Abstract
Pharmacologic inhibitors of cyclin-dependent kinases 4 and 6 (CDK4/6) were designed to induce cancer cell cycle arrest. Recent studies have suggested that these agents also exert other effects, influencing cancer cell immunogenicity, apoptotic responses and differentiation. Using cell-based and mouse models of breast cancer together with clinical specimens, we show that CDK4/6 inhibitors induce remodeling of cancer cell chromatin characterized by widespread enhancer activation, and that this explains many of these effects. The newly activated enhancers include classical super-enhancers that drive luminal differentiation and apoptotic evasion, as well as a set of enhancers overlying endogenous retroviral elements that are enriched for proximity to interferon-driven genes. Mechanistically, CDK4/6 inhibition increases the level of several activator protein-1 transcription factor proteins, which are in turn implicated in the activity of many of the new enhancers. Our findings offer insights into CDK4/6 pathway biology and should inform the future development of CDK4/6 inhibitors.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
Data availability
ATAC-seq, ChIP-seq, RNA-seq and HiChIP data that support the findings of this study have been deposited into the Gene Expression Omnibus (GEO) repository under accession codes GSE157097, GSE157082, GSE157222, GSE157218, GSE157216, GSE157214, GSE157211, GSE157385, GSE157384, GSE157383 and GSE157381. Previously deposited transcriptomic data (GSE99062 (ref. 6) and GSE93204 (ref. 42)) that are used in this study are also available at GEO4. Source data for Figs. 1–4 and 6–8 and Extended Data Figs. 1, 3, 4, 6 and 7 are provided with this paper. All other data that support the findings of this study are available from the corresponding author upon reasonable request. Source data are provided with this paper.
Code availability
No unpublished code was used in this manuscript. Source data are provided with this paper.
References
Sherr, C. J., Beach, D. & Shapiro, G. I. Targeting CDK4 and CDK6: from discovery to therapy. Cancer Discov. 6, 353–367 (2016).
Spring, L. M. et al. Cyclin-dependent kinase 4 and 6 inhibitors for hormone receptor-positive breast cancer: past, present, and future. Lancet 395, 817–827 (2020).
Finn, R. S. et al. PD 0332991, a selective cyclin D kinase 4/6 inhibitor, preferentially inhibits proliferation of luminal estrogen receptor-positive human breast cancer cell lines in vitro. Breast Cancer Res. 11, R77 (2009).
Goel, S. et al. Overcoming therapeutic resistance in HER2-positive breast cancers with CDK4/6 inhibitors. Cancer Cell 29, 255–269 (2016).
Herrera-Abreu, M. T. et al. Early adaptation and acquired resistance to CDK4/6 inhibition in estrogen receptor-positive breast cancer. Cancer Res. 76, 2301–2313 (2016).
Goel, S. et al. CDK4/6 inhibition triggers anti-tumour immunity. Nature 548, 471–475 (2017).
Schaer, D. A. et al. The CDK4/6 inhibitor abemaciclib induces a T cell inflamed tumor microenvironment and enhances the efficacy of PD-L1 checkpoint blockade. Cell Rep. 22, 2978–2994 (2018).
Hurvitz, S. et al. Biological effects of abemaciclib in a phase 2 neoadjuvant study for postmenopausal patients with HR+, HER2− breast cancer. Cancer Res. 77, S4–S06 (2017).
Choi, Y. J. et al. The requirement for cyclin D function in tumor maintenance. Cancer Cell 22, 438–451 (2012).
Yoshida, A., Lee, E. K. & Diehl, J. A. Induction of therapeutic senescence in vemurafenib-resistant melanoma by extended inhibition of CDK4/6. Cancer Res. 76, 2990–3002 (2016).
Narita, M. et al. Rb-mediated heterochromatin formation and silencing of E2F target genes during cellular senescence. Cell 113, 703–716 (2003).
Shay, J. W., Pereira-Smith, O. M. & Wright, W. E. A role for both RB and p53 in the regulation of human cellular senescence. Exp. Cell. Res. 196, 33–39 (1991).
