Abstract
Methane (CH4) emissions from ruminants are of a significant environmental concern, necessitating accurate prediction for emission inventories. Existing models rely solely on dietary and host animal-related data, ignoring the predicting power of rumen microbiota, the source of CH4. To address this limitation, we developed novel CH4 prediction models incorporating rumen microbes as predictors, alongside animal- and feed-related predictors using four statistical/machine learning (ML) methods. These include random forest combined with boosting (RF-B), least absolute shrinkage and selection operator (LASSO), generalized linear mixed model with LASSO (glmmLasso), and smoothly clipped absolute deviation (SCAD) implemented on linear mixed models. With a sheep dataset (218 observations) of both animal data and rumen microbiota data (relative sequence abundance of 330 genera of rumen bacteria, archaea, protozoa, and fungi), we developed linear mixed models to predict CH4 production (g CH4/animal·d, ANIM-B models) and CH4 yield (g CH4/kg of dry matter intake, DMI-B models). We also developed models solely based on animal-related data. Prediction performance was evaluated 200 times with random data splits, while fitting performance was assessed without data splitting. The inclusion of microbial predictors improved the models, as indicated by decreased root mean square prediction error (RMSPE) and mean absolute error (MAE), and increased Lin’s concordance correlation coefficient (CCC). Both glmmLasso and SCAD reduced the Akaike information criterion (AIC) and Bayesian information criterion (BIC) for both the ANIM-B and the DMI-B models, while the other two ML methods had mixed outcomes. By balancing prediction performance and fitting performance, we obtained one ANIM-B model (containing 10 genera of bacteria and 3 animal data) fitted using glmmLasso and one DMI-B model (5 genera of bacteria and 1 animal datum) fitted using SCAD. This study highlights the importance of incorporating rumen microbiota data in CH4 prediction models to enhance accuracy and robustness. Additionally, ML methods facilitate the selection of microbial predictors from high-dimensional metataxonomic data of the rumen microbiota without overfitting. Moreover, the identified microbial predictors can serve as biomarkers of CH4 emissions from sheep, providing valuable insights for future research and mitigation strategies.
Similar content being viewed by others
Introduction
As a potent greenhouse gas that contributes significantly to climate change, methane (CH4) emissions from ruminants pose a direct threat to the environment and sustainable agricultural production1. Methane emissions from ruminants also lead to a waste of part of the ingested feed energy2. In sheep, CH4 emissions result in approximately a 7% loss of gross energy intake3. In recent years, there has been growing interest in prediction models for CH4 emissions for several reasons. First, direct measurement of CH4 emissions from large herds or flocks of ruminants is technically difficult and time-consuming, and thus it can be done practically only with small numbers of animals in research settings4. Second, it is not practical or feasible to establish regional, national, or global inventories of CH4 emissions by direct measurement of CH4 emissions from individual animals. Third, statistical models can assist in describing and understanding the relationship among CH4 emissions, animal performance, and other factors (nutritional, microbial, and physiological).
A diverse set of prediction models has been developed to predict enteric CH4 emissions from sheep5 and cattle6,7. These prediction equations include predictor variables describing the dietary composition, feed intake, rumen fermentation profiles, and other animal traits such as body weight (BW), but not individual taxa (genera or species) of the rumen microbiota. These prediction models vary considerably in robustness and prediction accuracy8,9. The rumen microbiota is solely responsible for CH4 emissions from ruminants. Specifically, rumen bacteria, fungi, and protozoa digest and ferment the ingested feed and produce the substrates (methylamines, hydrogen, CO2, and formate) that are then used by the rumen methanogens to produce CH410. Thus, the rumen microbiota is directly responsible for the CH4 emitted by ruminants although it can be affected by several factors, including diet and host-related factors11. Indeed, several studies have reported strong associations or correlations of some bacteria, methanogens, or protozoa with or qualitatively indicative of CH4 production12,13. In this context, we hypothesized that the inclusion of individual taxa of the ruminal microbiota as predictor variables, in addition to animal and feed-related predictor variables, in CH4 prediction models would improve our ability to predict CH4 emissions. Methane prediction equations or models containing rumen microbial taxa as predictor variables may also help understand their roles and importance in enteric CH4 emissions. The objective of this study was to develop such prediction models using multiple statistical/machine learning (ML) methods and a dataset that contained both animal-related data and metataxonomic data of the rumen microbiota of a cohort of sheep that were not subjected to any dietary interventions.
Methods
Dataset
This study did not involve the use of any animals, and thus no approval of the Institutional Animal Care and Use Committee was required. The dataset (Supplementary Table 1) used in this study was generated in a previous study that compared the rumen microbiota and CH4 emissions among a group of New Zealand sheep that were not subjected to any dietary interventions13. The animal experiment and analyses were detailed in another report14. Briefly, 1225 sheep were fed a pelleted alfalfa diet (without any anti-methane inhibitors) twice daily at two times their maintenance requirement for energy (based on BW). CH4 emissions from 340 individual animals were measured with open-circuit respiration chambers twice in two independent measuring rounds (10–15 days apart) with each time lasting for two consecutive days14. CH4 production (g CH4/animal·d) and CH4 yield (g CH4/kg of dry matter intake, DMI) were calculated. Feed intake and BW of the 340 individual animals were also recorded. Rumen samples were collected from 118 of the 340 sheep, with 60 representing the highest and 58 representing the lowest CH4 yield sheep, immediately after each CH4 emission measurement (corresponding to 17–18 h after the last feeding) using the stomach tubing method. The two groups of sheep were balanced by BW (24–60 kg), sex, and breeds (indicated below). The rumen fermentation characteristics, including acetate:propionate (A:P) ratio, were analyzed using gas–liquid chromatography. We provided the density plots of the CH4 data (g/d and g/kg of DMI, Supplementary Fig. 1). Both density curves appeared to represent the mixture of the two groups, which is consistent with the data feature (high and low emission sheep) and with the patterns found in the previous study13.
