencing on 9 patient pools (Data Citations 1–9), as detailed in Table 1. Two of our 9 sequencing pools (Data Citations 8, 9) were from individuals diagnosed with influenza C infection via quantitative RT-PCR, from one of which we could partially assemble (ref. 7) the genome of a novel strain of influenza C, designated C/Lancaster/1/2015 (Data Citations 10–12). Within other sequencing pools we also detected the complete genomes of novel strains of human rhinovirus A22 (ref. 8), human papillomavirus 23 (ref. 9) and human papillomavirus 20 (ref. 10) (Data Citations 13–15).
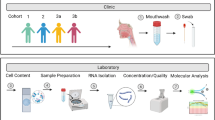
Figure 1 summarises our clinical workflow. From the 148 samples taken, 51 were prioritised for allocation to the 9 sequence pools, based on high yield of RNA and/or clinical phenotype. Our interest in influenza C is reflected in the decision to devote 2 of our 9 sequencing pools to single individuals positive for influenza C by reverse transcription polymerase chain reaction (RT-PCR), one of whom was extensively sequenced (14 Gb). Paediatric coryza was similarly also prioritised (ref. 11) with two pools of 2 and 3 individuals respectively (Data Citations 1, 2). Figure 2 summarises how deep sequencing reads were processed. The raw deep sequencing output was first trimmed of adapter and other extraneous sequences, then groomed to remove reads of low quality score. To avoid the potential ethical complication of sequence from the genomes of the volunteers being present in the sample, three versions of the human genome were used to screen out reads of human origin. It should be noted, however, that we have not attempted to remove any of the known common bacterial contaminants of deep sequencing reactions (ref. 12).
Since we performed deep sequencing on extracted RNA (see Methods), reads are expected to be greatly biased towards expressed regions. Therefore, although we have previously used the data to derive 3 complete short viral genomes (ref. 8–10), detectable genomes of cellular microbes are more likely to be fragmentary. Nevertheless, the dataset may be used for the discovery of further novel viruses, or for comparative studies of the whole virome or bacteriome between the 7 different clinical states defined in the 9 samples. Some of the samples may be compared with other microbiome depositions in the SRA, for instance with BioProject PRJNA310124 (Data Citation 16) on asthma in children (ref. 13). The smaller sequence read sets, for instance SRX2310765 (Data Citation 3) which contains 346 Mb of sequence, are suitable for use as training data for the bioinformatics tools used in metagenomics.
Methods
Ethics
Ethical approval was granted by the UK National Research Ethics Service (NRES), reference 14/LO/1634, Integrated Research Application System (IRAS) Project 147631. The project was registered with the National Institute of Health Research (NIHR), UK, as part of the NIHR Clinical Research Network (UKCRN) Portfolio, ID 17799. All methods were carried out in accordance with the relevant guidelines and regulations. Informed consent was obtained from all volunteers of 18 years and older. For those under 18 years, informed parental consent was obtained, together with supervised assent of the volunteer.
Patient recruitment & sample processing
Participants were approached in 3 locations in Lancaster, UK, (54.05°N 2.80°W) from November 2014 to May 2015: 1) Lancaster University, 2) a general medical practice, 3) hospital clinics. After informed consent was given, patients were allocated to a clinical category (Fig. 1). Nasopharyngeal swabs (MW951SENT, Medical Wire) were brushed over the rear wall of the nasopharynx of patients, and the tips then snapped off directly into Sigma Virocult medium.
RNA was extracted from the nasopharyngeal swabs using a MagMAX Viral RNA Isolation Kit (Ambion). cDNA was prepared using a High-Capacity RNA-to-cDNA Kit (Applied Biosystems, Life Technologies) and a Veriti Thermal Cycler (Applied Biosystems, Life Technologies). The samples were incubated at 37 °C for 60 min, before stopping the reaction at 95 °C for 5 min and then holding at 4 °C. Storage of completed reactions was at −20 °C. After selection of a subset of samples (Fig. 1), deep sequencing was performed using an Illumina Nextera XT library kit in 2×126 bp format on the Illumina HiSeq2500 system.