Munoz-Espin, D. & Serrano, M. Cellular senescence: from physiology to pathology. Nat. Rev. Mol. Cell Biol. 15, 482–496 (2014).
Tasdemir, N. et al. BRD4 connects enhancer remodeling to senescence immune surveillance. Cancer Discov. 6, 612–629 (2016).
Sen, P. et al. Histone acetyltransferase p300 induces de novo super-enhancers to drive cellular senescence. Mol. Cell 73, 684–698 (2019).
Buenrostro, J. D., Giresi, P. G., Zaba, L. C., Chang, H. Y. & Greenleaf, W. J. Transposition of native chromatin for fast and sensitive epigenomic profiling of open chromatin, DNA-binding proteins and nucleosome position. Nat. Methods 10, 1213–1218 (2013).
McLean, C. Y. et al. GREAT improves functional interpretation of cis-regulatory regions. Nat. Biotechnol. 28, 495–501 (2010).
Chicas, A. et al. H3K4 demethylation by Jarid1a and Jarid1b contributes to retinoblastoma-mediated gene silencing during cellular senescence. PNAS 109, 8971–8976 (2012).
Cejas, P. et al. Chromatin immunoprecipitation from fixed clinical tissues reveals tumor-specific enhancer profiles. Nat. Med. 22, 685–691 (2016).
Font-Tello, A. et al. FiTAc-seq: fixed-tissue ChIP-seq for H3K27ac profiling and super-enhancer analysis of FFPE tissues. Nat. Protoc. 15, 2503–2518 (2020).
Mumbach, M. R. et al. HiChIP: efficient and sensitive analysis of protein-directed genome architecture. Nat. Methods 13, 919–922 (2016).
Wang, S. et al. Target analysis by integration of transcriptome and ChIP-seq data with BETA. Nat. Protoc. 8, 2502–2515 (2013).
Whyte, W. A. et al. Master transcription factors and mediator establish super-enhancers at key cell identity genes. Cell 153, 307–319 (2013).
Loven, J. et al. Selective inhibition of tumor oncogenes by disruption of super-enhancers. Cell 153, 320–334 (2013).
Welch, R. P. et al. ChIP-Enrich: gene set enrichment testing for ChIP-seq data. Nucleic Acids Res. 42, e105 (2014).
Fu, N. Y. et al. Foxp1 is indispensable for ductal morphogenesis and controls the exit of mammary stem cells from quiescence. Dev. Cell 47, 629–644 (2018).
Hong, D. et al. Runx1 stabilizes the mammary epithelial cell phenotype and prevents epithelial to mesenchymal transition. Oncotarget 8, 17610–17627 (2017).
Sengupta, S. & George, R. E. Super-enhancer-driven transcriptional dependencies in cancer. Trends Cancer 3, 269–281 (2017).
Montero, J. et al. Drug-induced death signaling strategy rapidly predicts cancer response to chemotherapy. Cell 160, 977–989 (2015).
Del Gaizo Moore, V. & Letai, A. BH3 profiling–measuring integrated function of the mitochondrial apoptotic pathway to predict cell fate decisions. Cancer Lett. 332, 202–205 (2013).
De Cecco, M. et al. Genomes of replicatively senescent cells undergo global epigenetic changes leading to gene silencing and activation of transposable elements. Aging Cell 12, 247–256 (2013).
Criscione, S. W., Zhang, Y., Thompson, W., Sedivy, J. M. & Neretti, N. Transcriptional landscape of repetitive elements in normal and cancer human cells. BMC Genomics 15, 583 (2014).
Fuentes, D. R., Swigut, T. & Wysocka, J. Systematic perturbation of retroviral LTRs reveals widespread long-range effects on human gene regulation. eLife 7, e35989 (2018).
Chuong, E. B., Elde, N. C. & Feschotte, C. Regulatory evolution of innate immunity through co-option of endogenous retroviruses. Science 351, 1083–1087 (2016).
Zheng, R. et al. Cistrome Data Browser: expanded datasets and new tools for gene regulatory analysis. Nucleic Acids Res. 47, D729–D735 (2019).