Two rumen samples were collected (10 to 15 days apart) from each sheep and the composition and structure of the rumen microbiota were analyzed using metataxonomics sequencing the 16S rRNA genes of both bacteria and archaea, 18S rRNA gene of protozoa, and internal transcribed spacer 1 (ITS1) of fungi13. One sheep was excluded due to missing data, and 16 sheep were represented by only one sample that yielded complete metataxonomic data. This resulted in a cumulative 218 observations, with 101 sheep each contributing two observations and the remaining 16 sheep contributing one observation.
In this study, we retrieved the metataxonomic sequencing data of the sheep rumen samples from the EMBL database per the accession numbers (ERP003779 for Bacteria, ERP003773 for Archaea, ERP003772 for ciliate protozoa, and ERP003764 for anaerobic fungi). The sequences were then analyzed using DADA215. Briefly, the amplicon sequences were clustered into amplicon sequence variants (ASVs), which were then taxonomically assigned to individual taxa using specialty databases: the Silva database (version 138.1) for bacteria, archaea, and protozoa16; and the UNITE database for fungi17. The relative sequence abundance of each genus was calculated as the sum of the relative sequence abundance of all the ASVs assigned to the respective genus. The sequences of the ASVs that were classified to an unclassified genus (Uncl_genus) are listed in Supplementary Table 2. Only the genera that each had a relative sequence abundance reaching > 1% in all the samples were used as microbial variables. For an Uncl_genus, its higher and known taxon name was used to show its taxonomic lineage. In total, 330 microbial genera were obtained and used, including 308 genera of bacteria, 9 genera of archaea, 5 genera of protozoa, and 8 genera of fungi. The relative sequence abundance data of the microbial genera were log-transformed before the modeling process. In addition to the rumen microbiota data and CH4 emission data (CH4 production, g CH4/animal·d; CH4 yield, g CH4/kg of DMI), the dataset also contained the following five animal-related data of the individual sheep: DMI (kg/d), BW (kg), A:P ratio, sex, and breed (Coopworth, Corriedale, Finnish Landrace, Perendale, Romney, and Texel).
Linear mixed model
In our dataset, most animals (101 out of 118) had two records of CH4 emissions and other data (except for sex and breed). These repeated measurements from the same sheep were likely to be more akin to each other than to the measurements from a different sheep. Thus, the two records from the same sheep were expected to be correlated. Linear mixed-effects models were used to model the dependence of the data and account for correlations between the two repeated measurements. The general structure of the model was described as follows:
where yij is the observed CH4 emissions from the ith animal and jth observation (i = 1, …, 118 and j = 1, 2), set as either g CH4/animal·d (referred to as ANIM-B models) or g CH4/kg of DMI (referred to as DMI-B models); Xi = (1, DMI, BW, A:P ratio, sex, breed, microbial variables) is the design vector for the fixed effects containing the values of the predictor variables for the ith animal and the “1” entry for the intercept (DMI was not included as a variable in developing the DMI-B models because it is a dependent variable in those models); β is the coefficients vector of the fixed effects describing the relationships between CH4 emissions and the predictor variables; αi is the random animal effects to account for intra-animal correlation between the two measurements; and εij is the random error. The αi was assumed to be independent and identically distributed with mean zero and variance \({{\sigma }_{\alpha }}^{2}\), and the εij errors were assumed to be independent and identically distributed with mean zero and variance \({\sigma }^{2}\). Further, it was assumed that random effects and errors were mutually independent.
Machine learning methods
The rumen microbiota is diverse, containing many hundreds to thousands of distinct species of microbes at varying relative sequence abundance. The inclusion of such a large number of microbial variables in prediction models can lead to overfitting. Additionally, relative abundance data of microbiota are usually on different scales, zero-inflated, sparse, and over-dispersed. Thus, proper selection of microbial predictor variables to be included in prediction models is crucial in model development. In the present study, we used four ML methods to select microbial predictor variables: random forest combined with boosting (RF-B) using the randomForest and gbm packages18, least absolute shrinkage and selection operator (LASSO) using the glmnet package19, generalized linear mixed model with LASSO (glmmLasso) using the glmmLasso package20, and smoothly clipped absolute deviation (SCAD) using the splmm package21. We opted for both LASSO and glmmLasso because the former does not consider intra-animal correlation, while the latter takes these correlations into account by utilizing the linear mixed-effect model described in Eq. (1). Each of the ML methods was evaluated for both prediction performance and fitting performance. Specifically, to evaluate prediction performance (based on 200 evaluations with random data splitting into testing data and training data (see section ‘Evaluation of prediction performance’ below), each ML method was used to make variable selection (selection for both animal and microbial predictor variables). Using the lmer function of the lme4 package22, we also fitted conventional models utilizing solely the animal-related data (i.e., BW, A:P ratio, and DMI (DMI was used only in developing the ANIM-B models)) as predictors. The two categorical variables (sex and breed) were not included in the prediction models because their addition did not lead to improvement in prediction performance (data not shown). The sets of variables selected by glmmLasso, LASSO, SCAD, RF-B, and conventional modeling were then separately fitted with a linear mixed model, and the resultant models were used to predict CH4 emissions. Further, to evaluate fitting performances, linear mixed models were fitted with all available data by selecting the variables using the ML and the conventional method. The Akaike information criterion (AIC) and Bayesian information criterion (BIC) of those models were computed using the ANOVA function in R v4.0.223. The estimated standard deviation of the animal random effects (\({\widehat{\sigma }}_{\alpha }\)), coefficients, and associated standard errors (SE) of the predictor variables were computed using the lmer function22. The observations and residuals versus the model predictions were plotted to evaluate the linear mixed model assumptions for all the ML models.
