Abstract
Breast density, or the amount of fibroglandular tissue (FGT) relative to the overall breast volume, increases the risk of developing breast cancer. Although previous studies have utilized deep learning to assess breast density, the limited public availability of data and quantitative tools hinders the development of better assessment tools. Our objective was to (1) create and share a large dataset of pixel-wise annotations according to well-defined criteria, and (2) develop, evaluate, and share an automated segmentation method for breast, FGT, and blood vessels using convolutional neural networks. We used the Duke Breast Cancer MRI dataset to randomly select 100 MRI studies and manually annotated the breast, FGT, and blood vessels for each study. Model performance was evaluated using the Dice similarity coefficient (DSC). The model achieved DSC values of 0.92 for breast, 0.86 for FGT, and 0.65 for blood vessels on the test set. The correlation between our model’s predicted breast density and the manually generated masks was 0.95. The correlation between the predicted breast density and qualitative radiologist assessment was 0.75. Our automated models can accurately segment breast, FGT, and blood vessels using pre-contrast breast MRI data. The data and the models were made publicly available.
Similar content being viewed by others
Introduction
Algorithmic quantitative analysis of medical images could lead to improved diagnostic accuracy and prognosis for patients, but this is hindered by multiple factors. These include: (1) poor availability of shared data that could be used for the development of algorithmic models and as a common benchmark for model evaluation, (2) poor public availability of the developed models that would allow for comparison between different approaches and use for downstream tasks, and (3) imprecise definitions of the quantities measured in images, leading to inconsistent model development and evaluation with high inter-reader variability when these quantities are assessed by human readers, such as radiologists. An example of a measurement that suffers from all these drawbacks is breast density.
Breast density has strong clinical implications despite its qualitative assessment methods. Many studies have shown that women with a higher breast density are at a greater risk for developing breast cancer1,2,3,4. Breast density, or the amount of fibroglandular tissue (FGT) relative to overall breast volume, is typically assessed by a radiologist qualitatively or by using semi-automatic quantitative tools. However, both methods still have inter-user variability, as radiologists use qualitative descriptors to perform categorization (e.g. “scattered” vs. “heterogenous” vs. “extreme”) with kappa values ranging from 0.69 to 0.84 for inter-radiologist agreement, and some semi-automatic tools have been shown to underestimate breast density5,6,7,8,9. Therefore, there is a need for an objective and efficient method of automated breast density assessment.
The imaging modality that allows for the highest contrast between tissues within the breast is magnetic resonance imaging (MRI). It is used to screen high-risk women and stage women with newly diagnosed breast cancer, among other indications10. It has many advantages over standard mammography, including increased sensitivity, three-dimensional visualization of the breast and axilla, and dynamic information about blood flow11. Due to the nature of MRI usage in high-risk populations and the importance of accurate risk assessment models, an accurate assessment of FGT is needed.
Previous studies have utilized deep learning to perform automated segmentation of FGT on breast MRI but with significant drawbacks, mainly regarding private dataset usage12,13,14,15,16. The lack of annotated large-scale datasets has been widely recognized as one of the most significant challenges in medical imaging segmentation17,18. Previous works also do not clearly define the annotation methods used and there can be ambiguity when defining the boundaries of tissues, especially the breast which has limited surrounding anatomical landmarks. Providing full transparency facilitates the reproduction of results and allows others to further improve on the achieved results. Furthermore, to our knowledge, previous studies do not take blood vessels into account when performing manual segmentations. Even in pre-contrast MRI sequences, large blood vessels, namely branches of the internal mammary and lateral thoracic arteries, can still appear hyperintense and therefore be confused with FGT. The initial branches of these larger arteries can occupy a significant amount of volume within the breast, which can confound FGT calculations.