Processing of raw deep sequencing data for quality control and to remove reads of human genome origin
Quality of Illumina deep sequencing reads was assessed using FastQC (see Technical Validation). Reads were then trimmed of adapters and other non-genomic elements using CutAdapt 1.1 (ref. 14) (https://pypi.python.org/pypi/cutadapt), fastq-mcf 0.11.3 (ref. 15) (https://expressionanalysis.github.io/ea-utils), and trim_galore (http://www.bioinformatics.babraham.ac.uk/projects/trim_galore/), within the Read_cleaner pipeline (Gatherer, unpublished, see Code Availability). Reads of less than 25 bases after trimming were discarded.
Read_cleaner implements the following commands:
-
fastx_clipper -a ADAPTOR_SEQ -l 25 -n -v -M 5 -i INPUT_FILE -o TEMP_FILE
-
trim_galore --length 25 --adapter ADAPTOR_SEQ --stringency 5 TEMP_FILE
-
fastq-mcf adapters.fa TEMP_FILE -l 25 -o OUTPUT_FILE
cycling fastx_clipper and and trim_galore (which itself pipelines CutAdapt) over a set of common adaptor sequences, passing the output of each iteration to a temporary file, followed by application of fastq-mcf, which takes all the adapters in a single file. Minimum read length is specified as 25 for each of the 3 commands, but this is an operator-entered variable in Read_cleaner.
Ethical approval required that no genetic material remain within the samples which could enable identification of patients. Therefore, human genome and transcriptome sequences were removed by iterative alignment onto the NCBI, Ensembl and UCSC human iGenomes (http://support.illumina.com/sequencing/sequencing_software/igenome.html), first using bowtie 1.1.1 (ref. 16) (http://bowtie-bio.sourceforge.net/index.shtml), then BWA 0.7.12-r1039 (ref. 17) (http://bio-bwa.sourceforge.net). Following each alignment, extraction of unaligned reads was achieved using samtools 0.1.19 (ref. 18) (http://samtools.sourceforge.net) and the next alignment commenced. Bowtie, BWA and samtools were co-ordinated using the Vanator pipeline (ref. 19).
Vanator implements the following commands for bowtie and BWA:
-
bwa aln GENOME READS>SAI_INDEX
-
bwa samse GENOME SAI_INDEX READS>SAM
-
bowtie -S -p 8 GENOME READS SAM
-
samtools view -S SAM -b -o BAM
-
samtools sort BAM BAM_FILE
-
samtools index BAM
-
bam2fastq --no-aligned --force --strict -o READS BAM
cycling over the three reference human genomes, each time retrieving the unaligned reads using bam2fastq.
Quality improvements were checked, again using FastQC (see Technical Validation).
Identification of potential bacterial contaminants in the sequence pools
Potential contaminating genomes (produced by the presence of non-metagenome DNA contaminants in sequencing reagents) were hypothesised by reference to Salter et al. (ref. 12), and a representative screening set was selected as follows: NC010725 Methylobacterium populi BJ001; NC009485 Bradyrhizobium sp. BTAi1; NC014323 Herbaspirillum seropedicae SmR1; NC022438 Leifsonia xyli subsp. cynodontis DSM 46306; NC015675 Mesorhizobium opportunistum WSM2075; NC007794 Novosphingobium aromaticivorans DSM 12444; NC008313 Ralstonia eutropha H16 chromosome 1; NZCP009571 Sphingomonas taxi strain ATCC 55669; NZCP010409 Xanthomonas sacchari strain R1.
An assessment of the relative contribution of such contaminants was produced by aligning each sequence pool onto the genomes listed above using bowtie 1.1.1 (ref. 16) (http://bowtie-bio.sourceforge.net/index.shtml). However, no further action was taken to remove reads corresponding to these contaminants, or related species, and they remain in the data (see Technical Validation).
Code availability
The unpublished Read_cleaner pipeline is freely available under CC-BY license (Data Citation 17) along with instructions for its use. Vanator (ref. 19) is available at https://sourceforge.net/projects/vanator-cvr.
Data Records
The primary data discussed in this paper is the Illumina deep sequencing output available in Data Citations 1–9. The data format of the deep sequencing output is FASTQ (ref. 20), which adds sequencing quality data to the well-established FASTA nucleic acid sequence format (ref. 21). Viral genome sequences derived from the deep sequencing output by assembly (see Methods) are given in Data Citations 10–15. These are in GenBank format.
