Abstract
Mus minutoides is one of the smallest mammals worldwide; however, the regulatory mechanisms underlying its dwarfism have not been examined. Therefore, we aimed to establish M. minutoides induced pluripotent stem cells (iPSCs) using the PiggyBac transposon system for applications in developmental engineering. The established M. minutoides iPSCs were found to express pluripotency markers and could differentiate into neurons. Based on in vitro differentiation analysis, M. minutoides iPSCs formed embryoid bodies expressing marker genes in all three germ layers. Moreover, according to the in vivo analysis, these cells contributed to the formation of teratoma and development of chimeric mice with Mus musculus. Overall, the M. minutoides iPSCs generated in this study possess properties that are comparable to or closely resemble those of naïve pluripotent stem cells (PSCs). These findings suggest these iPSCs have potential utility in various analytical applications, including methods for blastocyst completion.
Similar content being viewed by others
Introduction
Mus minutoides, also known as the African pygmy mouse, has recently gained attention as a domestic pet. M. minutoides is considered one of the smallest mammals and belongs to the same genus as the common laboratory mouse (Mus musculus). Despite their common biological characteristics, such as lifespan and gestation period, M. minutoides is significantly smaller than M. musculus, measuring only one-tenth of the body size of M. musculus1. As previous studies involved the performance of phylogenetic analysis and focused on the sex-determination patterns and viruses carried by M. minutoides2,3,4,5,6, the dwarfism of M. minutoides has not been studied in detail.
In mammals, the Gh–Igf1 axis influences postnatal body size in mammals7,8. In our previous study, we analyzed the growth hormones in M. minutoides9; however, a study exploring the characteristics of growth-related factors, particularly growth hormones, has not been performed. The generation of genetically modified animals, such as knockout mice, is effective for conducting body size regulation-related molecular genetic analyses. However, as M. minutoides is not a laboratory animal, its use is difficult, and techniques for developmental engineering have not been established.
Pluripotent stem cells (PSCs) are defined as embryonic stem cells (ESCs) derived from the inner cell mass of blastocysts and induced pluripotent stem cells (iPSCs) reprogrammed from differentiated cells. iPSCs were first produced in M. musculus in 200610. M. minutoides is vulnerable to transport-related stress. Moreover, as only a few investigators use M. minutoides, obtaining individuals is a challenging task. In prior studies with M. minutoides, developmental engineering techniques, particularly superovulation, egg collection, and fertilized egg culture systems, were not employed. Consequently, obtaining fertilized eggs from M. minutoides to establish ESCs poses a significant challenge. Therefore, this study focused on iPSCs. iPSCs have garnered attention in regenerative medicine and have been applied to developmental process analysis and understanding of pathological conditions11,12,13. The potential applications of iPSCs in conserving endangered wildlife have also been considered in recent years14,15. iPSCs have been successfully induced in mice, rats16, humans17, cattle18, pigs19, goats20, sheep21, Bactrian camel22, Tokuno-shima spiny rats23, American plain voles24, and other rodents, including the naked mole rat25.
In this study, we aimed to establish M. minutoides iPSCs derived from M. minutoides fibroblasts. These iPSCs exhibit the requisite attributes of pluripotent stem cells and actively participate in ontogenesis. Overall, M. minutoides iPSCs are an important resource that may substantially improve our understanding of dwarfism, which is the main characteristic trait of M. minutoides.
Results
Establishment of the iPSC lines from fibroblasts
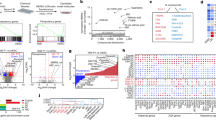
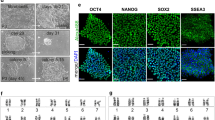
The four iPSC-inducing plasmid constructs used in this study are shown in Fig. 1a. Electroporation was employed as the plasmid transfer method to establish the M. minutoides iPSC strains (Fig. 1b). Primary fibroblasts were established from the tail tip of M. minutoides. The day of electroporation was designated as day 0; the M. minutoides fibroblasts transfected with the iPSC-inducing plasmid formed colonies and exhibited EGFP+ signals on feeder cells by day 7 (Fig. 1c). Upon subsequent passaging, the cells developed naïve ESC-like colonies when cultured with DOX. On day 12, DOX supplementation was discontinued; however, naïve ESC-like colonies were still observed on day 33 after several passages (Fig. 1d). A total of 16 M. minutoides iPSC lines were produced, with no apparent differences in growth rate or colony morphology. To confirm the origin of iPSCs from M. minutoides, genomic DNA was extracted from a cell line that had undergone at least 10 additional passages, and PCR was performed. M. minutoides and M. musculus genomes were differentiated based on the disparity in the length of the Ghr promoter region (unpublished data). PCR products from the M. minutoides genome were detected, confirming that all iPSC lines established in this study originated from M. minutoides (Fig. 1e).
Establishment of Mus minutoides induced pluripotent stem cells (iPSCs). (a) iPSC plasmid constructs. (b) Timeline of the establishment of M. minutoides iPSCs. (c) Fluorescence microscopic image of M. minutoides iPSC-like cells on day 7. Bar: 100 µm. (d) Colonies of M. minutoides iPSCs on days 0, 7, and 33 after electroporation. Bar: 100 µm. (e) PCR identification of M. minutoides-derived cells using genomic DNA. "mouse" and "apm” represent genomic DNA extracted from M. musculus and M. minutoides, respectively.