In our study, we address prior limitations to improve the assessment of FGT on breast MRI with our publicly available dataset and segmentation method. The contributions of the study are as follows: (1) We provide a detailed and clear methodology to annotate breast MRI that allows for a higher level of reproducibility with a clearer, quantitative definition of breast density. In addition to being explicit, our methodology is the first to clearly consider vessels as separate structures. (2) We generated an extensive set of detailed, pixel-wise, fully three-dimensional annotations, a result of hundreds of hours spent annotating by our team. The average annotation time per case is 8 hours. The annotations were all modified and/or approved by breast fellowship-trained radiologists. (3) The annotations are publicly available at The Cancer Imaging Archive website along with the corresponding MRIs previously made available by our group. (4) We developed and evaluated an accurate deep learning model to segment the breast, FGT, and blood vessels. (5) Finally, we made the segmentation model publicly available so that others can apply it to their own datasets.
Methods
Dataset
We randomly selected 100 cases from the publicly available Duke-Breast-Cancer-MRI dataset which is available at https://doi.org/10.7937/TCIA.e3sv-re9319. The full dataset contains MRI sequences from 922 biopsy-confirmed invasive breast cancer patients at the Duke University Medical Center.
Our study was determined to be exempt from review by the Duke Health Institutional Review Board. The review board also determined that receiving informed consent is waived as we are using a publicly available database. All methods were performed accordance with the relevant guidelines and regulations.
The MRIs were obtained from patients in the prone position using a 1.5 T or 3.0 T scanner (GE Healthcare and Siemens). Axial images were acquired with a voxel size ranging from 320 × 320 × 144 to 512 × 512 × 200 and a resolution ranging from 0.6 × 0.6 × 1.0 mm3 to 1.1 × 1.1 × 1.2 mm3. We used the T1-weighted fat-suppressed pre-contrast sequence for our study as it provides ample contrast between FGT and fatty tissue while minimizing visible blood vessels.
The 100 patients were randomly split into training, validation, and test sets with a 70%/15%/15% split. The validation set was used during model development and evaluation to improve model performance. The test set was used only after model development was completed and no changes were made to improve performance on the test set data after its use for evaluation.
Image annotations
All MRIs were manually annotated and reviewed using 3D Slicer. There were three target tissues for annotation: breast, FGT, and blood vessels. The annotators and reviewers were given the following instructions:
-
1.
Breast annotation: Trace the contours of the breast, excluding inner anatomical structures, such as the chest wall muscles and sternum. The superior edge of the annotation should stop approximately at the level of the clavicle. The inferior edge of the annotation should stop at the inframammary fold.
-
2.
FGT annotation: Mark all areas of FGT that do not appear to be blood vessels. Biopsy clips and lymph nodes should be excluded. All FGT should be within the breast.
-
3.
Blood vessel annotation: Mark all readily apparent blood vessels. All blood vessels should be within the breast.
The annotators consisted of one postdoctoral fellow, one upper-level medical student, and two undergraduate students who were all given the same annotation instructions, examples, and instructions to use the software. All annotators had a minimum of 5 hours spent learning how to properly perform annotations and reviewing practice annotations to ensure that instructions were followed properly. The medical student annotator reviewed and, if needed, edited all annotations. All annotations were then reviewed and, if needed, edited by one of three board-certified, fellowship-trained breast radiologists at our institution.
The annotations are publicly available online at https://doi.org/10.7937/TCIA.e3sv-re93, under “Data Access”, “Supplemental Segmentation”, “3D Breast and FGT MRI Segmentation Supplemental Data”. There are additional segmentations included in the database, but this study exclusively used the more extensive “3D Breast and FGT MRI Segmentation Supplemental Data” that were created using the guidelines described above.
Data processing and model architecture
Multiple steps were taken to improve data uniformity. First, spatial information from DICOM files was used to ensure all MRI volumes and their accompanying annotations were in the same position. Volumes or annotations that were rotated and/or flipped incorrectly were aligned accordingly. Following this, we performed basic preprocessing of the data. First, we capped the extreme values in the image (below 0.1 percentile and above 99.9 percentile) and normalized the intensity values using a z-score, as shown in Eq. (1).
Where \(x\) is the intensity value of the individual voxel, \(\mu\) is the mean intensity value within the MRI volume, and \(\sigma\) is the standard deviation for intensity values within the MRI volume.
