Abstract
Burkholderia pseudomallei, an etiological agent of melioidosis is an environmental bacterium that can survive as an intracellular pathogen. The biofilm produced by B. pseudomallei is crucial for cellular pathogenesis of melioidosis. The purpose of this investigation is to explore the role of biofilm in survival of B. pseudomallei during encounters with Acanthamoeba sp. using B. pseudomallei H777 (a biofilm wild type), M10 (a biofilm defect mutant) and C17 (a biofilm-complemented strain). The results demonstrated similar adhesion to amoebae by both the biofilm wild type and biofilm mutant strains. There was higher initial internalisation, but the difference diminished after longer encounter with the amoeba. Interestingly, confocal laser scanning microscopy demonstrated that pre-formed biofilm of B. pseudomallei H777 and C17 were markedly more persistent in the face of Acanthamoeba sp. grazing than that of M10. Metabolomic analysis revealed a significant increased level of 8-O-4′-diferulic acid, a superoxide scavenger metabolite, in B. pseudomallei H777 serially passaged in Acanthamoeba sp. The interaction between B. pseudomallei with a free-living amoeba may indicate the evolutionary pathway that enables the bacterium to withstand superoxide radicals in intracellular environments. This study supports the hypothesis that B. pseudomallei biofilm persists under grazing by amoebae and suggests a strategy of metabolite production that turns this bacterium from saprophyte to intracellular pathogen.
Similar content being viewed by others
Introduction
Burkholderia pseudomallei is an etiological agent of melioidosis. This bacterium is generally an environmental saprophyte dwelling in soil and water1,2,3,4,5. This pathogen can be transmitted to susceptible human hosts via ingestion, inhalation, or skin inoculation. It can become an intracellular pathogen, evading host immune surveillance using numerous virulence strategies and contribute to its pathogenicity and disease severity, resulting in mortality rates that range from 40 to 70%6,7,8,9. Melioidosis is of growing public health concern, causing an estimated 165,000 cases and 89,000 deaths per year10. Recently, there has been a call for WHO to officially recognise melioidosis as a neglected tropical disease11. There is clearly a need to understand how this saprophytic bacterium evolved to become a life-threatening pathogen.
Burkholderia pseudomallei can persist in non-living reservoirs, including distilled water, for 16 years12 and remains viable in a soil microcosm for at least 120 days2. The bacterium can also live in other living organisms including grasses13 and free-living amoebae from the genus Acanthamoeba14. Biofilm formation is a key factor for bacterial survival in diverse natural environments and in interactions with the host15. Biofilm formation by Pseudomonas aeruginosa and Vibrio cholerae facilitates their survival and persistence in the environment despite grazing by protozoans16,17. Previous research has established that B. pseudomallei biofilm promotes bacterial adhesion and internalisation in human epithelial A549 cells18. It has not yet been determined whether B. pseudomallei biofilm plays a role in interactions with living organisms other than human hosts.
Acanthamoeba offers a model for the development of intracellular pathogenicity in humans as they facilitate the intracellular survival of pathogens within themselves19,20. Both human professional phagocytes and amoebae produce reactive oxygen and nitrogen species such as nitric oxide (NO), superoxide (O2–) and hydrogen peroxide (H2O2) as antimicrobial molecules21,22. It has previously been observed that hypervirulent V. cholerae evolution after passing through protozoan predation acquired strategies to reduce intracellular stress responses including superoxide (O2−) and H2O223,24. Moreover, previous research has established that superoxide dismutase C production is essential to provide resistance against killing by reactive oxygen intermediates, leading to intracellular survival in host phagocytes and hence virulence of B. pseudomallei12. Much uncertainty remains about the intracellular adaptations that allow environmental B. pseudomallei to become an intracellular pathogen after interacting with organisms in the environment, including Acanthamoeba spp.
We investigated this question by co-cultivation of B. pseudomallei H777 (a clinical isolate, moderate biofilm producing wild-type), M10 (a biofilm-defect mutant of H777) and C17 (a biofilm-complemented of M10) (Table 1) with Acanthamoeba sp. to clarify the role of biofilm on B. pseudomallei-amoeba interaction. Bacterial adhesion, intracellular survival, metabolomic analyses of serially passaged B. pseudomallei, and persistence of bacterial biofilm grazed by amoebae were all investigated (Fig. 1). Our results demonstrated the persistence of B. pseudomallei biofilm formation against grazing by amoebae. The importance and originality of this study is that it explores for the first time the intracellular superoxide scavenger metabolites produced by B. pseudomallei following encounters with amoebae and demonstrates the persistence of B. pseudomallei biofilm despite grazing by amoebae. The presence of superoxide scavenger metabolites following passage through amoebae may indicate a pathway by which B. pseudomallei can become hypervirulent and a human pathogen.