Characteristics of M. minutoides iPSCs
To assess the pluripotency of M. minutoides iPSCs, we initially examined the activity of ALP, a commonly used stem cell marker. Notably, almost all colonies established in this study were ALP positive (Fig. 2a). cDNA was then synthesized using total RNA extracted from M. minutoides iPSCs, and the expression of the pluripotency marker genes was analyzed using RT-PCR. The expression of the pluripotency marker genes, Eras and Fgf4, was observed in conjunction with the genes delivered by the iPSC-inducing plasmids despite not being introduced through electroporation (Fig. 2b). To confirm the presence of Fgf4 in M. minutoides iPSCs, we performed RT-PCR using cDNA and primers specific to a region of the Fgf4 gene sequence. Sequencing of the amplified product revealed that several nucleotide sequences differed from those of M. musculus (Supplemental Fig. 1a); however, the predicted amino acid sequence derived from the nucleotide sequence was identical to that of M. musculus (Supplemental Fig. 1b). Furthermore, we quantitatively evaluated the expression of the pluripotency marker gene, Nanog, a representative marker, in multiple established cell lines using RT-qPCR. Nanog expression levels varied among the cell lines (Fig. 2c). In addition, we examined the expression of other pluripotency marker genes (Oct3/4, Sox2, Eras, and Fgf4) in M. minutoides iPSCs derived from lines 6 and 10, in which Nanog expression was comparable to that in M. musculus ESCs. The expression levels of these markers, particularly Fgf4, were significantly higher in M. minutoides iPSCs than in M. musculus ESCs (Fig. 2d). Correlation analysis of the gene expression levels revealed a weak positive correlation between Eras and other genes (Supplemental Fig. 2). We also examined the expression and localization of the pluripotency markers, NANOG and OCT3/4, using fluorescence immunostaining. Only EGFP-positive M. minutoides iPSC colonies stained positive for NANOG and OCT3/4 (Fig. 2e).
Analysis of pluripotency in Mus minutoides induced pluripotent stem cells (iPSCs). (a) Microscopic (left) and macroscopic image (right) of alkaline phosphatase-stained M. minutoides iPSCs (apm iPSC) in a 60-mm dish. M. minutoides fibroblast (apm fibroblast) was used as the negative control. Bar: 100 µm. (b) Expression analysis of pluripotency marker genes based on RT-PCR. Gapdh was used as the positive control. “mouse” and “apm” represent M. musculus and M. minutoides, respectively. (c) Relative Nanog expression in 16 established M. minutoides iPSC lines. Gapdh was used as the endogenous control, and mouse_ESC expression level was defined as 1.0. “mouse” and “apm” represent M. musculus and M. minutoides, respectively. (d) Relative Nanog, Oct3/4, Sox2, Eras, and Fgf4 expression levels in M. minutoides iPSCs (apm_iPSC-6, apm_iPSC-10). Gapdh was used as the endogenous control, and mouse_ESC expression level was defined as 1.0. “mouse” and “apm” represent M. musculus and M. minutoides, respectively. (e) Fluorescence immunostaining with the anti-NANOG and -OCT3/4 antibodies in M. minutoides iPSCs (apm iPSC). M. minutoides fibroblast (apm fibroblast) was used as the negative control. Bar: 100 µm.
To obtain a more in-depth clarification of gene expression in M. minutoides iPSCs, RNA-seq analysis was performed using iPSCs and fibroblasts. The 29,437 expressed genes exhibited significant differences in gene expression status between M. minutoides iPSCs and fibroblasts. The expression levels of 6503 genes varied significantly by more than twofold, such as Nanog, Oct3/4, Sox2, Eras, and Fgf4, which were upregulated based on RT-qPCR (Fig. 3a). Enrichment analysis of the differentially expressed genes (DEGs) from M. minutoides iPSCs and fibroblasts revealed several Gene Ontology (GO) terms related to pluripotency pathways and cell differentiation (Fig. 3b). Of the 292 genes in WP1763 with the highest rank in the enrichment analysis, 102 were upregulated in iPSC_6 and iPSC_10 compared to the levels found in fibroblasts (Supplemental Fig. 3a). Subsequent analyses were performed using mouse iPSCs (mouse_iPS_1 and mouse_iPS_2) obtained from the GEO database. Principal component analysis (PCA) revealed differences in the gene expression status of each cell group (Supplemental Fig. 3b).
RNA-seq analysis of M. minutoides iPS cells and fibroblasts. (a) Volcano plot of differentially expressed genes in M. minutoides iPSCs and fibroblasts. (b) Enrichment analysis of the upregulated genes in M. minutoides iPSCs, obtained using Metascape. (c) (Left) Venn diagram showing the twofold upregulated genes in M. minutoides and M. musculus iPSCs compared to fibroblasts. (Right) Enrichment analysis of the upregulated genes in M. minutoides and M. musculus iPSCs was obtained using Metascape. (d) (Left) Venn diagram showing the twofold downregulated genes in M. minutoides and M. musculus iPSCs compared to fibroblasts. (Right) Enrichment analysis of the downregulated genes in M. minutoides and M. musculus iPSCs was obtained using Metascape.
A comparison of the fibroblast and iPSC gene expression profiles revealed the shared upregulation of 631 genes in M. minutoides and M. musculus iPSCs (Fig. 3c). Table 1 shows some of the upregulated genes that are relevant to pluripotency in M. minutoides, M. musculus, or both iPSC types. Enrichment analysis of these genes revealed significant associations with GO terms related to pluripotency and the maintenance of stem cell populations, as illustrated in Fig. 3c. Conversely, the expression levels of 582 genes decreased in M. minutoides and M. musculus iPSCs, and this gene cluster encompassed numerous GO terms associated with fibroblast and connective tissue functions (Fig. 3d).
Differentiation potential of iPSCs
To assess the differentiation potential of M. minutoides iPSCs, we conducted in vitro differentiation experiments targeting the neuronal lineages. Although a considerable number of cells underwent apoptosis during the early stages of differentiation, the expression of neuronal marker genes (Tuj1, Nestin) was observed 14 days post-induction (Fig. 4a). Fluorescent immunostaining also revealed elongated processes originating from NESTIN-positive neurons (Fig. 4b). EB formation was examined in M. minutoides iPSCs. These iPSCs successfully formed EGFP-expressing EBs after 5 days of culture under EB-forming conditions (Fig. 4c,d). Using RT-PCR, we found that the EBs and cells migrating from the EB masses differentiated into three germ layers, the ectoderm, mesoderm, and endoderm, as well as the trophic ectoderm, which contributes to extraembryonic tissues (Fig. 4e).