We selected the U-Net architecture as it has been successfully applied to many medical imaging segmentation tasks20,21,22. The 3D U-Net architecture features a fully connected convolutional residual network with a contraction and expansion phase. During the contraction phase, each step applies two 3 × 3 × 3 convolutional layers with batch normalization and a rectified linear unit (ReLU) followed by a 2 × 2 × 2 max pooling layer with a stride 2. During the expansion phase, each step up-samples the input, concatenates it with its corresponding feature map from the contracting path, then applies two 3 × 3 × 3 convolutional layers with batch normalization and a ReLU. We also used a 2D U-Net that featured a similar architecture with one less dimension used in all layers.
Two models were trained for our segmentation task. The first model, named Breast U-Net, used the MRI volume to output a binary prediction on each voxel to perform a segmentation of the breast. For the second model, named FGT-Vessel U-Net, we incorporated information on the predicted location of the FGT and vessels, since they will always be contained within the breast. To accomplish this, we included the outputs of Breast U-Net as an additional channel alongside the MRI volume. An overview of the model architecture and data input is shown in Figs. 1 and 2.
We tested two segmentation methods: one with 2D inputs and one with 3D inputs. For 2D inputs, each slice of the volume was inputted into the model individually. For 3D inputs, we developed different methods of inputting the data into the models. To predict the breast, the full MRI volume was input into the model after downsizing to 144 × 144 × 96. To predict the FGT and blood vessels, we divided the 3D MRI volume into 3D sub-volumes of size 96 × 96 × 96. To obtain each sub-volume, we used a random sub-sampling method, where all areas of the volume had an equal chance to be sampled, to generate sub-volumes for model input. Data augmentations that added random motion, noise, and bias fields were used in both models. To obtain segmentations for a full volume, 192 evenly spaced sub-volumes are predicted from the full volume. The prediction for a voxel in the full volume was the mean of all predictions in sub-volumes that contain it. These different 3D approaches for breast and FGT were selected based on experimental results using the training and validation data.
To select hyperparameters, we experimentally evaluated various total epochs, learning rates, learning rate decays, batch sizes, sub-volume sizes, and sub-volume counts using the validation set. Total epochs, learning rate, and learning rate decay were adjusted with the aim of reducing overfitting. We also compared using cross-entropy and dice loss functions. Dice loss in particular was used as blood vessels occupy a small number of voxels and therefore face a class imbalance issue, which can be overcome using dice loss or similar loss functions23,24. We chose the model that had the best DSC values for each task after testing various hyperparameter combinations using the validation set. We used the Adam optimizer during gradient back-propagation.
All code and trained models used in the study are publicly available at: https://github.com/mazurowski-lab/3D-Breast-FGT-and-Blood-Vessel-Segmentation.
Model evaluation
The models were evaluated using the Dice Similar Coefficient (DSC) as it is commonly used to evaluate the performance of image segmentation methods. The minimum DSC value of 0 indicates no overlap between segmentations and the maximum DSC value of 1 indicates complete overlap. For breast segmentation predictions, we applied a sigmoid function to model outputs and thresholded values at 0.5 to generate breast masks. For FGT and blood vessel segmentation predictions, we applied a softmax function to model outputs and assigned each voxel the class with the highest value (FGT, blood vessel, or neither). DSC scores were then calculated for each predicted segmentation individually compared to their respective annotations. The final model evaluation was performed using the test set. The Mann-Whitney test was utilized to compare DSC scores of the test set data.
We used the Pytorch open-source machine learning framework. Our program was run on multiple GPUs (NVIDIA RTX 3090 24GB).
Additional analysis
Three fellowship-trained and board-certified radiologists were asked to assess the breast density of each image in the test set. Breast density is defined in the Breast Imaging-Reporting and Data System (BI-RADS) Atlas as four categories: (a) almost entirely fat, (b) scattered FGT, (c) heterogeneous FGT, and (d) extreme FGT25. We compared assessments by each pair of radiologists using Cohen’s kappa.
FGT percentage was used to assess the model’s predictions in relation to breast density assessments and is calculated using Eq. (2). We used Pearson’s correlation coefficient to compare FGT percentages and radiologist breast density assessments.