Schematic flow chart of the co-cultivation of two different B. pseudomallei biofilm phenotypes and Acanthamoeba sp. Adhesion and intracellular-survival assays at MOI 100 using non-encapsulated biofilm cells were performed. Burkholderia pseudomallei were passaged through Acanthamoeba sp. up to three times and were then collected for metabolomic analysis using ultra-high-performance liquid chromatography-electrospray ionization-quadruple time-of-flight mass spectrometry in parallel with observations of colony morphology on Ashdown’s agar. In addition, preformed 24-h and 48-h B. pseudomallei biofilm was cocultured with Acanthamoeba sp. to monitor the biofilm structure and biofilm biomass using confocal laser scanning microscopy. Amoeba cells were counted using a hemocytometer.
Results
Non-encapsulated biofilm cells of B. pseudomallei H777 and M10 showed similar adhesion to and survive within Acanthamoeba sp. cells
Non-encapsulated biofilm cells of B. pseudomallei H777 and M10 cells co-cultured with Acanthamoeba sp. at MOI 100 for 1 h revealed similar levels of bacterial adhesion to Acanthamoeba sp.. However, this was not the case for the C17 strain (Fig. 2a and b). After the kanamycin protection assay, the number of B. pseudomallei of all three strains within the amoebae exhibited comparable levels at 1.5 h p.i. and 4.5 h p.i. (Fig. 2c).
Percentage of B. pseudomallei cells adhering to amoebae and intracellular survival after co-cultivation within Acanthamoeba sp. Similar percentages of planktonic B. pseudomallei H777 and M10 cells adhered to Acanthamoeba sp. after 1 h but not in the case of C17 (a). Bright field microscope visualization of B. pseudomallei H777, M10 and C17 adhering to Acanthamoeba sp. (black arrow) at 1,000 × magnification, scale bars = 10 µm (b); Cells of B. pseudomallei H777, M10 and C17 exhibited similar internalised at the early phase of infection at 1.5 h post-infection (p.i.) in Acanthamoeba sp. and similar survived at 4.5 p.i. (c). The experiment was performed in four replicates in three independent experiments (n = 12). Error bars represent mean ± SD. Statistical significance was tested using one-way ANOVA followed by Tukey post hoc test.
Non-encapsulated biofilm cells of Burkholderia pseudomallei H777, M10 and C17 co-cultured with Acanthamoeba sp. at MOI 100 for 1 h were then monitored under an inverted microscope for another 10 min. The results revealed the amoeba exhibited grazing actions on bacterial cells (Supplement video 1).
Burkholderia pseudomallei biofilm persist to Acanthamoeba sp. predation
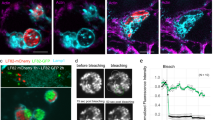
To investigate the persistence of B. pseudomallei biofilm despite grazing by amoebae, the 24-h and 48-h preformed biofilms of B. pseudomallei H777, M10 and C17 were co-cultured with Acanthamoeba sp. for an additional 24 h. Confocal images and COMSTAT analysis revealed the disruption of the 48-h amoeba-challenged biofilm biomass of both B. pseudomallei H777, M10 and C17 (Fig. 3a–g) and the 72-h amoeba-challenged biofilm biomass of both B. pseudomallei H777, M10 and C17 (Fig. 4a–g) compared to the untreated control (p < 0.01, 0.001). Notably, B. pseudomallei H777 and C17 biofilms were better able persist against grazing by amoebae than was that of M10 (Figs. 3h and 4h). The numbers of amoebae when co-cultured with the 48-h B. pseudomallei H777, M10 and C17 biofilms, amoebae numbers were comparable to those cultured alone (Fig. 3i,j). However, the co-cultured with the 72-h B. pseudomallei H777, M10 and C17 biofilms resulted in a significant increase in amoebae numbers to amoebae cultured alone (p < 0.05) (Fig. 4i,j). These findings suggest that the 72-h B. pseudomallei biofilms may serve as a food source for Acanthamoeba sp., leading to the increased amoeba cell numbers.