In vitro differentiation potential analysis of Mus minutoides induced pluripotent stem cells (iPSCs). (a) RT-PCR analysis of neuronal differentiation-induced M. minutoides iPSCs. Gapdh was used as the positive control. (b) Fluorescence immunostaining in neuronal differentiation-induced M. minutoides iPSCs with the anti-NESTIN antibody. Bar: 50 µm. (c) Bright field image of embryoid bodies (EBs) formed from M. minutoides iPSCs on day 4 of floating culture. Bar: 200 µm. (d) Fluorescence microscopic image of M. minutoides EBs on day 5 of floating culture. Bar: 100 µm. (e) RT-PCR analysis of EBs (EB d10, EB d20) and EB migrating cells (EB-cells) cultured for 10 or 20 days. Gapdh was used as the positive control. (f) Fluorescence microscopic image of M. minutoides EB-cells differentiated into cardiomyocytes. Bar: 100 µm.
We also performed an adhesion culture of EBs derived from M. minutoides iPSCs. A population of cells with filament-like structures and self-beating behavior, characteristic of cardiomyocytes, was observed within the layered sheets of adherent cells (Fig. 4f and Supplemental Movie 1). Based on RT-PCR using this cell population, Tnnt2, a specific marker of cardiomyocytes, was robustly expressed (Fig. 4e).
Teratoma
To evaluate the in vivo differentiation potential of M. minutoides iPSCs, M. minutoides iPSCs were subcutaneously injected into nude mice with specific immune cell dysfunction. Two weeks after the injection, teratoma formation was observed (Fig. 5a). Subsequent histological analysis revealed that these teratomas contained tissues representing all three germ layers: neural tissue (ectoderm), muscle (mesoderm), and intestinal epithelium (endoderm) (Fig. 5b). To determine the genomic origin of the teratomas, we extracted genomic DNA from paraffin-embedded sections to detect M. minutoides iPSC colonies. Our analyses confirmed that the genomic DNA of teratomas corresponded to that of M. minutoides (Fig. 5c).
In vivo differentiation and contribution potential analysis of Mus minutoides induced pluripotent stem cells (iPSCs). (a) Teratoma formed from M. minutoides iPSCs. (b) Ectoderm (neural epithelium), mesoderm (muscle), and endoderm (gastrointestinal epithelium) observed in the teratoma. Bar: 50 µm. (c) Identification of the cell origin based on PCR using genomic DNA. “mouse" and "apm" represent genomic DNA extracted from M. musculus and M. minutoides, respectively. (d) The upper and lower images represent 3- and 11-day-old mice, respectively. The coat color (wild color) from M. minutoides is observed with the coat color (white) from ICR (M. musculus). Representative chimeric mice are marked with *.
Chimeric mice
To evaluate the developmental contribution of M. minutoides iPSCs in animals, we used the blastocyst complementation method to generate chimeric mice. By integrating M. minutoides iPSCs into M. musculus (ICR mouse) blastocysts, chimeric pups with M. minutoides coats were successfully generated and allowed to develop for two weeks (5/14 pups) (Fig. 5d).
Discussion
In the present study, we successfully generated iPSCs from M. minutoides fibroblasts, which formed dome-shaped colonies. In addition, we demonstrated that M. minutoides iPSCs are pluripotent and can contribute to the generation of chimeric mice when transplanted into the embryos of M. musculus, a closely related species within the Mus genus.
To establish M. minutoides iPSCs, we used a combination of seven genes, including Yamanaka factors (Oct3/4, Sox2, Klf4, c-Myc), Glis1, Nanog, and Lin28a, which enhance the efficiency of iPSC generation26,27. Previously unpublished data revealed successful iPSC generation from M. musculus (C57BL/6N, C57BL/6 J, ICR) and Rattus norvegicus (Wistar-Imamichi) using the same vector. Although the gene sequences necessary for iPSC establishment in M. minutoides are currently unknown, we used vectors containing Homo sapiens sequences to facilitate successful conversion of M. minutoides fibroblasts into undifferentiated cells. Owing to the close evolutionary relationship between M. minutoides and M. musculus9, the successful establishment of M. minutoides iPSCs in this study is a significant achievement. The successful establishment of M. minutoides iPSCs was largely due to the use of highly efficient vectors that combine seven factors. The expression of EGFP was observed in colonies and persisted after passaging. PCR was performed to confirm that the iPSC colonies established in this study were derived from M. minutoides and not M. musculus. The difference in the length of the Ghr promoter region between M. minutoides and M. musculus, as observed in our previous study (unpublished data), was used to identify M. musculus and M. minutoides DNA via PCR, with the genome as a template. The genomic DNA extracted from M. minutoides iPSCs amplified not only M. minutoides DNA but also a small amount of M. musculus-derived products, which are speculated to be of feeder cell origin. Attempting a feeder-free culture of M. minutoides iPSCs was unsuccessful due to a decline in growth rate and a reduction in the number of colonies during cultivation.