Results
An overview of the demographics and basic characteristics of the 100 patients used in model development and testing is shown in Table 1.
The average DSC values for breast, FGT, and blood vessel segmentation can be found in Table 2. The following describes model performance on the test set. The 3D Breast U-Net achieved a DSC score of 0.92 for breast. The following 3D FGT-Vessel U-Net that combined MRI data with predicted breast masks achieved a DSC score of 0.86 for FGT and 0.65 for blood vessels. The 3D FGT-Vessel U-Net that utilized MRI input alone achieved a DSC score of 0.86 for FGT and 0.61 for blood vessels. Between these two 3D FGT-Vessel U-Net, there was no difference between the performance of FGT (p = 0.53) or blood vessel (p = 0.25) segmentation. The 2D Breast U-Net found a DSC score of 0.95 for breast which was statistically different from the 3D performance (p ≤ 0.001). The following 2D FGT-Vessel U-Net that combined MRI data with predicted breast masks achieved a DSC score of 0.84 for FGT and 0.53 for blood vessels. The same model with MRI input alone found a DSC score of 0.84 for FGT and 0.37 for blood vessels. The 2D FGT-Vessel U-Net version had no difference between FGT segmentation performance (p = 0.53) but a difference was present between blood vessel predictions (p = 0.001). Comparing 3D and 2D FGT-Vessel U-Nets that utilized both predicted breast masks and MRI data input, there was no difference between FGT segmentation performance (p = 0.18) but a difference was present between blood vessel performance (p < 0.001).
On average, completing the segmentation of a case took 9.6s for the 3D Breast U-Net and 72.3s for the 3D FGT-Vessel U-Net. Sample outputs, taken from the validation set, of the best performing model are shown in Fig. 3.
Table 3 demonstrates the confusion matrices for the breast density assessments of each pair of the 3 radiologists on the test set images. The Cohen’s kappa coefficient was 0.38 for radiologists 1 and 2, 0.65 for radiologists 1 and 3, and 0.43 for radiologists 2 and 3. The cumulative breast density assessments by the 3 radiologists on the 15 test set images were the following: 1 (2.2%) almost entirely fat, 25 (55.6%) scattered FGT, 14 (31.1%) heterogeneous FGT, and 5 (11.1%) extreme FGT.
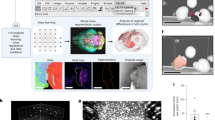
Figure 4 compares ground-truth FGT percentages with predicted FGT percentages in the test set. The average difference between the ground-truth FGT percentage and model-predicted FGT percentage was 1.1% and the Pearson coefficient comparing these percentages was 0.95.
Scatter plot of ground truth FGT percentages versus predicted FGT percentages for images in the test set. The dashed line represents perfect correlation between the two FGT percentages. The average difference between ground-truth and predicted FGT percentage was 1.1% and the Pearson coefficient was 0.95.
Figure 5 compares the breast density assessments of the 3 radiologists and the percentage of FGT present within the breast based on model predictions on the test set. The Pearson coefficient was 0.80 for ground-truth FGT percentages versus radiologist assessments and 0.75 for model-predicted FGT percentages versus radiologist assessments.
Discussion
In our study, we developed two CNNs to accurately segment breast, FGT, and blood vessels using pre-contrast fat-suppressed T1-weighted MRI volumes. We created a radiologist-reviewed dataset using well-defined criteria to train these models. The dataset, code, and models were made publicly available.
Our deep learning method achieved a segmentation performance similar to or greater than previous methods. There have been a variety of studies that have used non-deep learning techniques to segment the breast and FGT. The most common methods are atlas-based fuzzy C-means methods and level-set-based methods, achieving DSC values ranging from 0.61 to 0.97 for breast segmentation and DSC values ranging from 0.80 to 0.84 for FGT segmentation26,27,28,29,30,31,32,33,34. There are also many studies that utilized deep learning, also employing either 2D or 3D U-Net models or similarly designed CNNs. They found DSC values ranging from 0.86 to 0.94 for breast segmentation and DSC values ranging from 0.83 to 0.92 for FGT segmentation12,13,14,15,16. Past works have compared a single U-net versus consecutive U-nets, similar to our work, for segmentation of breast and FGT, with Dalmış et al demonstrating superior performance with a single U-net12. However, our work includes blood vessel segmentation which is rarer tissue within the breast to segment, which may emphasize the need for consecutive U-nets to provide a breast mask for context.