Table 1 cross-references the SRA BioSample, Experiment, Accession and Run references against the clinical sources and other data on each sequencing pool. These different accessions provide alternative access points to the data within SRA. They are intended to operate in a hierarchical manner, allowing for instance several sequencing ‘Runs’ to be derived from each ‘Experiment’, and several ‘Experiments’ to be performed on a single ‘BioSample’, and so on. In our work, the hierarchy is sparse, in that each ‘Sample’ was treated as a separate ‘Experiment’, and only one deep sequencing ‘Run’ performed on each.
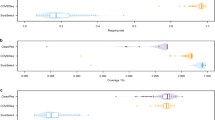
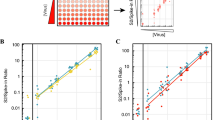
Table 2 gives the proportion of reads aligning to the top two contaminants of each sequencing pool. In all pools, Methylobacterium populi is the most common contaminant, with more than 3% of reads in all pools except F, and up to 21% of reads in pool A, and the second most common is Ralstonia eutropha, with under 0.7% of reads in all pools. Other contaminants total 0.1 to 1.2% of reads.
Technical Validation
Deep sequencing read technical quality
FastQC (https://www.bioinformatics.babraham.ac.uk/projects/fastqc/) was used to check the quality of the Illumina sequencing reads, both in their raw state and after the cleaning processes described above in Methods. FastQC output is deposited in Data Citation 18.
Deep sequencing reagent contaminants
Alignments of each sequencing pool to a set of representative common reagent contaminant genomes (ref. 12) was performed using bowtie 1.1.1 (ref. 16) (http://bowtie-bio.sourceforge.net/index.shtml). The BAM files, their BAI indexes and the contaminant reference genomes are deposited in Data Citation 19.
Additional information
How to cite this article: Atkinson, K. V. et al. Nasopharyngeal metagenomic deep sequencing data, Lancaster, UK, 2014–2015. Sci. Data 4:170161 doi: 10.1038/sdata.2017.161 (2017).
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
References
References
Keech, M. & Beardsworth, P. The impact of influenza on working days lost: a review of the literature. PharmacoEconomics 26, 911–924 (2008).
WHO. Influenza update. Available at http://www.who.int/influenza/surveillance_monitoring/updates/latest_update_GIP_surveillance/en/ (2017).
Nickbakhsh, S. et al. Extensive multiplex PCR diagnostics reveal new insights into the epidemiology of viral respiratory infections. Epidemiology and infection 144, 2064–2076 (2016).
Gong, Y. N. et al. A metagenomics study for the identification of respiratory viruses in mixed clinical specimens: an application of the iterative mapping approach. Archives of virology 162, 2003–2012 (2017).
Schlaberg, R. et al. Viral Pathogen Detection by Metagenomics and Pan Viral Group PCR in Children with Pneumonia Lacking Identifiable Etiology. The Journal of infectious diseases 215, 1407–1415 (2017).
Zou, X. et al. Simultaneous virus identification and characterization of severe unexplained pneumonia cases using a metagenomics sequencing technique. Science China. Life sciences 60, 279–286 (2017).
Atkinson, K. V. et al. Influenza C in Lancaster, UK, in the winter of 2014-2015. Scientific reports 7, 46578 (2017).
Atkinson, K. V. et al. Genome Sequence of Human Rhinovirus A22, Strain Lancaster/2015. Genome announcements 5, e01713–16 (2017).
Atkinson, K. V. et al. Genome Sequence of Human Papillomavirus 23 Strain HPV-23/Lancaster/2015. Genome announcements 5, e00257–17 (2017).
Atkinson, K. V. et al. Genome Sequence of Human Papillomavirus Type 20, Strain HPV-20/Lancaster/2015. Genome announcements 5, e00712–17 (2017).
Matsuzaki, Y. et al. Clinical features of influenza C virus infection in children. The Journal of infectious diseases 193, 1229–1235 (2006).