Evaluation of prediction performance
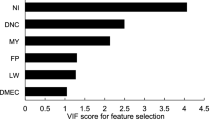
Prediction performance was validated in five steps. Step 1 involved screening the independent variables using the sure independence screening (SIS) test implemented in the SIS package in R24. This step was omitted for the RF-B method because this method can handle high-dimensional data without the need for data prescreening when building the decision trees. To check for potential multicollinearity, the variance inflation factor (VIF) was examined, and variables with a VIF < 5 were retained as recommended by Niu et al.7.
Step 2 began by randomly splitting the dataset into testing data (20% of the data) and training data (the remaining data). The training data were standardized by mean centering (subtracting each variable from the corresponding mean) and scaling (dividing each variable by its corresponding standard deviation). The testing data were standardized based on the centering and scaling from the training data. To prevent data leaking of the repeated records from the same animal, the two repeated records were always allocated to either the testing data or the training data.
In step 3, glmmLasso, LASSO, SCAD, and RF-B were individually used to select predictor variables from the training data. For glmmLasso and SCAD, the optimal tuning parameter λ was determined using BIC. Two increments of λ, 0.1 and 0.5, were also evaluated, but the results were virtually the same (data not shown) except for a substantial increase in computing time. For LASSO, tenfold cross-validation was used to determine the optimal tuning parameter λ. For RF-B, the top 10 important variables identified using the randomForest package were selected when the number of decision trees was set as 500, and the number of variables randomly sampled as candidates at each split (using mtry in the randomForest package) was set as 40. The top 10 important variables determined by boosting using the gbm package were also selected with the default settings applied. The union of the variables selected by random forest and boosting was kept.
Step 4 involved separately refitting a linear mixed model with the sets of selected variables using each of the ML methods and the training data. Predictions were then made using the values of the variables from the testing data. All the predictions and observations were recorded.
Step 5 repeated steps 2 to 4 for 200 times (sufficiently large and not too computationally intensive) by re-splitting the data into testing and training data in each iteration. Mean squared prediction error (MSPE), root MSPE (RMSPE), mean absolute errors (MAE), and Lin’s concordance correlation coefficients (CCC) were calculated using the equations specified in the section ‘Model evaluation equations’ below. We utilized RMSPE, MAE, and CCC to provide a comprehensive evaluation of the model performance. In particular, MAE is a robust assessment that is less sensitive to extreme values compared to RMSPE, whereas CCC can provide information on the precision and accuracy of the prediction.
Model evaluation equations
The values of MSPE, RMSPE, and MAE of all the models were calculated using the equations shown below25. The unit of MSPE was (g/d)2 for the ANIM-B models and (g/kg)2 for the DMI-B models. Lin’s CCC was calculated by the epi.ccc function in the epiR package in R26.
where \({O}_{i}\) is the CH4 emissions of ith observation; \({P}_{i}\) is CH4 prediction from the model; N is the total number of observations; \({S}_{o}\) is the standard deviation of the observations; \({S}_{p}\) is the standard deviation of the predictions; \(\overline{o }\) is the mean of the observations; \(\overline{p }\) is the mean of the predictions; \({S}_{op}\) is the covariance of the observations and the predictions.
The R codes we used in the present study were available at GitHub (https://github.com/yu2269/CH4_prediction_model).
Results
Model performance
Tables 1 and 2 summarize the prediction performances of the ANIM-B and the DMI-B models developed using both the ML and the conventional methods, based on random data splitting (200 times). The ML methods notably improved the performance of ANIM-B models, reducing RMSPE (by − 1.25% for SCAD and 3.8% for glmmLasso) and MAE (by − 0.39% for SCAD and 5.08% for LASSO), while increasing CCC (by 2.64% for SCAD and 9.8% for LASSO), as compared to the conventional method. Similarly, the ML methods enhanced the performance of DMI-B models, reducing RMSPE (by 6.22% for glmmLasso and 11.12% for LASSO) and MAE (by 7.36% for RF-B and 13.31% for LASSO) and increasing CCC (by 68.73% for glmmLasso and 106.18% for LASSO). Tables 3 and 4 present the fitting performance of the ANIM-B and the DMI-B models developed using both the ML methods and the conventional method, based on the entire dataset. Compared to the conventional method, the ML methods decreased BIC (by − 33 for LASSO and 19 for glmmLasso) and AIC (by − 152 for LASSO and 53 for glmmLasso) for the ANIM-B models. Similarly, for the DMI-B models, the ML methods decreased BIC (by − 201 for RF-B and 5 for glmmLasso and SCAD) and AIC (by − 157 for RF-B and 18 for glmmLasso), as compared to the conventional method.