We analyzed the gene expression profiles of M. minutoides iPSCs and observed the expression of various pluripotency marker genes. Immunostaining revealed the presence of NANOG and OCT3/4 in EGFP-positive M. minutoides iPSCs. Moreover, the colonies were observed to maintain their undifferentiated state. Although we did not determine whether the gene expression was endogenous or exogenous, interestingly, Eras and Fgf4, which are not expressed in the gene expression vectors commonly used for iPSC induction, were expressed. This finding suggests that the induction vectors employed in our study initiated reprogramming in M. minutoides fibroblasts. Based on Fgf4 sequence analysis, although the amino acid sequence remained identical, the cDNA sequence partially differed. This disparity in cDNA sequences indicated that fibroblast initialization triggered the reactivation of genes associated with the original pluripotent state specific to M. minutoides. Prior to induction, we observed the expression of the Klf4 gene in fibroblasts despite its inclusion in the gene expression vectors employed for iPSC induction. The Klf4 gene was originally isolated from NIH3T3, a cell line derived from mouse fetal skin28. Notably, Klf4 is expressed in the dermal fibroblasts of M. musculus and the fibroblasts of Tokudaia tokunoshimensis (Amami spiny rat) and Myotis lucifugus (little brown bat), which have been successfully used to generate iPSCs using similar induction methods23,29. These observations strongly suggest that Klf4 may also serve as an endogenous gene expressed in M. minutoides tail fibroblasts. Klf4 plays diverse roles, including the maintenance of skin barrier function. In stem cells, Klf4 actively regulates pluripotency gene expression and inhibits the induction of differentiation-related genes30. Therefore, Klf4 may exhibit different functions between M. minutoides iPSCs and fibroblasts.
Quantitative analysis revealed differences in the expression of pluripotency markers between the M. minutoides iPSC lines. In particular, the expression levels of the pluripotency marker genes, particularly Nanog, were nearly identical between the two iPSC lines (apm_iPSC_6 and iPSC_10) and M. musculus ES cells. In these two iPSC lines, the expression levels of Oct3/4, Sox2, Eras, and Fgf4 were significantly higher than those in M. musculus ESCs. Notably, the expression levels of Fgf4 were substantially high. Fgf4 is involved in the proliferation and differentiation of ESCs and tissue stem cells in mice31. Moreover, the absence of Fgf4 impairs differentiation into nerve and mesoderm lineages32, indicating its importance in specific differentiation. Based on our findings, the M. minutoides iPSCs established in this study readily differentiated into neurons, and a substantial number of cardiomyocyte colonies emerged from migrating cells within the EBs. This enhanced differentiation potential may be attributed to the elevated expression of Fgf4. RNA-seq analysis also revealed significant differences in gene expression between the established M. minutoides iPSCs and fibroblasts, highlighting the expression of numerous pluripotency-related genes that could not be elucidated by RT-PCR. Principal component analysis using gene expression data from M. musculus iPSCs (mouse_iPS_1, mouse_iPS_2) and fibroblasts (mouse_fib_1, mouse_fib_2) revealed that M. minutoides iPSCs and M. musculus iPSCs have different gene expression states. Moreover, distinct gene expression patterns were discovered between the fibroblasts of M. minutoides and M. musculus. Although we cannot definitively exclude the possibility that these differences are due to technical variations in the extraction processes, the availability of M. musculus iPSCs data from the GEO database implies that each cell population of M. minutoides and M. musculus may possess a unique gene expression profile that must be further investigated. In the enrichment analysis, the highest-ranked pathway matched pluripotency, and many other GO terms related to embryonic development and the maintenance of undifferentiated potential were observed. Based on the RT-qPCR results, a strong positive correlation was found among the expression levels of the pluripotency markers, suggesting that the regulatory mechanism governing pluripotency gene expression in M. minutoides was similar to that observed in M. musculus33,34; however, of the 292 genes included in the top hit WP1763: Mechanisms associated with pluripotency, only 102 genes (104 genes in iPSC_6 and 109 in iPSC_10) had significantly increased expression in M. minutoides iPSCs. As shown in Fig. 3c,d, 1661 upregulated and 1227 downregulated genes were unique to M. minutoides iPSCs. Thus, although M. minutoides belongs to the same genus (Mus) as mice, pathways involved in maintaining pluripotency in M. minutoides may differ from those in mice. GO:0006305 DNA alkylation is attributed to mitomycin C-treated feeder cells.
We proceeded to evaluate the pluripotency of M. minutoides iPSCs. Based on the in vitro results, these iPSCs successfully differentiated into neurons, exhibiting numerous elongated processes originating from dome-shaped colonies. This successful neural differentiation suggests that functional cells can be directly derived from the M. minutoides iPSCs established in this study. Subsequently, the floating culture method was employed to generate EBs from M. minutoides iPSCs. EBs are three-dimensional cellular aggregates formed from floating PSC cultures. EBs possess a two-layered structure comprising the EB ectoderm, which differentiates into neurons and other cell types, and the primitive endoderm, which differentiates into the mesoderm and endoderm. Using a floating-culture technique, we successfully formed EBs from M. minutoides iPSCs. These EBs exhibited a cell mass, with initiation of the cellular fate of the three germ layers (ectoderm, mesoderm, and endoderm). These results indicate that M. minutoides iPSCs are pluripotent and can differentiate into diverse cell lineages in vitro. Although EBs primarily differentiate into cell lineages comprising the EB, our gene expression analysis revealed slight expression of Cdx2, a marker associated with the trophic ectoderm. Although further detailed analysis of this finding is warranted, it highlights an intriguing outcome regarding EB differentiation in M. minutoides. The expression of the T (Brachyury) gene decreased significantly as the EB culture period progressed. T is a transcription factor expressed during the early stages of mesoderm formation35. According to previous reports, the expression of T is initially upregulated during the early culture of EBs derived from mouse ESCs, followed by a gradual decrease36. In EBs derived from M. minutoides iPSCs, an extended culture period resulted in the differentiation of EBs accompanied by a decrease in T expression, indicating differentiation into the mesoendodermal lineage, similar to M. musculus iPSCs.