Our analysis which compared breast density assessments between radiologists showed that there can be a large amount of variability between density assessments, with Cohen’s kappa values of 0.38, 0.65, and 0.43. This further illustrates the need for a standardized method of breast density assessment. FGT percentage is the calculation that can be employed from our model’s predictions of breast and FGT. When comparing FGT percentages and radiologist assessments, we demonstrated Pearson’s correlation coefficients of 0.80 and 0.75 for ground-truth and model-predicted FGT percentages, respectively. These indicate that i) FGT percentages are moderately correlated with breast density assessments, as expected, and ii) our model is able to retain a very similar level of correlation.
Although we achieved similar DSC values for breast and FGT segmentation to previous studies, a unique distinction of our study is the addition of blood vessel segmentations. Within the breast, there are other structures that can appear similar to FGT as they all appear similarly hyperintense on T1w fat-suppressed pre-contrast images. These hyperintense structures are mainly blood vessels and lymph nodes. Lymph nodes are most commonly present in the axillary region but intramammary lymph nodes can occur within the breast. When present, intramammary lymph nodes are usually small, few in number, and represent a very small proportion of the breast. As a result, we excluded them from FGT annotations and did not perform a separate annotation for them. In contrast, blood vessels are always present in the breast and can represent a sizeable proportion of the breast. In our annotations, out of all voxels that were labeled as either FGT or blood vessels, blood vessels accounted for 5.7% of the volume. As far as we are aware, previous studies that performed FGT segmentation did not account for blood vessels and therefore may be overestimating FGT by approximately 5.7% if blood vessels were included in FGT annotations.
The main limitation we faced in our study was the number of fully annotated MRI volumes. Despite using techniques such as intensity thresholding and 3D annotating tools, annotating a single MRI volume took approximately 8 hours. However, since we randomly selected patients, we believe that our dataset provides a heterogenous cohort for analysis with patients in a variety of scanners and with a variety of demographics. Furthermore, the size of our test set may have limited our comparison of models. There was no statistically significant difference between 3D models that included and did not include breast mask predictions, but such a difference was present when comparing 2D models. This is likely due to the small number of 3D volumes (n = 15) but many 2D slices (n = 2578).
Conclusion
Our study performed accurate segmentation of breast, FGT, and blood vessels using two consecutive 3D U-Nets. Additionally, all code and data used are provided online for public use. Our deep learning method has the potential to objectively evaluate breast density in women to improve breast cancer risk assessments.
Data availability
The MRI studies used in this article are from the Duke Breast Cancer MRI dataset which is available online at https://doi.org/10.7937/TCIA.e3sv-re93, under “Data Access”, “Supplemental Segmentation”, “3D Breast and FGT MRI Segmentation Supplemental Data”. All code and trained models used in the study are publicly available at: https://github.com/mazurowski-lab/3D-Breast-FGT-and-Blood-Vessel-Segmentation.
References
Advani, S. M. et al. Association of breast density with breast cancer risk among women aged 65 years or older by age group and body mass index. JAMA Netw. Open 4, e2122810. https://doi.org/10.1001/jamanetworkopen.2021.22810 (2021).
Boyd, N. F., Martin, L. J., Yaffe, M. J. & Minkin, S. Mammographic density and breast cancer risk: Current understanding and future prospects. Breast Cancer Res 13, 223. https://doi.org/10.1186/bcr2942 (2011).
McCormack, V. A. & dos Santos Silva, I. Breast density and parenchymal patterns as markers of breast cancer risk: A meta-analysis. Cancer Epidemiol. Biomark. Prev. 15, 1159–1169. https://doi.org/10.1158/1055-9965.Epi-06-0034 (2006).