The final models were selected by balancing the prediction performance and the fitting performance. Among the ANIM-B models, the glmmLasso model had the lowest BIC and AIC (Table 3). It also had the lowest RMSPE in prediction performance (Table 1). Thus, we selected the ANIM-B glmmLasso model as the final model. For the prediction performance of the DMI-B models, the LASSO model had a lower RMSPE than the SCAD model (1.89 vs. 1.92), which was lower than that of the glmmLasso model (Table 2), but the LASSO model had a higher BIC and AIC than the SCAD model and the glmmLasso model (Table 4). The DMI-B SCAD model also had the fewest variables (6 in total, Table 4) among the ML models. Thus, the DMI-B SCAD model was selected as the final DMI-B model.
We assessed the performance of the models by plotting the observed CH4 emissions against the CH4 emissions predicted using the final ANIM-B and the DMI-B models, along with the corresponding studentized residuals (Supplementary Figs. 2 and 3, respectively). Both the ANIM-B glmmLasso and the DMI-B SCAD models showed more tightly clustered and less scattered patterns in the observation vs. prediction plots, as compared to the ANIM-B and the DMI-B conventional models (Supplementary Figs. 2A,B and 3A,B, respectively). On the studentized residuals vs. predictions plots, the two ANIM-B and the DMI-B ML models also had tighter and more symmetric patterns around the regression lines that had smaller slopes compared to the ANIM-B and the DMI-B conventional models (Supplementary Figs. 2a,b and 3a,b, respectively).
Animal-based (g CH4/animal·d) prediction models
The final ANIM-B glmmLasso model contained three animal-related predictor variables (DMI, A:P ratio, and BW) and 10 bacterial genera (or equivalents) of rumen bacteria out of the 308 genera (Table 5). None of the protozoal, fungal, or archaeal genera was selected as predictor variables. The three animal-related variables all had a positive coefficient. Of the 10 microbial predictor variables, four had a positive coefficient: 0.59 for the Uncl_genus of Oscillospiraceae, 0.49 for the Uncl_genus of the order Clostridia, 0.46 for the Uncl_Family of the order Gastranaerophilales, and 0.24 for the genus Moryella. The remaining six genera had a negative coefficient: Uncl_Family of the order RF39 (‒0.27), the genus Prevotella_7 (‒0.30), the Uncl_Genus of Marinifilaceae (‒0.34), the genus Syntrophococcus (‒0.37), the genus Oribacterium (‒0.40), and the Uncl_Family of the order Oscillospirales (‒0.44). The ANIM-B conventional model had the same three of the five animal-related data as predictor variables (DMI, A:P ratio, and BW). Compared to the ANIM-B conventional model, the ANIM-B glmmLasso model had a reduced \({\widehat{\sigma }}_{\alpha }\) (1.59 vs. 2.04), RMSPE (by 3.8%), and MAE (by 4.6%), and increased CCC (by 9.49%). The ANIM-B glmmLasso model also had a lower AIC and BIC than the ANIM-B conventional model (by 53 and 19, respectively).
Dry matter intake-based (g CH 4 /kg of DMI) prediction model
The final DMI-B SCAD model had six predictor variables: one animal-related predictor variable (A:P ratio) and five bacterial genera as microbial predictor variables (Table 6). No protozoal, fungal, or archaeal genera were selected as predictor variables. The A:P ratio predictor variable had a positive coefficient. Except for Uncl_Family of the order Gastranaerophilales (with a coefficient of 0.50), the other four microbial predictor variables were all known genera, all of which had a negative coefficient: − 0.24 for Pseudoramibacter, − 0.33 for Megasphaera, − 0.39 for Selenomonas, and − 0.40 for Oribacterium. The DMI-B conventional model had two animal-related predictor variables (A:P ratio and BW), and both had a positive coefficient. Compared to the DMI-B conventional model, the DMI-B SCAD model had a reduced \({\widehat{\sigma }}_{\alpha }\) (0.98 vs. 1.47), RMSPE (by 9.66%), and MAE (by 9.97%), and increased CCC (by 84.36%). Compared to the DMI-B conventional model, the DMI-B SCAD model reduced both AIC and BIC (by 15 and 5, respectively).
Discussion
The rumen microbiota is responsible for about 90% of the CH4 emitted from sheep27. With respect to CH4 production, rumen microbes can be categorized as producers of methanogenesis substrates (primarily bacteria and protozoa), CH4 producers (i.e., methanogens), and those that influence CH4 production by interacting with the above two categories either positively (e.g., through mutualism and commensalism) or negatively (i.e., through amensalism, competition, and predation) interactions. Thus, some of the rumen microbes could have a quantitative relationship with the overall CH4 emissions. Using four statistical/ML methods and one dataset (218 observations) that contains both animal-related data and metataxonomic data of the rumen microbiota (330 genera in total), we developed ML models for improved prediction of CH4 production (g CH4/animal·d) and CH4 yield (g CH4/kg of DMI) from sheep, which include rumen microbes as predictor variables.
New ML models including microbial predictor variables
As evaluated for prediction performance, it is evident that all the ML models, except for the ANIM-B SCAD model, improved prediction of CH4 emissions in terms of error, bias, and accuracy when compared with the conventional models. Additionally, the improvement in prediction performance demonstrated in the evaluation with testing data randomly selected from the entire dataset suggests the potential usefulness of these models over a broad range of CH4 emissions. The improvement in CH4 prediction by the final ANIM-B glmmLasso model and the final DMI-B SCAD model is also evident in the plots of the predictions vs. the observations along with the studentized residuals. In addition to the same animal-related predictor variables included in the conventional models (except for the exclusion of BW in the DMI-B SCAD model), the final ML included genera of bacteria as predictor variables. The improvement in prediction performance strongly suggests that the inclusion of these microbial predictor variables might have fine-tuned the models. Moreover, the improved predictions of the final ML models align well with the assumptions of linear mixed models. These findings suggest that microbial predictor variables can differentiate sheep based on CH4 production levels, offering promising prospects for capturing variations in CH4 emissions and helping achieve accurate CH4 prediction.