Pellacani, D. et al. Analysis of normal human mammary epigenomes reveals cell-specific active enhancer states and associated transcription factor networks. Cell Rep. 17, 2060–2074 (2016).
Dravis, C. et al. Epigenetic and transcriptomic profiling of mammary gland development and tumor models disclose regulators of cell state plasticity. Cancer Cell 34, 466–482 (2018).
Nead, M. A., Baglia, L. A., Antinore, M. J., Ludlow, J. W. & McCance, D. J. Rb binds c-Jun and activates transcription. EMBO J. 17, 2342–2352 (1998).
Nishitani, J. et al. Recruitment of the retinoblastoma protein to c-Jun enhances transcription activity mediated through the AP-1 binding site. J. Biol. Chem. 274, 5454–5461 (1999).
Angel, P., Hattori, K., Smeal, T. & Karin, M. The jun proto-oncogene is positively autoregulated by its product, Jun/AP-1. Cell 55, 875–885 (1988).
Fanjul, A. et al. A new class of retinoids with selective inhibition of AP-1 inhibits proliferation. Nature 372, 107–111 (1994).
Ma, C. X. et al. NeoPalAna: neoadjuvant palbociclib, a cyclin-dependent kinase 4/6 inhibitor, and anastrozole for clinical stage 2 or 3 estrogen receptor-positive breast cancer. Clin. Cancer Res. 23, 4055–4065 (2017).
Haines, E. et al. Palbociclib resistance confers dependence on an FGFR-MAP kinase-mTOR-driven pathway in KRAS-mutant non-small cell lung cancer. Oncotarget 9, 31572–31589 (2018).
Walter, D. M. et al. RB constrains lineage fidelity and multiple stages of tumour progression and metastasis. Nature https://doi.org/10.1038/s41586-019-1172-9 (2019).
Leverson, J. D. et al. Exploiting selective BCL-2 family inhibitors to dissect cell survival dependencies and define improved strategies for cancer therapy. Sci. Transl. Med. 7, 279ra240 (2015).
Corces, M. R. et al. An improved ATAC-seq protocol reduces background and enables interrogation of frozen tissues. Nat. Methods 14, 959–962 (2017).
Buenrostro, J. D., Wu, B., Chang, H. Y. & Greenleaf, W. J. ATAC-seq: a method for assaying chromatin accessibility genome-wide. Curr. Protoc. Mol. Biol. 109, 21–29 (2015). 21 29.
Savic, D., Gertz, J., Jain, P., Cooper, G. M. & Myers, R. M. Mapping genome-wide transcription factor binding sites in frozen tissues. Epigenetics Chromatin 6, 30 (2013).
Servant, N. et al. HiC-Pro: an optimized and flexible pipeline for Hi-C data processing. Genome Biol. 16, 259 (2015).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl Acad. Sci. USA 102, 15545–15550 (2005).
Ritchie, M. E. et al. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 43, e47 (2015).
Qin, Q. et al. ChiLin: a comprehensive ChIP-seq and DNase-seq quality control and analysis pipeline. BMC Bioinf. 17, 404 (2016).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows–Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
Zhang, Y. et al. Model-based analysis of ChIP-Seq (MACS). Genome Biol. 9, R137 (2008).
Shin, H., Liu, T., Manrai, A. K. & Liu, X. S. CEAS: cis-regulatory element annotation system. Bioinformatics 25, 2605–2606 (2009).
Quinlan, A. R. & Hall, I. M. BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics 26, 841–842 (2010).
Ramirez, F., Dundar, F., Diehl, S., Gruning, B. A. & Manke, T. deepTools: a flexible platform for exploring deep-sequencing data. Nucleic Acids Res. 42, W187–W191 (2014).
Cornwell, M. et al. VIPER: visualization pipeline for RNA-seq, a snakemake workflow for efficient and complete RNA-seq analysis. BMC Bioinf. 19, 135 (2018).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Robinson, M. D., McCarthy, D. J. & Smyth, G. K. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26, 139–140 (2010).
Ay, F., Bailey, T. L. & Noble, W. S. Statistical confidence estimation for Hi-C data reveals regulatory chromatin contacts. Genome Res. 24, 999–1011 (2014).