Vachon, C. M. et al. Mammographic density, breast cancer risk and risk prediction. Breast Cancer Res. 9, 217. https://doi.org/10.1186/bcr1829 (2007).
Brandt, K. R. et al. Comparison of clinical and automated breast density measurements: Implications for risk prediction and supplemental screening. Radiology 279, 710–719. https://doi.org/10.1148/radiol.2015151261 (2016).
Alomaim, W. et al. Subjective versus quantitative methods of assessing breast density. Diagnostics (Basel) 10, 331. https://doi.org/10.3390/diagnostics10050331 (2020).
Tagliafico, A. et al. Mammographic density estimation: Comparison among BI-RADS categories, a semi-automated software and a fully automated one. Breast 18, 35–40. https://doi.org/10.1016/j.breast.2008.09.005 (2009).
Redondo, A. et al. Inter- and intraradiologist variability in the BI-RADS assessment and breast density categories for screening mammograms. Br. J. Radiol. 85, 1465–1470. https://doi.org/10.1259/bjr/21256379 (2012).
Irshad, A. et al. Effects of changes in BI-RADS density assessment guidelines (fourth versus fifth edition) on breast density assessment: Intra- and interreader agreements and density distribution. Am. J. Roentgenol. 207, 1366–1371. https://doi.org/10.2214/ajr.16.16561 (2016).
Chiarelli, A. M. et al. Performance measures of magnetic resonance imaging plus mammography in the high risk ontario breast screening program. J. Natl. Cancer Inst. 112, 136–144. https://doi.org/10.1093/jnci/djz079 (2020).
Sardanelli, F. et al. Sensitivity of MRI versus mammography for detecting foci of multifocal, multicentric breast cancer in Fatty and dense breasts using the whole-breast pathologic examination as a gold standard. AJR Am. J. Roentgenol. 183, 1149–1157. https://doi.org/10.2214/ajr.183.4.1831149 (2004).
Dalmış, M. U. et al. Using deep learning to segment breast and fibroglandular tissue in MRI volumes. Med. Phys. 44, 533–546. https://doi.org/10.1002/mp.12079 (2017).
Zhang, Y. et al. Automatic breast and fibroglandular tissue segmentation in breast MRI using deep learning by a fully-convolutional residual neural network U-Net. Acad. Radiol. 26, 1526–1535. https://doi.org/10.1016/j.acra.2019.01.012 (2019).
Ivanovska, T. et al. A deep learning framework for efficient analysis of breast volume and fibroglandular tissue using MR data with strong artifacts. Int. J. Comput. Assist. Radiol. Surg. 14, 1627–1633. https://doi.org/10.1007/s11548-019-01928-y (2019).
Ha, R. et al. Fully automated convolutional neural network method for quantification of breast MRI fibroglandular tissue and background parenchymal enhancement. J. Digit. Imaging 32, 141–147. https://doi.org/10.1007/s10278-018-0114-7 (2019).
van der Velden, B. H. M., Janse, M. H. A., Ragusi, M. A. A., Loo, C. E. & Gilhuijs, K. G. A. Volumetric breast density estimation on MRI using explainable deep learning regression. Sci. Rep. 10, 18095. https://doi.org/10.1038/s41598-020-75167-6 (2020).
Altaf, F., Islam, S. M. S., Akhtar, N. & Janjua, N. K. Going deep in medical image analysis: Concepts, methods, challenges, and future directions. IEEE Access 7, 99540–99572. https://doi.org/10.1109/access.2019.2929365 (2019).
Chan, H. P., Samala, R. K., Hadjiiski, L. M. & Zhou, C. Deep learning in medical image analysis. Adv. Exp. Med. Biol. 1213, 3–21. https://doi.org/10.1007/978-3-030-33128-3_1 (2020).
Saha, A. et al. A machine learning approach to radiogenomics of breast cancer: A study of 922 subjects and 529 DCE-MRI features. Br. J. Cancer 119, 508–516. https://doi.org/10.1038/s41416-018-0185-8 (2018).
Liu, X., Song, L., Liu, S. & Zhang, Y. A review of deep-learning-based medical image segmentation methods. Sustainability 13, 1224. https://doi.org/10.3390/su13031224 (2021).