The bacterial genera included in the ML models
Despite the dataset containing 330 genera of microbes, only a small number of genera were selected as microbial predictor variables, and their inclusion did not lead to model overfitting. Notably, all the microbial predictor variables in both the ANIM-B and the DMI-B prediction models are genera of bacteria, with the majority belonging to Firmicutes, an abundant phylum of rumen microbiota. Interestingly, no archaea, protozoa, or fungi were selected as predictor variables in any of the ML models. The absence of methanogen genera in the models might seem counterintuitive, given their role as CH4 producers. However, previous studies have shown that the abundance of rumen methanogens, even that of the most abundant genus, Methanobrevibacter28, has very a weak to no correlation with CH4 production12,29. Furthermore, changes in diets that significantly affected CH4 production were not found to lead to substantial alterations in the relative sequence abundance of Methanobrevibacter30. Thus, the lack of methanogens in the final prediction models is not surprising. At low abundance, methanogens, fungi, and protozoa might have little predicting power for CH4 emissions.
Both the ANIM-B and DMI-B ML models included the genus Oribacterium and one unclassified genus of the order Gastranaerophilales (the Uncl_Family of the order Gastranaerophilales) with a negative and a positive coefficient, respectively. Although the function of this unclassified genus is not known, its large coefficients in both prediction models suggest a high predicting power. Interestingly, Gastranaerophilales has been shown to be negatively associated with milk yield in dairy cattle31 and negatively associated with growth in lambs32. Regarding the genus Oribacterium, it had a negative coefficient in both the final ANIM-B and the DMI-B models. This aligns with Oribacterium being a potential H2 sink in the rumen of sheep33 and its negative association with CH4 emissions documented in dairy cows34. Future research is warranted to explore and verify if Oribacterium has a quantitative relationship with rumen CH4 production and can be used as a predictor variable or biomarker of CH4 emissions.
In addition to the two genera shared with the DMI-B SCAD model, the ANIM-B glmmLasso model included another eight genera, five of which were unclassified, and their functions remain unknown. Among these genera, Moryella, Prevotella_7, and Syntrophococcus are known genera of bacteria, but their association or correlation with CH4 emissions in ruminants has not been reported in the literature. Prevotella_7 is a newly established genus35, consisting of P. multiformis, P. multisaccharivorax, and uncultured bacteria from the oral cavities and the rumen (https://www.arb-silva.de/). Studies have shown that both P. multiformis and P. multisaccharivorax were enriched by concentrate-based diets correlated with decreased CH4 emissions36,37. In buffalos, a higher relative sequence abundance of Prevotella (closely related to Prevotella_7) was associated with low CH4 emissions38. Prevotella is considered to be hydrogen-consuming39, making the inclusion of Prevotella_7 as a predictor variable with a negative coefficient not surprising. As for Moryella and Syntrophococcus, the literature has no information about their relationship with CH4 emissions, except for one in vitro study that showed a positive correlation between Syntrophococcus and maximum CH4 production, but a negative correlation with the appearance of peak CH4 production40. Future quantitative studies are needed to verify the qualitative relationship between these two genera and CH4 emissions.
The DMI-B SCAD model did not include BW as a predictor variable, but the inclusion of bacterial predictor variables improved the prediction accuracy. This suggests that these bacterial predictor variables might have reduced the variation in CH4 prediction introduced by BW. In addition to the two unknown genera discussed earlier, the final DMI-B SCAD model included three known genera (Pseudoramibacter, Megasphaera, and Selenomonas), which were not included in the final ANIM-B glmmLasso model. While the literature has no information about any potential association between Pseudoramibacter and CH4 emissions, Pseudoramibacter was reported to be positively correlated with feed efficiency in sheep41. Similarly, it remains unknown if Megasphaera or Selenomonas has any association or correlation with CH4 emissions in the absence of anti-methane inhibitors. However, one in vitro study has shown that calcium salts of long-chain fatty acids42, tucuma oil43, and ginkgo extract44 decreased CH4 emissions while increasing propionate and the abundance of Megasphaera and Selenomonas. Encapsulated nitrate has also been found to reduce CH4 emissions while increasing Selenomonas in steers45. Additionally, Megasphaera was more abundant in low than in high CH4-yield sheep46. Thus, the negative coefficients of both Megasphaera and Selenomonas in the DMI-B SCAD model align with their negative association with CH4 emissions and their ability to consume hydrogen and produce propionate33,39,47,48. It should be noted that many of these correlations were observed in ruminants that were subjected to dietary interventions using anti-methane inhibitors, which could directly or indirectly affect these genera, skewing their relationships with CH4 production. Future research is warranted to determine their relationship with CH4 emissions in the absence of anti-methane inhibitors and their predicting power of CH4 emissions.