Langmead, B., Trapnell, C., Pop, M. & Salzberg, S. L. Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol. 10, R25 (2009).
Bhattacharyya, S., Chandra, V., Vijayanand, P. & Ay, F. Identification of significant chromatin contacts from HiChIP data by FitHiChIP. Nat. Commun. 10, 4221 (2019).
Acknowledgements
This work is supported by the National Health and Medical Research Council of Australia (Investigator Grant GNT1177357 to S.G.); Susan G. Komen for the Cure (to I.K. and M.B. and CCR18547966 to S.G.); the Breast Cancer Alliance (Young Investigator Grant to S.G.); the Royal Australasian College of Physicians (Research Establishment Fellowship to S.G.); Eli Lilly and Co (to S.G.); the NIH/NCI (P50 CA168504 to S.G., I.K., E.W. and J.J.Z.; R35 CA210057 to J.J.Z.; P50 CA165962-06A1 to J.J.Z.; P01CA080111 to M.B.; R01CA193910 to M.L.F.; R02HG005220 to K.D.K. (PI Rafael A. Irizarry), R01GM083084 to K.D.K. (PI Rafael A. Irizarry), U41HG004059 to K.D.K. (PI Martin T. Morgan)); the US Department of Defense CDMRP (W81XWH-18-1-0491 to J.J.Z.); the Breast Cancer Research Foundation (BCRF-18-179 to J.J.Z. and O.M.-F.), the Ministry of Economy and Competitiveness (Instituto de Salud Carlos III) PI18-01604 to P.C.; the Chan Zuckerberg Initiative (DAF grant 2018-183142 to K.D.K. (PI Rafael A. Irizarry)); Stand Up To Cancer (SU2C) and The V Foundation (TVF) SU2C-TVF Convergence Scholar Awards (D2015-037 to J.M.); Ramon y Cajal Programme, Ministerio de Economia y Competitividad (RYC-2015-18357 to J.M.); Terri Brodeur Breast Cancer Foundation (to V.W.D.); and the European Union’s Horizon 2020 Research and Innovation Programme (Marie Skłodowska-Curie grant agreement no. 754490 to C.G. and the MINDED project).
Author information
Authors and Affiliations
Contributions
S.G. developed the concept behind the study and supervised the project, with important advice from H.W.L., A.C.W., P.C. and M.J.D. Contributions include: model development (S.G., J.J.Z., M.J.D., A.C.W. and H.B.J.); formal analysis preclinical data, including computational analyses (S.G., P.C., N.K., H.W.L., M.J.D., A.C.W., H.B.J., P.C., X.Q., E.Y.N.L., J.M., V.W.D., L.T., C.A.M., K.D.K., O.G. and C.C.B.); formal analysis of clinical data (O.M.-F., S.G., P.C., H.W.L., X.Q., C.M., M.J.E., E.Y.N.L. and A.C.W.); preclinical experiments (A.C.W., P.C., M.J.D., H.B.J., R.C., A.F.T., K.L., J.M., R.V., V.W.D. and S.G.); computational tool development (N.K., L.T., X.Q., E.Y.N.L., H.W.L., P.C., S.G., C.G., B.P., J.H.S. and M.L.F.); resource provision (S.G., J.J.Z., M.B., C.M., M.J.E., E.W., I.K., M.A.D. and A.L.); writing (S.G., A.C.W., M.J.D., P.C., H.W.L. and N.K.); supervision and oversight (S.G., J.J.Z. and H.W.L.); and funding acquisition (S.G. and J.J.Z.).