In the present study, M. minutoides iPSCs differentiated into three germ layers in vivo, similar to M. musculus ESCs. Chimeric mice were successfully generated by injecting M. minutoides iPSCs into M. musculus blastocysts. The resulting chimeras exhibited a predominantly white coat color, characteristic of ICR (M. musculus), with additional patches displaying a wild coat color derived from M. minutoides. This observation indicates that M. minutoides iPSCs actively contribute to inner cell mass formation in conjunction with M. musculus-derived cells, demonstrating their functional and physiological capabilities in vivo.
PSCs are classified into two distinct states, naïve and primed, based on their morphological characteristics and capacity to form chimeric organisms37. For M. musculus, ESCs are recognized as naïve whereas epiblast stem cells (EpiSCs) are considered primed. These two states exhibit notable differences in their genetic expression profiles and their ability to contribute to chimerism38,39. RNA-seq analysis revealed that certain genes that were highly expressed in M. musculus ESCs, such as Zfp42, Dppa2, and Tbx3, were also significantly upregulated in M. minutoides iPSCs, whereas other genes were not significantly upregulated. Conversely, among the genes highly expressed in M. musculus EpiSCs, Fgf5 and Foxa2 exhibited significantly higher expression in M. minutoides iPSCs, whereas no significant differences were observed in M. musculus iPSCs. Considering the colony morphology and outcomes obtained in our study, including the ability to contribute to chimeric mice, the established M. minutoides iPSCs can be inferred to possess properties that are either comparable to or very closely aligned with those of naïve PSCs.
In summary, the findings of our in vitro and in vivo investigations confirmed that M. minutoides iPSCs generated by reprogramming M. minutoides fibroblasts possess the essential characteristics of PSCs and contribute to their ontogeny. The establishment of M. minutoides iPSCs has remarkable potential to advance the analysis of dwarfism, which is the most distinct M. minutoides phenotype. Thus, the biology of non-model animals that lack established developmental engineering techniques and experience challenges in terms of ESC establishment can be explored, which may also advance research on interspecies chimera creation.
Methods
Animal experiment
Mus minutoides (obtained from the Pet Shop YANOHASHI) was exsanguinated via cervical dislocation under anesthesia with medetomidine, midazolam, and butorphanol and used for each experimental procedure. The animal experiments complied with the regulations and guidelines of the Yamaguchi University Experimental Animal Committee (No. 291) and the Animal Care and Use Committee of the University of Tokyo (P22-092), and their protocols were duly approved by the respective committees. The results of the study were also reported in accordance with the ARRIVE guidelines.
Fibroblast establishment
Primary fibroblasts were isolated from the tail tips of dead M. minutoides and M. musculus. The tail tip was attached to gelatin-coated plates and cultured in Dulbecco’s modified Eagle’s medium (DMEM; FUJILIM Wako) supplemented with 10% fetal bovine serum (FBS), penicillin (100 U/mL), and streptomycin (100 µg/mL) at 37 °C with 5% CO2. The culture medium was replaced weekly, and the cells were passaged as necessary to monitor their migration.
Plasmid construction
PiggyBac inverted terminal repeat (ITR) sequences were synthesized by overlap extension PCR using oligonucleotides obtained from Eurofins (Japan). The amplicons were inserted into the AatII–AflIII sites of pUC19 to construct pUC19-PB. To construct the plasmid, PB-TRE-Oct3/4-2A-Klf4-2A-Sox2-2A-cMyc (PB-TRE-OKSM), we cloned the TRE-CMV minimal promoter and a polyA signal sequence fragment from a previously constructed plasmid DNA40. The OKSM fragment was obtained from pMaster3 (Addgene plasmid #58526)41, which was modified from a previously reported plasmid42. These fragments were inserted into the ITRs of pUC19-PB.
Similarly, to construct PB-TRE-Glis1-2A-Nanog-2A-lin28a (PB-TRE-GNL), the TRE-CMV minimal promoter and poly (A) signal sequence fragments were obtained as described above. The Glis1 open reading frame (ORF) was cloned using C57BL/6-derived testis cDNA, and the NL fragment was obtained from pMaster3 (Addgene plasmid #58526)41. These fragments were inserted into the ITRs of pUC19-PB.
To construct PB-CAG-rtTA-IRES-eGFP-pGK-NeoR, the CAG-rtTA and IRES-eGFP sequences were obtained from a previously constructed plasmid DNA40. These fragments, along with pGK-NeoR (Addgene plasmid #13442)43, were inserted into pUC19-PB.
The above plasmids will be deposited in Addgene.
For pCAG-T3-hyPBase-pA construction, the hyPBase ORF was cloned from pCMV-hyPBase (kindly provided by Dr. A Bradley, Wellcome Sanger Institute) using PCR and inserted into the NotI–ClaI site of pCAG-T3-hCas9-pA (Addgene plasmid #48625)44.
The constructed vectors were sequenced using a commercial sequencing kit (Applied Biosystems, Foster City, CA, USA) and a DNA sequencer (Applied Biosystems), according to the manufacturer’s instructions. All iPSC-inducing plasmid constructs are shown in Fig. 1a.
iPSC establishment
Fibroblasts from M. minutoides were dissociated, washed, and suspended in Opti-MEM (Gibco). The cell suspension was then transferred to a 4-mm cuvette electrode (SE-204; BEX Co., LTD), and 2.5 μg/100 μL of plasmid was added individually for iPSC induction (PB-TRE-OKSM, PB-TRE-GNL, PB2-CAG-rtTA-ires-eGFP pGK-Neo, and pCAG-T3-hyPBase-pA(95)). Electroporation was performed using a CUY21EDIT II electroporator (BEX) with a single 10 ms pulse at 350 V, followed by five 50 ms pulses at 40 V at 50 ms intervals. After electroporation, the cells were reseeded on feeder cells in ESGRO Complete Basal Medium (Merck) supplemented with penicillin (100 U/mL), streptomycin (100 μg/mL), 20% KnockOut Serum Replacement (KSR) (Gibco), and ESGRO-2i Supplement Kit (Merck), with 2 μg/mL (final concentration) doxycycline (DOX) (Takara). Emerging colonies with iPSC-like morphology were passaged on feeder cells and mitomycin C-treated mouse embryonic fibroblasts and cultured with DOX until the reappearance of the colonies. Subsequently, DOX supplementation was discontinued, and the cells were further cultured. Individual colonies that maintained domed-morphology were selected under a stereomicroscope and single cells from a colony were seeded on feeder cells in 96-well plates to establish multiple M. minutoides iPSC lines.