Potential limitations of the sheep dataset and the selected models
The dataset used in developing the prediction models was from one study using a relatively small number of sheep that were fed the same diet13. A larger database from multiple studies using different diets would improve prediction models and enhance their applicability across a broad range of feeding systems. However, currently, no such a database exists. A few studies in the literature have reported rumen microbiota data together with CH4 emission data, and we attempted to combine these data into a larger dataset. However, most of the studies used anti-methane inhibitors, likely skewing the microbiota data. Additionally, different studies sequenced different regions of the 16S rRNA or 18S rRNA gene, making it impossible to combine the sequence data for microbiota analysis. To facilitate the combination of sequence data from multiple studies, a standardized metataxonomic protocol should be adopted.
Metataxonomic analysis of the rumen microbiota incurs additional cost, which may hinder the adoption and application in the livestock industry, but it is relatively small and justifiable for several reasons. First, amplicon sequences are routinely generated and analyzed in numerous studies to understand the microbiological underpinning of treatment effects or group differences, making sequencing cost a part of the study cost. Second, the cost of amplicon sequencing has been steadily decreasing, and with the advent of new sequencing technologies (e.g., the Illumina NovaSeq 6000), it is expected to further decrease. Third, accurate and precise CH4 emission prediction is becoming increasingly important for precision livestock farming, which is an essential approach to increase feed efficiency while reducing CH4 emissions for a sustainable industry of ruminants. Improved prediction of CH4 emissions can assist researchers and farmers in improving the overall feed efficiency and sustainability of livestock production.
Conclusion
Using machine learning methods, we successfully developed animal-based and dry matter intake-based CH4 prediction models that include both animal data and microbial genera as predictor variables. The inclusion of microbial predictor variables improved CH4 emission prediction in terms of error, bias, and accuracy compared to the conventional models that solely contain animal-related data, particularly for predicting CH4 yield (g CH4/kg of DMI). Notably, the coefficients (positive or negative) of some of the predictor variables from known bacterial genera (e.g., Oribacterium, Prevotella_7, Megasphaera, and Selenomonas) align well with their known associations or correlations (positive or negative) with CH4 emissions and/or with their known functions in the rumen ecosystem, such as hydrogen consumption. However, the role and association of other predictor variables of known genera (e.g., Moryella, Pseudoramibacter, and Syntrophococcus), as well as the unclassified genera, with CH4 emissions remain unknown.
While further improvements in CH4 prediction models would benefit from extensive databases encompassing rumen bacteria, archaea, fungi, and protozoa, along with animal-related data from multiple studies, this study demonstrated the potential and feasibility to include rumen microbes as predictor variables to improve CH4 predictions from sheep, likely from cattle also. A standardized approach to generate and analyze rumen microbial data is imperative to facilitate the compilation of multiple datasets from numerous studies. Such efforts will contribute to a better understanding of the rumen microbiota's impact on CH4 emissions and improved prediction models for sustainable agricultural practices.
Data availability
The sequence and animal data used in this study were published by Kittelmann, et al.13 and can be downloaded from the EMBL database (accession numbers: ERP003779 for Bacteria, ERP003773 for Archaea, ERP003772 for ciliate protozoa, and ERP003764 for anaerobic fungi). The R codes used in this study are available at https://github.com/yu2269/CH4_prediction_model.
Abbreviations
- CH4 :
-
Methane
- ANIM-B:
-
Animal-based
- A:P ratio:
-
Acetate:propionate ratio
- ITS1:
-
Internal transcribed spacer 1
- ASV:
-
Amplicon sequence variant
- VIF:
-
Variance inflation factor
- Uncl:
-
Uncultured
- BW:
-
Body weight
- DMI:
-
Dry matter intake
- DMI-B:
-
Dry matter-based
- ML:
-
Machine learning
- RF:
-
Random forest
- RF-B:
-
Random forest combined with boosting
- LASSO:
-
Least absolute shrinkage and selection operator
- glmmLasso:
-
Generalized linear mixed with LASSO
- SCAD:
-
Smoothly clipped absolute deviation
- AIC:
-
Akaike information criterion
- BIC:
-
Bayesian information criterion
- SE:
-
Standard errors
- SIS:
-
Sure independence screening
- MSPE:
-
Mean square prediction error
- RMSPE:
-
Root mean square prediction error
- MAE:
-
Mean absolute errors
- CCC:
-
Concordance correlation coefficient
References
Masson-Delmotte, V. et al. Climate Change 2021: The Physical Science Basis. Contribution of Working Group I to the Sixth Assessment Report of the Intergovernmental Panel on Climate Change (2021).
Johnson, K. A. & Johnson, D. E. Methane emissions from cattle. J. Anim. Sci. 73, 2483–2492. https://doi.org/10.2527/1995.7382483x (1995).
Pelchen, A. & Peters, K. J. Methane emissions from sheep. Small Rumin. Res. 27, 137–150 (1998).
Hammond, K. J. et al. Review of current in vivo measurement techniques for quantifying enteric methane emission from ruminants. Anim. Feed Sci. Technol. 219, 13–30. https://doi.org/10.1016/j.anifeedsci.2016.05.018 (2016).
Zhao, Y. G., O’Connell, N. E. & Yan, T. Prediction of enteric methane emissions from sheep offered fresh perennial ryegrass (Lolium perenne) using data measured in indirect open-circuit respiration chambers. J. Anim. Sci. 94, 2425–2435. https://doi.org/10.2527/jas.2016-0334 (2016).
van Lingen, H. J. et al. Prediction of enteric methane production, yield and intensity of beef cattle using an intercontinental database. Agric. Ecosyst. Environ. 283, 106575. https://doi.org/10.1016/j.agee.2019.106575 (2019).
Niu, M. et al. Prediction of enteric methane production, yield, and intensity in dairy cattle using an intercontinental database. Glob. Chang. Biol. 24, 3368–3389. https://doi.org/10.1111/gcb.14094 (2018).