Corresponding authors
Ethics declarations
Competing interests
S.G. is the recipient of research funding from Eli Lilly and Co., which has been used to support a part of this work. S.G. has served as a paid advisory board member for Eli Lilly and Co., G1 Therapeutics, Pfizer and Novartis. S.G. also conducts laboratory research sponsored by G1 Therapeutics and clinical research sponsored by Eli Lilly and Co. and by Novartis. J.J.Z. is founder and board director of Crimson Biotech and Geode Therapeutics. J.M. is a consultant for Oncoheroes Biosciences and Vivid Biosciences. M.B. receives sponsored research support from Novartis. M.B. is a consultant to H3 Biomedicine and serves on the SAB of Kronos Bio and GV20 Therapeutics. C.M. is a consultant for Pfizer, Novartis, Seattle Genetics and Eli Lilly and Co. and receives institutional research funding from Pfizer and Puma. I.K. receives institutional research funding and grants from Genentech/Roche and Pfizer. I.K. is an advisory board participant, consultant and has received honoraria from Daiichi/Sankyo, Macrogenics and Genentech/Roche. I.K. is an advisory board participant and has received honoraria from Context Therapeutics, Taiho Oncology and Seattle Genetics. I.K. is a data monitoring board member at Novartis. E.W. is a consultant at and has received honoraria from Carrick Therapeutics, DragonFly, Genentech/Roche, Genomic Health, GSK, Jounce, Eli Lilly and Co., Seattle Genetics and Merck. E.W. is a scientific advisory board member and has received honoraria from Leap. M.E. has patents and receives royalties from prosigna/Nanostring. O.M.-F. receives institutional research funding from AbbVie, Genentech, Roche and Pfizer, and has received honoraria from Roche. M.A.D. has been a member of the advisory boards for CTX CRC, Storm Therapeutics, Celgene and Cambridge Epigenetix. The Dawson laboratory receives research funding from CTX CRC. All other authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Transcriptional, phenotypic, and epigenetic effects of CDK4/6 inhibitors in human breast cancer cells.
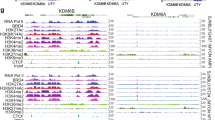
a, Relative RNA-seq normalized reads of representative E2F target genes in breast cancer cell lines treated with abemaciclib for the indicated times (n=2 in MCF7, n=1 in MDA-MB-453, independent cultures). b, Percentage of cells in S-phase in cell lines treated with dimethyl sulfoxide (DMSO; control) or abemaciclib (abema) for 24 hours (n=3 independent cultures). c, Representative senescence-associated β-galactosidase staining (blue) of cells treated with DMSO or abemaciclib for 1, 3, or 7 days. Scale bars represent 200 μm. Representative images of two independent experiments in MCF7 and two technical replicates from one experiment in MDA-MB-453. d, Genomic distribution of regions of significantly reduced ATAC-seq signal in cells treated with abemaciclib, compared to DMSO. e, GREAT (Genomic Regions Enrichment of Annotations) analysis of regions of significantly reduced ATAC-seq signal within 10 kb of the single nearest gene in cells treated with abemaciclib for 7 days (compared to DMSO). f, Number of regions with significantly increased ATAC-seq signal in cells treated with abemaciclib as indicated. g, Heatmap of regions with significantly increased ATAC-seq peak signal after abemaciclib treatment in MDA-MB-361. Up-peaks were determined by a threshold of adjusted P<0.05 calculated by DESeq2. h, Western blot for RB in MCF7 shLuc and shRB1 cells, representative images from two independent experiments. Western blots are cropped; uncropped blot images for the experiments in this figure are shown in Source Data Extended Data Fig. 1. i, Relative RNA-seq normalized reads of cell cycle-related genes in MCF7 shRB1 cells and MCF7 shLuc cells (n=2 independent cultures) treated with DMSO or abemaciclib. j,k, Composite profiles of H3K27ac (j) and H3K9me3 (k) ChIP-seq signals at regions of significantly increased ATAC-seq signal in MCF7 and MDA-MB-453 treated with DMSO or abemaciclib for 7 days. l, Heatmap of H3K27ac ChIP-seq profiles in MCF7 treated with DMSO, abemaciclib, or palbociclib at abemaciclib-induced H3K27ac up-peak regions. m, Sample-sample correlation between RNA-seq samples of MCF7 and MDA-MB-453 treated with DMSO or abemaciclib (parental: n=3; shRB1: n=2).