The 48-h B. pseudomallei H777, M10 and C17 biofilms after challenged with Acanthamoeba sp. The 24-h B. pseudomallei H777, M10 and C17 pre-formed biofilms were co-cultured with Acanthamoeba sp. for 24 h and then biofilm structure and biomass were assessed by CLSM, and numbers of amoebae counted using a hemocytometer. CLSM images of the 48-h B. pseudomallei H777, M10 and C17 biofilms (a–c). CLSM images of the 48-h amoeba-challenged B. pseudomallei H777, M10 and C17 biofilms (630 × magnification, scale bars = 10 µm.) (d–f). Biomass was compared between the co-cultured biofilms with amoebae and controls (g). Δ Biomass of B. pseudomallei H777, M10 and C17 after co-cultivation with amoebae (h). The biomass and Δ Biomass data from 72 images (24 image z-stacks from 4 cover slips in three independent experiments) were used in each analysis. Numbers of amoeba cells after incubation with bacterial biofilm from duplicates of the three independent experiments (n = 6) (i). Amoeba population after co-cultivation the biofilms (100 × magnification, scale bars = 50 µm) (j). Error bars represent mean ± SD. Statistical significance was tested using one-way ANOVA followed by Tukey post hoc test.
The 72-h B. pseudomallei H777, M10 and C17 biofilms after challenged with Acanthamoeba sp. The 48-h B. pseudomallei H777, M10 and C17 biofilms were co-cultured with Acanthamoeba sp. for 24 h and then biofilm structure and biomass were assessed by CLSM, and numbers of amoebae counted using a hemocytometer. CLSM images of the 72-h B. pseudomallei H777, M10 and C17 biofilms (a-c). CLSM images of the 72-h amoeba-challenged B. pseudomallei H777, M10 and C17 biofilms (630 × magnification, scale bars = 10 µm.) (d–f). Biomass was compared between the co-cultured biofilms with amoebae and controls (g). Δ Biomass of H777, M10 and C17 after co-cultivation with amoebae (h). The biomass and Δ Biomass data from 72 images (24 image z-stacks from 4 cover slips in three independent experiments) were used in each analysis. Numbers of amoeba cells after incubation with bacterial biofilm from duplicates of the three independent experiments (n = 6) (i). Amoeba population after co-cultivation the biofilms (100 × magnification, scale bars = 50 µm) (j). Error bars represent mean ± SD. Statistical significance was tested using one-way ANOVA followed by Tukey post hoc test.
Metabolic phenotypes of internalised B. pseudomallei H777 after three passages through Acanthamoeba sp
To monitor the metabolic alterations in the repeatedly internalised B. pseudomallei in Acanthamoeba sp., B. pseudomallei H777 was grown on Ashdown's agar after three passages through amoebae. The colony morphology of B. pseudomallei H777 liberated from each passage was similar to the control (Supplement Fig. 1a, b).
The intracellular metabolites following three passages of B. pseudomallei in Acanthamoeba sp. were subjected to UHPLC-ESI-QTOF-MS/MS analysis. Peak chromatography with retention time in positive and negative ESI modes showed no differences between experimental and control groups (Fig. 5).
Untargeted profile chromatogram of MS/MS spectra intensities with retention time. Untargeted profile chromatogram of MS/MS spectra intensities with retention time in positive (a and b) and negative (c and d) ionisation mode. Blue peak, B. pseudomallei H777 without Acanthamoeba sp. (n = 5). Red peak, B. pseudomallei H777 co-cultured with Acanthamoeba sp. (n = 5).
The metabolome datasets contained 1587 and 1052 features in positive and negative ESI modes, respectively (data not shown). Subsequently, PCA and O-PLS-DA models were constructed using Pareto the scaling method. PCA scores plot revealed that the metabolic profiles of B. pseudomallei and control groups were generally similar in both positive and negative ESI modes (Supplement Fig. 2a,b). In addition, the O-PLS-DA models showed no significant difference between groups in positive (p = 0.43) and negative (p = 0.83) ESI modes (Supplement Figs. 2c,d).
Relative concentrations of identified metabolites in each ESI mode analysed using fold-change with cut-off > 1.2 (Table 2). Nine metabolites of positive ESI mode including 3-hydroxyhexanoic acid (C6H12O3), N-(5,6-dioctyltriazin-4-yl) butanamide (C23H42N4O), 4-hydroxy-2,6-di (phenanthrene-9-yl)-4H-4lambda ~ 5 ~ -dinaphtho[2,1-d:1′,2′-f][1,3,2]dioxaphos -phepin-4-one (C48H29O4P), N-nervonoyl threonine (C28H53NO4), penta-2,4-diynoic acid (C5H2O2), MG(22:4(7Z,10Z,13Z,16Z)/0:0/0:0) (C25H42O4), cholin acetate (C7H17NO3), linoleyl carnitine (C25H45NO4), and acetic acid—1,2,2-triethoxyethan-1-ol (1/1) (C10H22O6) were significantly observed in internalised B. pseudomallei compared to that in LB growth. In addition, three metabolites of negative ESI mode including 8-O-4′-diferulic acid (C14H10Cl4), sumarotene (C24H30O2S), and 3-hydroxy-3-phenylpropanoic acid (C9H10O3) were detected.