Genomic PCR
Primers were designed based on the promoter region sequence of the M. musculus growth hormone receptor (Ghr) gene, and genomic PCR was performed using BIOTAQ DNA polymerase (NIPPON Genetics). The primer sequences are listed in Table 2. The following PCR cycling conditions were employed: initial denaturation at 94 °C for 3 min, followed by 35 cycles of denaturation at 94 °C for 30 s, annealing at 64 °C for 45 s, extension at 72 °C for 45 s, and a final extension at 72 °C for 1 min.
Alkaline phosphatase staining
Alkaline phosphatase (ALP) staining was conducted using an Alkaline Phosphatase detection kit (Merck), according to the manufacturer’s protocol.
Genomic DNA and mRNA extraction and reverse transcription
Total RNA and genomic DNA (gDNA) were extracted using the NucleoSpin TriPrep kit (Takara) or ReliaPrep RNA Cell Miniprep System (Promega), respectively, according to the respective protocols. Total RNA was reverse-transcribed using the QuantiTect Reverse Transcription Kit (QIAGEN), according to the manufacturer’s protocol.
RT-PCR
The primer design for RT-PCR was based on highly conserved regions identified by comparing the DNA sequences of M. musculus, Mus pahari, and Mus spretus. Primers were designed for the pluripotency markers: Nanog, Oct3/4, Sox2, Klf4, Eras, and Fgf4; neural markers: Tuj1 and Nestin; markers for ectoderm (Tuj1, Nestin), mesoderm (T, Acta2), cardiomyocyte (Tnnt2), endoderm (Gata4, AFP), and trophectoderm (Cdx2) differentiation in embryoid bodies (EBs) and migrating cells; and primers for the Gapdh gene. The primer sequences are listed in Table 3. PCR amplification was performed using BIOTAQ DNA polymerase (Nippon Genetics) and the following cycling conditions: initial denaturation at 94 °C for 3 min, followed by 35 cycles of denaturation at 94 °C for 30 s, annealing at 64 °C for 45 s, extension at 72 °C for 45 s, and a final extension step at 72 °C for 1 min.
Sequencing
After PCR, bands corresponding to the desired sizes were excised from the electrophoresed gel, and the amplified DNA fragments were extracted using the FastGene Gel/PCR Extraction Kit (Nippon Genetics). The extracted DNA was sequenced at Yamaguchi University Center for Gene Research. The obtained sequence data were aligned at the nucleotide and amino acid levels using CLUSTALW 2.1 software (https://www.genome.jp/tools-bin/clustalw) to determine the corresponding amino acid sequences.
RT-qPCR
Real-time PCR was conducted using the KAPA SYBR Fast qPCR Kit (Nippon Genetics) with synthesized cDNA as the template. The reaction was performed on the CFX96 Touch Deep Well real-time PCR analysis system (Bio-Rad) using the following cycling conditions: initial denaturation at 95 °C for 3 min, followed by 40 cycles of denaturation at 95 °C for 3 s, and annealing/extension at 60 °C for 20 s. Each gene expression level was normalized to that of Gapdh and analyzed using relative cycle threshold (CT). The values are presented as mean ± standard deviation.
Cellular immunostaining
The cells were fixed with 4% paraformaldehyde for 30 min. After three washes with Tris-buffered saline (TBS), the cells were permeabilized with 0.2% Triton X-100 in TBS (TBST) for 20 min. After three additional washes with TBS, the cells were blocked with 5% goat serum in TBST for 30 min. Rabbit anti-mouse NANOG (diluted 200-fold, ab80892; Abcam) and rabbit anti-human mouse OCT4 (diluted 200-fold, C30A3C1; Cell Signaling Technology) were used as the primary antibodies for the pluripotency markers. In this study, a specific OCT4 antibody was used to recognize the pluripotency-associated OCT4A isoform rather than the more prevalent OCT4B isoform. These primary antibodies were diluted in 5% goat serum in TBST and incubated with the cells for 17 h at 4 °C. Thereafter, Alexa Fluor 594-conjugated goat anti-rabbit IgG (diluted 500-fold, ab150080; Abcam) was used as the secondary antibody. For neuronal differentiation, rat anti-mouse NESTIN antibody (diluted 200-fold, 012-26843, FUJIFILM Wako) was used as the primary antibody, and Alexa Fluor 568-conjugated goat anti-rat IgG (diluted 500-fold, A-11077; Thermo Fisher Scientific) was used as the secondary antibody. Following three additional washes with TBS, contrast staining was performed by incubating the cells with 1 g/mL DAPI in TBS for 15 min under shielded light.
RNA-seq analysis
Total RNA was extracted from M. minutoides iPSCs and fibroblasts using the Maxwell RSC RNA Cell Kit (Promega), and RNA-seq analysis was performed using the NovaSeq 6000 (Illumina) at the Yamaguchi University Center for Gene Research. The number of reads was 33.5 million for fib_1, 33.1 million for fib_2, 36.7 million for iPS-6, and 33.9 million for iPS-10. The expression levels were analyzed after TPM normalization. The RNA-seq data for mouse iPSCs and fibroblasts (GSE46104; GSM1123730, GSM1123731, GSM1123732, and GSM1123733) were obtained from NCBI Gene Expression Omnibus (GEO) database using GEO RNA-seq Experiments Interactive Navigator (GREIN) (http://www.ilincs.org/apps/grein/?gse =).