Extended Data Fig. 2 Characterization of CDK4/6 inhibitor-activated super-enhancers in breast cancer cell lines.
a, Signal distribution at abemaciclib-activated H3K27ac-marked enhancers in MCF7 and MDA-MB-453 cells. Arrows indicate the positioning of super-enhancers (to the right of dashed line). b, Schema for analysis of biological processes regulated by abemaciclib-activated super-enhancers. c, GO analysis of non-TSS ends of H3K27ac-decorated genomic loops (HiChIP) identified exclusively in abemaciclib-treated MCF7 and MDA-MB-453 cells. Odds ratios and P-values were calculated by ChIP-Enrich. d, Gene Set Enrichment Analysis (GSEA) of RNA-seq data from MCF7 and MDA-MB-453 cells treated with abemaciclib compared to DMSO. e, RNA-seq log2 fold change of gene expression of all genes within indicated GSEA gene sets in MCF7 and MDA-MB-453 cells treated with DMSO or abemaciclib for 7 days, calculated by DESeq2. Each dot represents one gene: red, predicted to be regulated by abemaciclib-activated super-enhancers; blue, not predicted to be regulated by abemaciclib-activated super-enhancers. f, GSEA of RNA-seq data from MCF7 and MDA-MB-453 cells treated with palbociclib compared to DMSO. FDR q values in d and f were calculated by GSEAPreranked.
Extended Data Fig. 3 Enhanced expression of luminal differentiation-related genes is associated with increased H3K27 acetylation at nearby super-enhancers.
a, Representative H3K27ac ChIP-seq genome browser tracks with co-localized H3K27ac-decorated genomic loops (HiChIP) in MCF7 cells treated with DMSO or abemaciclib for 7 days. Gene promoters are highlighted in grey. b, Relative RNA-seq normalized reads of genes in MCF7 shRB1 cells treated with DMSO or abemaciclib for 7 days (n=2 independent cultures). c, Relative mRNA expression of genes shown in Fig. 3d in MMTV-rtTA/tetO-Her2 tumors treated for up to 59 days with vehicle or abemaciclib (n=3 independent tumors). Means ± s.d. are shown. P-values were determined by two-tailed unpaired t-tests corrected for multiple comparisons by Holm-Sidak method.
Extended Data Fig. 4 CDK4/6 inhibition promotes apoptotic evasion via BCL2L1.
a,b, ChIP-Enrich analysis of regions gaining H3K27ac after 7 days of abemaciclib treatment of PDX 14-07 tumors (21-28 days of treatment) (a), or of non-TSS ends of genomic loops (measured by HiChIP) detected only in abemaciclib-treated MCF7 and MDA-MB-453 cells (b). c, Dose-response curves showing percent mitochondrial depolarization in MDA-MB-361 and BT474 treated with DMSO or abemaciclib for 7 days and after exposure to Bim peptide (dynamic BH3 profiling; n=2 technical replicates, data are representative of two independent experiment). d-f, Relative RNA-seq normalized reads of MCL1 (d), BCL2 (e), and BCL2L2 (f) in human breast cancer cell lines treated with DMSO or abemaciclib, measured by RNA-seq (MDA-MB-361 and T47D: n=2; MCF7, MDA-MB-453, and BT474: n=3 independent cultures). Mean ± s.d. are shown. DESeq2 was used to determine adjusted P-values. g,h, Relative RNA-seq normalized reads of BCL2L1 and MCL1 in MCF7 shRB1 cells (g; n=2 independent cultures) and MDA-MB-468 (h, RB1 null; n=3 independent cultures) treated with DMSO or abemaciclib. Data are presented as mean ± s.d. DESeq2 was used to calculate adjusted P-values. i, Relative Bcl2l1 expression by transcriptomic profiling of MMTV-rtTA/tetO-Her2 tumors treated with vehicle (n=23) or abemaciclib (n=25) for 12 days. Mean ± s.d. of normalized reads are shown. P-values were determined using a two-tailed unpaired t-test. j, H3K27ac ChIP-seq tracks and co-localized H3K27ac-decorated genomic loops (identified by HiChIP) adjacent to BCL2L1 in MDA-MB-453 cells treated with DMSO or abemaciclib for 7 days. Gene promoters are highlighted in grey. k, Western blot showing cleaved PARP in MCF7 cells treated with DMSO or abemaciclib (500 nM) for 3 days, followed by treatment with A-1155463 (1 μM) for 24 hours and BYL719 (1 μM) for 8 hours as indicated. Representative blots of two independent experiments in MCF7. Western blots are cropped; uncropped blot images for the experiments in this figure are shown in Source Data Extended Data Fig. 4.