Du, G., Cao, X., Liang, J., Chen, X. & Zhan, Y. Medical image segmentation based on u-net: A review. J. Imaging Sci. Technol. 64, 1–12 (2020).
Pérez-García, F. fepegar/unet: PyTorch implementation of 2D and 3D U-Net. Zenodo https://doi.org/10.5281/zenodo.3697931 (2020).
Zhao, R. et al. in 2020 IEEE International Conference on Data Mining (ICDM). 851–860.
Sudre, C. H., Li, W., Vercauteren, T., Ourselin, S. & Jorge Cardoso, M. in Deep Learning in Medical Image Analysis and Multimodal Learning for Clinical Decision Support. (eds M. Jorge Cardoso et al.) 240–248 (Springer International Publishing).
D’Orsi, C. J. 2013 ACR BI-RADS Atlas: Breast Imaging Reporting and Data System (American College of Radiology, 2014).
Wu, S., Weinstein, S. P., Conant, E. F. & Kontos, D. Automated fibroglandular tissue segmentation and volumetric density estimation in breast MRI using an atlas-aided fuzzy C-means method. Med. Phys. 40, 122302. https://doi.org/10.1118/1.4829496 (2013).
Ivanovska, T. et al. A level set based framework for quantitative evaluation of breast tissue density from MRI data. PLoS ONE 9, e112709. https://doi.org/10.1371/journal.pone.0112709 (2014).
Wu, S., Weinstein, S. & Kontos, D. Atlas-based probabilistic fibroglandular tissue segmentation in breast MRI. Med. Image Comput. Comput. Assist. Interv. 15, 437–445. https://doi.org/10.1007/978-3-642-33418-4_54 (2012).
Razavi, M. et al. in Image Analysis and Processing — ICIAP 2015. (eds Vittorio Murino & Enrico Puppo) 528–536 (Springer International Publishing).
Gubern-Mérida, A., Kallenberg, M., Mann, R. M., Martí, R. & Karssemeijer, N. Breast segmentation and density estimation in breast MRI: A fully automatic framework. IEEE J. Biomed. Health Inform. 19, 349–357. https://doi.org/10.1109/JBHI.2014.2311163 (2015).
Takhtawala, R. et al. Automated artificial intelligence quantification of fibroglandular tissue on breast MRI. J. Clin. Oncol. 37, e12071–e12071. https://doi.org/10.1200/JCO.2019.37.15_suppl.e12071 (2019).
Huo, L. et al. Segmentation of whole breast and fibroglandular tissue using nnU-Net in dynamic contrast enhanced MR images. Magn. Reson. Imaging 82, 31–41. https://doi.org/10.1016/j.mri.2021.06.017 (2021).
Ma, X. et al. Automated fibroglandular tissue segmentation in breast MRI using generative adversarial networks. Phys. Med. Biol. 65, 105006. https://doi.org/10.1088/1361-6560/ab7e7f (2020).
Wei, D. et al. Fully automatic quantification of fibroglandular tissue and background parenchymal enhancement with accurate implementation for axial and sagittal breast MRI protocols. Med. Phys. 48, 238–252. https://doi.org/10.1002/mp.14581 (2021).
Author information
Authors and Affiliations
Contributions
C.O.L. led the study design, created and reviewed most of the annotations, and wrote the codebase. M.H. assisted with the study creation and created many annotations. E.R.K. and E.J.K. created many of the annotations. H.D. and H.G. reviewed the codebase and tested various methods to improve model performance. L.J.G., R.W., and D.A.L. reviewed the annotations and provided clinical input. M.A.M. oversaw the study and assisted with all aspects of the study design. C.O.L. drafted the manuscript and all authors contributed to the finalized version.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Lew, C.O., Harouni, M., Kirksey, E.R. et al. A publicly available deep learning model and dataset for segmentation of breast, fibroglandular tissue, and vessels in breast MRI. Sci Rep 14, 5383 (2024). https://doi.org/10.1038/s41598-024-54048-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-024-54048-2
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.