Neural differentiation
To induce the differentiation of M. minutoides iPSCs into neurons, the culture medium was replaced with the Ndiff227 medium (Takara).
Embryoid body formation
Mus minutoides iPSCs cultured on the feeder cells were detached and collected. The cells were then suspended in DMEM supplemented with 10% KSR, MEM non-essential amino acid solution (100×) (Fujifilm Wako), StemSureR 10 mmol/L 2-mercaptoethanol solution (100×) (Fujifilm Wako), penicillin (100 U/mL), and streptomycin (100 µg/mL). The supernatant containing floating iPSCs was collected and cultured in a suspension for EB formation. These EBs were subjected to adherent culture to obtain migrating cells. Total RNA was extracted from EBs, and migrating cells were derived from EBs and reverse-transcribed.
Teratoma formation
A total of 2 × 105 cells were collected, suspended in Matrigel (Corning), and subcutaneously injected into 8-week-old KSN male mice. After a 2-week incubation period, the teratomas were extracted, fixed with 3.7% paraformaldehyde, embedded in paraffin, sliced into 4 µm-thick sections, and stained with hematoxylin and eosin. Genomic DNA was extracted from teratomas using the NucleoSpin DNA FFPE XS kit (Takara), according to the manufacturer’s protocol.
Chimeric mice production
Interspecies chimeras between M. musculus and M. minutoides were generated using blastocyst complementation, according to the protocols described in Nagy et al.45, with some modifications. Briefly, ICR zygotes were obtained via in vitro fertilization46 and cultured in KSOMaa-BSA medium until the blastocyst stage. M. minutoides iPSCs were microinjected into the blastocoel of 4.0–4.5 dpc ICR blastocysts, which were then transferred into the uteri of pseudopregnant recipient ICR females on day 2.5 of the pseudopregnancy, the pups were obtained through natural birth.
Statistical analysis
Statistical analysis was performed using a t-test to determine significant differences between values. p < 0.05 was considered statistically significant.
References
Willan, K. & Meester, J. Breeding biology and postnatal development of the African dwarf mouse. Acta Theriol. 23, 55–74 (1978).
Bryja, J. et al. Pan-African phylogeny of Mus (subgenus Nannomys) reveals one of the most successful mammal radiations in Africa. BMC Evol. Biol. 14, 256 (2014).
Chevret, P., Robinson, T. J., Perez, J., Veyrunes, F. & Britton-Davidian, J. A phylogeographic survey of the pygmy mouse Mus minutoides in South Africa: Taxonomic and karyotypic inference from cytochrome b sequences of museum specimens. PLoS One 9, e98499 (2014).
Rahmoun, M. et al. Anatomical and molecular analyses of XY ovaries from the African pygmy mouse Mus minutoides. Sex Dev. 8, 356–363 (2014).
Zhao, L., Quinn, A., Ng, E. T., Veyrunes, F. & Koopman, P. Reduced activity of SRY and its target enhancer Sox9-TESCO in a mouse species with X*Y sex reversal. Sci. Rep. 7, 41378 (2017).
Vučak, M. et al. Genome sequences of five arenaviruses from pygmy mice (Mus minutoides) in Sierra Leone. Microbiol. Resour. Announc. 11, e0009522 (2022).
Zhou, Y. et al. A mammalian model for Laron syndrome produced by targeted disruption of the mouse growth hormone receptor/binding protein gene (the Laron mouse). Proc. Natl. Acad. Sci. USA 94, 13215–13220 (1997).
Lupu, F., Terwilliger, J. D., Lee, K., Segre, G. V. & Efstratiadis, A. Roles of growth hormone and insulin-like growth factor 1 in mouse postnatal growth. Dev. Biol. 229, 141–162 (2001).
Matsuya, S., Imai, H., Kiso, Y., Kusakabe, K. T. & Kano, K. Characterization and expression of DNA sequences encoding the growth hormone gene in African Pygmy Mouse (Mus minutoides). J. Vet. Med. Sci. 83, 1244–1247 (2021).
Takahashi, K. & Yamanaka, S. Induction of pluripotent stem cells from mouse embryonic and adult fibroblast cultures by defined factors. Cell 126, 663–676 (2006).
Thoma, E. C. et al. Establishment of a translational endothelial cell model using directed differentiation of induced pluripotent stem cells from Cynomolgus monkey. Sci. Rep. 6, 35830 (2016).
Dejosez, M. et al. Bat pluripotent stem cells reveal unusual entanglement between host and viruses. Cell 186, 957-974.e928 (2023).
Mandai, M. Pluripotent stem cell-derived retinal organoid/cells for retinal regeneration therapies: A review. Regen. Ther. 22, 59–67 (2023).
Hayashi, M. et al. Robust induction of primordial germ cells of white rhinoceros on the brink of extinction. Sci. Adv. 8, eabp683 (2022).
Katayama, M. et al. Induced pluripotent stem cells of endangered avian species. Commun. Biol. 5, 1049 (2022).
Liao, J. et al. Generation of induced pluripotent stem cell lines from adult rat cells. Cell Stem Cell 4, 11–15 (2009).
Takahashi, K. et al. Induction of pluripotent stem cells from adult human fibroblasts by defined factors. Cell 131, 861–872 (2007).
Han, X. et al. Generation of induced pluripotent stem cells from bovine embryonic fibroblast cells. Cell Res. 21, 1509–1512 (2011).
Mao, J. et al. Epigenetic modifiers facilitate induction and pluripotency of porcine iPSCs. Stem Cell Rep. 8, 11–20 (2017).