Extended Data Fig. 5 Effect of CDK4/6 inhibition on H3K27ac signal at transposable elements and on interferon-stimulated gene expression.
a, Fold change in H3K27ac signal at LTRs, LINEs, and SINEs (long and short interspersed nuclear elements) identified by RepEnrich in MCF7 and MDA-MB-453 treated with DMSO or abemaciclib for 7 days (red, FC>1.5). b, Cross-species conservation of a representative LTR enhancer activated by abemaciclib treatment in MCF7 cells. c, GSEA using RNA-sequencing data from MCF7 cells treated with abemaciclib compared to DMSO for 7 days, or genome-wide transcriptomic profiling of PDX 14-07 tumors treated with abemaciclib compared with vehicle for 21-28 days. d, GSEA using RNA-sequencing data from MCF7 cells treated with palbociclib compared to DMSO for 7 days. FDR q values in c, d were calculated by GSEAPreranked.
Extended Data Fig. 6 CDK4/6 inhibitor-activated LTR enhancers are predicted to regulate immune genes.
a, Gene Ontology analysis of interferon-related signatures using genes predicted to be regulated by all abemaciclib-activated enhancers, abemaciclib-activated SEs, or abemaciclib-activated LTR enhancers in MCF7 cells treated with DMSO or abemaciclib for 7 days. Odds ratios and P-values calculated by ChIP-Enrich. b, ATAC-seq and H3K27ac ChIP-seq tracks near representative immune genes in MCF7 and MDA-MB-453 treated with DMSO or abemaciclib for 7 days. LTRs annotated using Repeat Masker are shown as blue bars. Yellow highlights indicate H3K27ac up-peaks that align with an LTR. c, Relative RNA-seq normalized reads of immune genes in MCF7 parental and shRB1 cells treated with DMSO or abemaciclib for 7 days, measured by RNA-seq (parental: n=3; shRB1: n=2). Means ± s.d. are shown. DESeq2 was used to calculated adjusted P-values.
Extended Data Fig. 7 Effect of CDK4/6 inhibition on AP-1 gene expression.
a, Similarity (GIGGLE) scores between regions of increased H3K27ac signal genome-wide in MCF7 cells after palbociclib treatment and GEO-archived datasets of ChIP-seq for transcription factors (using “Cistrome Toolkit”). Top 20 factors identified are shown - AP-1 factors and steroid hormone receptors are labeled. b, c, Relative RNA-seq normalized reads of AP-1 members in MCF7 and MDA-MB-453 (b), and in MCF7 shRB1 and MDA-MB-468 (c) treated with DMSO or abemaciclib (MCF7, MDA- MB-453, MDA-MB-468, n=3; MCF7 shRB1, n=2). Means ± s.d. are shown. DESeq2 was used to calculated adjusted P-values.
Extended Data Fig. 8 AP-1 binding increases at CDK4/6 inhibitor-induced enhancers and drives transcriptional activity that can be reversed by an AP-1 inhibitor.