Appuhamy, J. A., France, J. & Kebreab, E. Models for predicting enteric methane emissions from dairy cows in North America, Europe, and Australia and New Zealand. Glob. Chang. Biol. 22, 3039–3056. https://doi.org/10.1111/gcb.13339 (2016).
Hristov, A. N. et al. Symposium review: Uncertainties in enteric methane inventories, measurement techniques, and prediction models. J. Dairy Sci. 101, 6655–6674. https://doi.org/10.3168/jds.2017-13536 (2018).
Patra, A. K. & Yu, Z. Combinations of nitrate, saponin, and sulfate additively reduce methane production by rumen cultures in vitro while not adversely affecting feed digestion, fermentation or microbial communities. Bioresour. Technol. 155, 129–135. https://doi.org/10.1016/j.biortech.2013.12.099 (2014).
Newbold, C. J. & Ramos-Morales, E. Review: Ruminal microbiome and microbial metabolome: Effects of diet and ruminant host. Animal 14, s78–s86. https://doi.org/10.1017/S1751731119003252 (2020).
Tapio, I., Snelling, T. J., Strozzi, F. & Wallace, R. J. The ruminal microbiome associated with methane emissions from ruminant livestock. J. Anim. Sci. Biotechnol. 8, 7. https://doi.org/10.1186/s40104-017-0141-0 (2017).
Kittelmann, S. et al. Two different bacterial community types are linked with the low-methane emission trait in sheep. PLoS ONE 9, e103171. https://doi.org/10.1371/journal.pone.0103171 (2014).
Pinares-Patino, C. S. et al. Heritability estimates of methane emissions from sheep. Animal 7(Suppl 2), 316–321. https://doi.org/10.1017/S1751731113000864 (2013).
Callahan, B. J. et al. DADA2: High-resolution sample inference from Illumina amplicon data. Nat. Methods 13, 581–583. https://doi.org/10.1038/nmeth.3869 (2016).
Quast, C. et al. The SILVA ribosomal RNA gene database project: Improved data processing and web-based tools. Nucleic Acids Res. 41, D590-596. https://doi.org/10.1093/nar/gks1219 (2013).
Nilsson, R. H. et al. The UNITE database for molecular identification of fungi: Handling dark taxa and parallel taxonomic classifications. Nucleic Acids Res. 47, D259–D264. https://doi.org/10.1093/nar/gky1022 (2019).
Greg, R. gbm: Generalized Boosted Regression Models v. 2.1.8.1 (2010).
Friedman, J., Hastie, T. & Tibshirani, R. Regularization paths for generalized linear models via coordinate descent. J. Stat. Softw. 33, 1–22. https://doi.org/10.18637/jss.v033.i01 (2010).
Groll, A. & Tutz, G. Variable selection for generalized linear mixed models by L 1-penalized estimation. Stat. Comput. 24, 137–154. https://doi.org/10.1007/s11222-012-9359-z (2012).
Yang, L. Model-Based Clustering of Longitudinal Data in High Dimensions Thesis (Ph.D.) thesis (University of Rochester, 2021).
Bates, D., Mächler, M., Bolker, B. & Walker, S. Fitting linear mixed-effects models using lme4. J. Stat. Softw. 67, 1–48. https://doi.org/10.18637/jss.v067.i01 (2015).
Team, R. C. R: A Language and Environment for Statistical Computing (v. 4.0. 2) [Computer Software] (R Foundation for Statistical Computing, 2020).
Saldana, D. F. & Feng, Y. SIS: An R package for sure independence screening in ultrahigh-dimensional statistical models. J. Stat. Softw. https://doi.org/10.18637/jss.v083.i02 (2018).
James, G., Witten, D., Hastie, T., Tibshirani, R. & Taylor, J. An Introduction to Statistical Learning with Applications in R 2nd edn, 445 (Springer, 2021).
Stevenson, M. et al. EpiR: An R Package for the Analysis of Epidemiological Data v. 2.0.62 (2013).
Murray, R. M., Bryant, A. M. & Leng, R. A. Rates of production of methane in the rumen and large intestine of sheep. Br. J. Nutr. 36, 1–14. https://doi.org/10.1079/bjn19760053 (1976).
Zhou, M., Hernandez-Sanabria, E. & Guan, L. L. Characterization of variation in rumen methanogenic communities under different dietary and host feed efficiency conditions, as determined by PCR-denaturing gradient gel electrophoresis analysis. Appl. Environ. Microbiol. 76, 3776–3786. https://doi.org/10.1128/AEM.00010-10 (2010).
Cunha, C. S. et al. Compositional and structural dynamics of the ruminal microbiota in dairy heifers and its relationship to methane production. J. Sci. Food Agric. 99, 210–218. https://doi.org/10.1002/jsfa.9162 (2019).
van Gastelen, S. et al. Linseed oil and DGAT1 K232A polymorphism: Effects on methane emission, energy and nitrogen metabolism, lactation performance, ruminal fermentation, and rumen microbial composition of Holstein-Friesian cows. J. Dairy Sci. 100, 8939–8957. https://doi.org/10.3168/jds.2016-12367 (2017).
Bach, A. et al. Changes in the rumen and colon microbiota and effects of live yeast dietary supplementation during the transition from the dry period to lactation of dairy cows. J. Dairy Sci. 102, 6180–6198. https://doi.org/10.3168/jds.2018-16105 (2019).