In addition, we also performed univariate analysis to compare spectral intensities between B. pseudomallei and the control group of 12 metabolites using the Mann–Whitney U test (Table 2). The results revealed that acetic acid-1,2,2-triethoxyethan-1-ol (1/1) (p = 0.02), 8-O-4′-diferulic acid (p = 0.04), and sumarotene (p = 0.01) in B. pseudomallei were significantly different compared to the control group.
Discussion
Burkholderia pseudomallei is an environmental bacterium that thrives in soil and water, commonly establishes interactions with a variety of organisms, including plants and amoebae, particularly in melioidosis-endemic regions. Bacterial biofilm plays a crucial role in the survival of bacteria in diverse environments and contributes to their ability to cause diseases in human hosts. This study set out with the aim of assessing the role of B. pseudomallei biofilm on its survival against grazing by amoebae. The metabolic differences between the amoeba-internalised B. pseudomallei and control cultures were analysed. The results revealed that bacterial biofilm was dispersed after co-cultivation, but the B. pseudomallei biofilm-forming strain H777 and C17 persisted better against Acanthamoeba sp. grazing than did the biofilm-defect mutant strain, M10. A possible explanation for this finding might be that biofilm formation partly protects against grazing by amoebae. Furthermore, a significant increased level of 8-O-4′-diferulic acid, a superoxide scavenging metabolite, from the B. pseudomallei cells passaged three times in Acanthamoeba was observed. Hence, it could conceivably be hypothesized that grazing pressure from free-living amoebae may serve as a “training ground” stimulating the environmental saprophytic B. pseudomallei to produce compounds and may assist its survival in host cells.
Environmental saprophytes are commonly constrained by protozoan predation in natural food webs16. Biofilm formation provides bacterial cells with some shelter from these threats15. The persistence of the B. pseudomallei wild-type biofilm against grazing by Acanthamoeba sp. broadly supports the role of biofilm as an antipredator mechanism. The opportunistic bacterial pathogen, Vibrio cholerae, can survive protozoan grazing in biofilm form while non-encapsulated biofilm cells are eliminated. The environmental persistence of V. cholerae biofilms correlated with the principal cause of seasonal cholera epidemics16. Furthermore, the P. aeruginosa biofilms were demonstrated effectively defended against A. castellanii grazing25. This opportunistic pathogenic, P. aeruginosa was exhibited the type 3 secretory system components to kill biofilm-associated amoebae and may associate with the evolution of opportunistic bacterial pathogens26.
In this study, different biofilm phenotypes of B. pseudomallei (H777, M10 and C17) were apparently grazed and used as food by Acanthamoeba sp. as indicated by increased numbers of amoebae. This finding is consistent with our previous results on predator–prey relationships between B. pseudomallei and Acanthamoeba sp.27 and broadly supports the work of other studies in this area linking bacteria and Acanthamoeba sp.. Acanthamoeba castellanii was demonstrated as a biofilm grazer of mixed biofilms communities of Klebsiella pneumoniae, P. fluorescens and S. epidermidis28. Cell-free supernatant of A. castellanii, A. lenticulate and A. polyphaga disrupted the preformed biofilms of methicillin-resistant Staphylococcus aureus and Mycobacterium bovis. Biofilm dispersion by predatory amoebae highlights the potential for biofilm-busting, suggesting the possibility of identifying active molecules that can be applied as novel anti-biofilm compounds for management of biofilm-associated infections in conjunction with antimicrobial agents29. A possible explanation for the higher number of Acanthamoeba sp. cells following cultivation with B. pseudomallei biofilm is the consumption of non-encapsulated biofilm cells of B. pseudomallei by the amoebae after the dispersal of the biofilm. The presence of amoebae is crucial for maintaining nutrient cycling and balancing bacterial populations in ecosystems30. Acanthamoeba sp. may feed on extracellular polymeric substances (EPSs) in B. pseudomallei biofilm structure including capsular polysaccharides (CPS), exopolysaccharide, proteins, or lipids31.