a, Composite profile of c-Jun, JunB, and Fra2 ChIP-seq signals in MCF7 cells treated with DMSO or abemaciclib over regions of abemaciclib-induced H3K27ac up-peaks. b, H3K27ac, c-Jun, JunB, and Fra2 ChIP-seq tracks at PRICKLE2 and CDH1 loci in MCF7 cells treated with DMSO or abemaciclib. DNA loops (called by FDR<0.01) from H3K27ac HiChIP are depicted for PRICKLE2. Grey highlights indicate gene promoters. Green highlights indicate regions with abemaciclib-induced increases in both H3K27ac signal and c-Jun binding. c, Composite profile of c-Jun ChIP-seq signal over abemaciclib-activated LTR enhancers in MCF7 cells treated with DMSO or abemaciclib. d, H3K27ac, c-Jun, JunB, and Fra2 ChIP-seq tracks at RIPK2, HLA-C, and HLA-G loci in MCF7 cells treated with DMSO or abemaciclib. Green highlights as in b. LTRs annotated with Repeat Masker are shown in blue. e, Heatmap of H3K27ac, c-Jun, JunB, Fra2, and estrogen receptor (ER) ChIP-seq profiles at regions showing increased binding for any of c-Jun, JunB, or Fra2 (combined) after 7 days of abemaciclib treatment in MCF7 cells. The cluster “ER” denotes regions with increased ER binding after abemaciclib treatment and contains 1,124 regions. The cluster “no ER or ER unchanged” denotes regions with no change in ER binding or no ER at all and contains 14,504 regions. f, Analysis of GSEA signatures associated with luminal differentiation and interferon response using Principle Component 2 loadings from Fig. 7c. NES and FDR q values were calculated using GSEAPreranked.
Extended Data Fig. 9 Patterns of enhancer activation in breast cancers treated with combined CDK4/6 inhibition and endocrine therapy.
a, ChIP-Enrich analysis of regions gaining H3K27ac in MCF7 cells treated with abemaciclib plus fulvestrant (100 nM) versus DMSO for 7 days. b, Composite profiles of H3K27ac ChIP-seq signal from patient biopsies (baseline and tamoxifen + palbociclib, as in Fig. 8a), centered over ATAC peak-intersected, abemaciclib-induced H3K27ac up-peaks in MCF7 cells. c, H3K27ac ChIP-seq tracks at CDH1 and BCL2L1 from MCF7 cells and tumor biopsies from same patient as in b. d, H3K27ac ChIP-seq tracks at MKI67 and TOP2A from tumor biopsies from same patient as in b.
Supplementary information
Supplementary Information
Supplementary Fig. 1.
Supplementary Tables
Supplementary Table 1–5.
Source data
Source Data Fig. 1
Statistical source data.
Source Data Fig. 2
Statistical source data.
Source Data Fig. 3
Statistical source data.
Source Data Fig. 4
Statistical source data.
Source Data Fig. 4
Unprocessed western blot images.
Source Data Fig. 6
Statistical source data.
Source Data Fig. 6
Unprocessed western blot images.
Source Data Fig. 7
Statistical source data.
Source Data Fig. 8
Statistical source data.
Source Data Extended Data Fig. 1
Statistical source data.
Source Data Extended Data Fig. 1
Unprocessed western blot images.
Source Data Extended Data Fig. 3
Statistical source data.
Source Data Extended Data Fig. 4
Statistical source data.
Source Data Extended Data Fig. 4
Unprocessed western blot images.
Source Data Extended Data Fig. 6
Statistical source data.
Source Data Extended Data Fig. 7
Statistical source data.
Rights and permissions
About this article
Cite this article
Watt, A.C., Cejas, P., DeCristo, M.J. et al. CDK4/6 inhibition reprograms the breast cancer enhancer landscape by stimulating AP-1 transcriptional activity. Nat Cancer 2, 34–48 (2021). https://doi.org/10.1038/s43018-020-00135-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s43018-020-00135-y
This article is cited by
-
The BRD4 inhibitor JQ1 augments the antitumor efficacy of abemaciclib in preclinical models of gastric carcinoma
Journal of Experimental & Clinical Cancer Research (2023)
-
Etiology of super-enhancer reprogramming and activation in cancer
Epigenetics & Chromatin (2023)
-
Spatial transcriptomic analysis of Sonic hedgehog medulloblastoma identifies that the loss of heterogeneity and promotion of differentiation underlies the response to CDK4/6 inhibition
Genome Medicine (2023)
-
Ratios of monocytes and neutrophils to lymphocytes in the blood predict benefit of CDK4/6 inhibitor treatment in metastatic breast cancer
Scientific Reports (2023)
-
Risk of secondary tumours in patients with non-metastatic and metastatic human retinoblastoma
Eye (2023)