Salter, S. J. et al. Reagent and laboratory contamination can critically impact sequence-based microbiome analyses. BMC Biol 12, 87 (2014).
Perez-Losada, M., Alamri, L., Crandall, K. A. & Freishtat, R. J. Nasopharyngeal Microbiome Diversity Changes over Time in Children with Asthma. PLoS ONE 12, e0170543 (2017).
Martin, M. Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet.Journal 17, 10–12 (2011).
Aronesty, E. Comparison of sequencing utility programs. The Open Bioinformatics Journal 7, 1–8 (2013).
Langmead, B., Trapnell, C., Pop, M. & Salzberg, S. L. Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome biology 10, R25 (2009).
Li, H. & Durbin, R. Fast and accurate long-read alignment with Burrows-Wheeler transform. Bioinformatics 26, 589–595 (2010).
Li, H. et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Jarrett, R. F., Gallagher, A. & Gatherer, D. Molecular methods of virus detection in lymphoma. Methods Mol Biol 971, 277–293 (2013).
Cock, P. J., Fields, C. J., Goto, N., Heuer, M. L. & Rice, P. M. The Sanger FASTQ file format for sequences with quality scores, and the Solexa/Illumina FASTQ variants. Nucleic acids research 38, 1767–1771 (2010).
Pearson, W. R. & Lipman, D. J. Improved tools for biological sequence comparison. Proceedings of the National Academy of Sciences of the United States of America 85, 2444–2448 (1988).
Data Citations
NCBI Sequence Read Archive SRX2310763 (2016)
NCBI Sequence Read Archive SRX2310764 (2016)
NCBI Sequence Read Archive SRX2310765 (2016)
NCBI Sequence Read Archive SRX2310766 (2016)
NCBI Sequence Read Archive SRX2310759 (2016)
NCBI Sequence Read Archive SRX2310760 (2016)
NCBI Sequence Read Archive SRX2310761 (2016)
NCBI Sequence Read Archive SRX2310762 (2016)
NCBI Sequence Read Archive SRX2310758 (2016)
GenBank KY075640 (2016)
GenBank KY075641 (2016)
GenBank KY075642 (2016)
GenBank KY342346 (2017)
GenBank KY652675 (2017)
GenBank KY969593 (2017)
NCBI BioProject PRJNA310124 (2016)
Gatherer, D. Lancaster University https://doi.org/10.17635/lancaster/researchdata/111 (2016)
Gatherer, D. Lancaster University https://doi.org/10.17635/lancaster/researchdata/153 (2017)
Gatherer, D. Lancaster University https://doi.org/10.17635/lancaster/researchdata/171 (2017)
Acknowledgements
Funding was provided under the following grants: ‘Influenza C—a pilot study on a neglected respiratory virus’ (Grant ID NIHR-17799, University Hospitals of Morecambe Bay National Health Service Foundation Trust). ‘How important is influenza C virus?’ (Grant ID M395, Rosetrees Trust).
Author information
Authors and Affiliations
Contributions
K.V.A. collected the samples and performed laboratory work, except deep sequencing. L.A.B. supervised laboratory work. G.R. supervised laboratory work. N.S. provided positive control samples for influenza C detection. N.R.M. performed deep sequencing. M.J.H. performed deep sequencing. J.R. assisted in sample collection. N.H. assisted in sample collection. S.W. provided clinical supervision in the general practice surgery. R.M.L. provided executive laboratory supervision. R.W.P. provided executive laboratory supervision and was a named co-supervisor for the MSc degree of KVA. M.W. provided clinical supervision in the hospital and was a named co-supervisor for the MSc degree of KVA. D.G. wrote the grant and ethics applications, drafted and corrected the paper, and was principal supervisor for the MSc degree of K.V.A.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
ISA-Tab metadata
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/ The Creative Commons Public Domain Dedication waiver http://creativecommons.org/publicdomain/zero/1.0/ applies to the metadata files made available in this article.
About this article
Cite this article
Atkinson, K., Bishop, L., Rhodes, G. et al. Nasopharyngeal metagenomic deep sequencing data, Lancaster, UK, 2014–2015. Sci Data 4, 170161 (2017). https://doi.org/10.1038/sdata.2017.161
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/sdata.2017.161