Chen, H. et al. Inducing goat pluripotent stem cells with four transcription factor mRNAs that activate endogenous promoters. BMC Biotechnol. 17, 11 (2017).
Liu, M. et al. Generation of sheep induced pluripotent stem cells with defined DOX-inducible transcription factors via piggyBac transposition. Front. Cell Dev. Biol. 9, 785055 (2021).
Li, Z. et al. Establishment of bactrian camel induced pluripotent stem cells and prediction of their unique pluripotency genes. Int. J. Mol. Sci. 24, 1917 (2023).
Honda, A. et al. Flexible adaptation of male germ cells from female iPSCs of endangered Tokudaia osimensis. Sci. Adv. 3, e1602179 (2017).
Katayama, M. et al. Induced pluripotent stem cells with six reprogramming factors from prairie vole, which is an animal model for social behaviors. Cell Transpl. 25, 783–796 (2016).
Miyawaki, S. et al. Tumour resistance in induced pluripotent stem cells derived from naked mole-rats. Nat. Commun. 7, 11471 (2016).
Yu, J. et al. Induced pluripotent stem cell lines derived from human somatic cells. Science 318, 1917–1920 (2007).
Maekawa, M. et al. Direct reprogramming of somatic cells is promoted by maternal transcription factor Glis1. Nature 474, 225–229 (2011).
Shields, J. M., Christy, R. J. & Yang, V. W. Identification and characterization of a gene encoding a gut-enriched Kruppel-like factor expressed during growth arrest. J. Biol. Chem. 271, 20009–20017 (1996).
Mo, X., Li, N. & Wu, S. Generation and characterization of bat-induced pluripotent stem cells. Theriogenology 82, 283–293 (2014).
Ghaleb, A. M. & Yang, V. W. Krüppel-like factor 4 (KLF4): What we currently know. Gene 611, 27–37 (2017).
Kosaka, N., Sakamoto, H., Terada, M. & Ochiya, T. Pleiotropic function of FGF-4: Its role in development and stem cells. Dev Dyn 238, 265–276 (2009).
Kunath, T. et al. FGF stimulation of the Erk1/2 signalling cascade triggers transition of pluripotent embryonic stem cells from self-renewal to lineage commitment. Development 134, 2895–2902 (2007).
Ambrosetti, D. C., Scholer, H. R., Dailey, L. & Basilico, C. Modulation of the activity of multiple transcriptional activation domains by the DNA binding domains mediates the synergistic action of Sox2 and Oct-3 on the fibroblast growth factor-4 enhancer. J. Biol. Chem. 275, 23387–23397 (2000).
Niwa, H. How is pluripotency determined and maintained?. Development 134, 635–646 (2007).
Showell, C., Binder, O. & Conlon, F. L. T-box genes in early embryogenesis. Dev. Dyn. 229, 201–218 (2004).
Evans, A. L. et al. Genomic targets of Brachyury (T) in differentiating mouse embryonic stem cells. PLoS One 7, e33346 (2012).
Nichols, J. & Smith, A. Naive and primed pluripotent states. Cell Stem Cell 4, 487–492 (2009).
Brons, I. G. et al. Derivation of pluripotent epiblast stem cells from mammalian embryos. Nature 448, 191–195 (2007).
Tesar, P. J. et al. New cell lines from mouse epiblast share defining features with human embryonic stem cells. Nature 448, 196–199 (2007).
Yoshioka, S., Fujii, W., Ogawa, T., Sugiura, K. & Naito, K. Development of a mono-promoter-driven CRISPR/Cas9 system in mammalian cells. Sci. Rep. 5, 18341 (2015).
Wu, S., Wu, Y., Zhang, X. & Capecchi, M. R. Efficient germ-line transmission obtained with transgene-free induced pluripotent stem cells. Proc. Natl. Acad. Sci. USA 111, 10678–10683 (2014).
Kim, S. I. et al. KLF4 N-terminal variance modulates induced reprogramming to pluripotency. Stem Cell Rep. 4, 727–743 (2015).
Soriano, P., Friedrich, G. & Lawinger, P. Promoter interactions in retrovirus vectors introduced into fibroblasts and embryonic stem cells. J. Virol. 65, 2314–2319 (1991).
Fujii, W., Kawasaki, K., Sugiura, K. & Naito, K. Efficient generation of large-scale genome-modified mice using gRNA and CAS9 endonuclease. Nucleic Acids Res. 41, e187 (2013).
Nagy, A., Gertsenstein, M., Vintersten, K. & Behringer, R. Manipulating the Mouse Embryo: A Laboratory Manual, 3rd edn, (Cold Spring Harbor Laboratory Press, 2003).
Holmlund, H., Yamauchi, Y., Durango, G., Fujii, W. & Ward, M. A. Two acquired mouse Y chromosome-linked genes, Prssly and Teyorf1, are dispensable for male fertilitydouble dagger. Biol. Reprod. 107, 752–764 (2022).
Funding
This study was funded by JSPS KAKENHI (20H03169, JP23H02365), Grants-in-aid and The Foundation for Growth Science.
Author information
Authors and Affiliations
Contributions
S.M. prepared all figures; and S.M. and K.F. were involved in conception and design, collection and assembly of data, data analysis and interpretation, manuscript writing, and final approval of the manuscript; H.I. and K.T.K. were involved in data analysis and interpretation, financial support, and final approval of the manuscript; W.F. and K.K. were involved in conception and design, financial support, administrative support, collection and assembly of data, data analysis, and interpretation, manuscript writing, and final approval of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Matsuya, S., Fujino, K., Imai, H. et al. Establishment of African pygmy mouse induced pluripotent stem cells using defined doxycycline inducible transcription factors. Sci Rep 14, 3204 (2024). https://doi.org/10.1038/s41598-024-53687-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-024-53687-9
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.