Perea, K. et al. Feed efficiency phenotypes in lambs involve changes in ruminal, colonic, and small-intestine-located microbiota. J. Anim. Sci. 95, 2585–2592. https://doi.org/10.2527/jas.2016.1222 (2017).
Greening, C. et al. Diverse hydrogen production and consumption pathways influence methane production in ruminants. ISME J. 13, 2617–2632. https://doi.org/10.1038/s41396-019-0464-2 (2019).
Lopez-Garcia, A. et al. Fungal and ciliate protozoa are the main rumen microbes associated with methane emissions in dairy cattle. Gigascience 11, 88. https://doi.org/10.1093/gigascience/giab088 (2022).
Henderson, G. et al. Improved taxonomic assignment of rumen bacterial 16S rRNA sequences using a revised SILVA taxonomic framework. PeerJ 7, e6496. https://doi.org/10.7717/peerj.6496 (2019).
Ellison, M. J. et al. Diet and feed efficiency status affect rumen microbial profiles of sheep. Small Rumin. Res. 156, 12–19. https://doi.org/10.1016/j.smallrumres.2017.08.009 (2017).
Hristov, A. N. et al. Special topics–mitigation of methane and nitrous oxide emissions from animal operations: I. A review of enteric methane mitigation options. J. Anim. Sci. 91, 5045–5069. https://doi.org/10.2527/jas.2013-6583 (2013).
Aguilar-Marin, S. B., Betancur-Murillo, C. L., Isaza, G. A., Mesa, H. & Jovel, J. Lower methane emissions were associated with higher abundance of ruminal Prevotella in a cohort of Colombian buffalos. BMC Microbiol. 20, 364. https://doi.org/10.1186/s12866-020-02037-6 (2020).
Denman, S. E., Martinez Fernandez, G., Shinkai, T., Mitsumori, M. & McSweeney, C. S. Metagenomic analysis of the rumen microbial community following inhibition of methane formation by a halogenated methane analog. Front. Microbiol. 6, 1087. https://doi.org/10.3389/fmicb.2015.01087 (2015).
Li, R. et al. Effect of different forage-to-concentrate ratios on ruminal bacterial structure and real-time methane production in sheep. PLoS ONE 14, e0214777. https://doi.org/10.1371/journal.pone.0214777 (2019).
McLoughlin, S. et al. Rumen microbiome composition is altered in sheep divergent in feed efficiency. Front. Microbiol. 11, 1981. https://doi.org/10.3389/fmicb.2020.01981 (2020).
Sato, Y. et al. Calcium salts of long-chain fatty acids from linseed oil decrease methane production by altering the rumen microbiome in vitro. PLoS ONE 15, e0242158. https://doi.org/10.1371/journal.pone.0242158 (2020).
Ramos, A. F. O. et al. Tucuma oil shifted ruminal fermentation, reducing methane production and altering the microbiome but decreased substrate digestibility within a RUSITEC fed a mixed hay–concentrate diet. Front. Microbiol. 9, 1647. https://doi.org/10.3389/fmicb.2018.01647 (2018).
Oh, S., Koike, S. & Kobayashi, Y. Effect of ginkgo extract supplementation on in vitro rumen fermentation and bacterial profiles under different dietary conditions. Anim. Sci. J. 88, 1737–1743. https://doi.org/10.1111/asj.12877 (2017).
Granja-Salcedo, Y. T. et al. Long-term encapsulated nitrate supplementation modulates rumen microbial diversity and rumen fermentation to reduce methane emission in grazing steers. Front. Microbiol. 10, 614. https://doi.org/10.3389/fmicb.2019.00614 (2019).
Kamke, J. et al. Rumen metagenome and metatranscriptome analyses of low methane yield sheep reveals a Sharpea-enriched microbiome characterised by lactic acid formation and utilisation. Microbiome 4, 56. https://doi.org/10.1186/s40168-016-0201-2 (2016).
Chen, L. et al. Megasphaera elsdenii lactate degradation pattern shifts in rumen acidosis models. Front. Microbiol. 10, 162. https://doi.org/10.3389/fmicb.2019.00162 (2019).
Marchandin, H., Juvonen, R. & Haikara, A. Megasphaera. In Bergey’s Manual of Systematics of Archaea and Bacteria (eds Trujillo, M. E. et al.) 1–16 (Wiley, 2015).
Acknowledgements
This study is part of the Joint Programming Initiative on Agriculture, Food Security and Climate Change (FACCE-JPI)’s “GLOBAL NETWORK” project and the “Feeding and Nutrition Network” (http://animalscience.psu.edu/fnn) of the Livestock Research Group within the Global Research Alliance for Agricultural Greenhouse Gases (www.globalresearchalliance.org).
Funding
The authors gratefully acknowledge funding for this project from the USDA National Institute of Food and Agriculture (Award number: 2014-67003-21979). The animal and microbial data originated from a study funded by the Pastoral Greenhouse Gas Research Consortium (www.pggrc.co.nz).
Author information
Authors and Affiliations
Contributions
Z.Y. conceived and designed the study. P.J. provided the data. Z.Y., B.Z., and L.M. set up the database. B.Z. did the modeling. L.M. and Z.Y. supervise the study. B.Z. wrote the manuscript with input from all co-authors. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Zhang, B., Lin, S., Moraes, L. et al. Methane prediction equations including genera of rumen bacteria as predictor variables improve prediction accuracy. Sci Rep 13, 21305 (2023). https://doi.org/10.1038/s41598-023-48449-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-023-48449-y
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.