The levels of adhesion of non-encapsulated biofilm cells of B. pseudomallei H777, M10 and C17 to Acanthamoeba sp. is consistent with that of our previous study which has suggested the biofilm phenotypes of B. pseudomallei on initial adhesion and invasion in human lung epithelial cells18. As environmental predators, trophozoites of Acanthamoeba spp. approach different microbes using their universal receptors to bind with various bacterial surface components including capsules, peptidoglycan, lipopolysaccharide and β-(1–4)-N-acetylmuramic acid20. The actively grazing of Acanthamoeba sp. towards and target B. pseudomallei cells provide additional evidence for the amoebae-bacteria interactions. While, a mass spectrophotometry-based metabolomics approach demonstrated that Burkholderia agricolaris and B. hayleyella use chemotaxis to actively search for their host, the social amoeba, Dictyostelium discoideum32. However, further work should be undertaken to investigate how Acanthamoeba sp. attack B. pseudomallei to widen the understanding of the ecological interaction that may transform an environmental saprophyte to a potential pathogen.
Bacteria subject to attack by protozoa have evolved defensive mechanisms that allow them to survive within protozoa. These mechanisms also pre-adapt them as opportunistic pathogens to escape the harmful attentions of phagocytes33,34,35. Ours is the first study using UHPLC ESI-QTOF-MS/MS-based metabolic profiling to investigate the differences between B. pseudomallei cells following interactions with amoebae and cells grown in LB without the presence of amoebae. A remarkably elevated amount of 8-O-4´-diferulic acid was detected in B. pseudomallei after repeated encounters with amoeba. Nevertheless, colonies of B. pseudomallei H777 after three passages in Acanthamoeba sp. demonstrated similar morphology.
Amoebae and mammalian phagocytes share core mechanisms and molecular processes concerning phagocytosis and intracellular killing of pathogens33. Bacteria that can evade the digestion process to survive in amoebae may use similar mechanisms to avoid or survive in nonphagocytic and mammalian phagocytic cells. Therefore, Acanthamoeba is recognized for its influence on the evolution, persistence, and transmission of potential human pathogens20,36. Bacterial pathogens engage antioxidant strategies using superoxide dismutase and catalase to neutralise reactive oxygen species such as superoxide (O2−), hydrogen peroxide (H2O2) and hydroxyl (HO·), which are crucial pathogen-eradication mediators24. Burkholderia pseudomallei exhibits superoxide dismutase activity to detoxified the superoxide for its intracellular survival and virulence37,38. Likewise, ferulic acid and dimers of ferulic acid, commonly obtained from plants, have antioxidant properties as superoxide-scavenging molecules39,40,41. The detection of 8-O-4′-diferulic acid, a superoxide scavenging metabolite, in B. pseudomallei passaged in amoebae may offer preliminary insights into a possible strategy for evading amoebae-mediated destruction and enhancing survival within host cells. This result corroborates the findings of much of the previous work by Wan et al.42 that demonstrated the antioxidant defence in V. cholerae by utilizing catalase to scavenge reactive oxygen species. Furthermore, V. cholerae biofilms produce pyomelanin pigment and reactive oxygen species correlated with resistance against A. castellanii predation43. In addition, as well as replicating in the amoeba A. castellanii, intracellular V. cholerae could ultimately return to the aquatic habitat using quorum sensing involving a Vibrio polysaccharide44. Vibrio cholerae that survived intracellular killing might gain specific strategies that enhance their hypervirulent performance in human hosts. Hence, environmental V. cholerae that have passed through protozoa may be preadapted to become human pathogens23. Furthermore, extensive investigations are required to fully comprehend the role of the metabolic changes observed in B. pseudomallei following passage through amoebae in enhancing B. pseudomallei survival.
A possible limitation of our metabolomics study is that we only used short periods of 4.5 h for the thrice-passaged B. pseudomallei in Acanthamoeba sp. to obtain the intracellular-surviving bacteria. In addition, culture of the liberated internalised bacteria to obtain sufficient bacterial cells for the MOI 100 co-cultivation and metabolomics analysis might have influenced the results. Metabolites in B. pseudomallei may be altered during different stages of bacterial growth45,46,47 leading to the appearance of similar metabolites between treated and control groups. Further work is required to investigate metabolites produced by amoeba-internalised bacteria without their subsequent growth in bacterial culture medium48. Moreover, in the biofilm grazing observations, conducting an enumeration of B. pseudomallei H777, M10 and C17 both with and without co-cultivation with Acanthamoeba sp. may offer valuable insights into the impact of amoeba grazing.
To the best of our knowledge, this is the first report on interactions between B. pseudomallei biofilm and Acanthamoeba sp. The principal theoretical implication of this study is that B. pseudomallei biofilm provides general protection against grazing by Acanthamoeba sp. Metabolomic analysis identified 8-O-4′-diferulic acid, a superoxide scavenging metabolite, that may play a role in predator-driven B. pseudomallei adaptation. The ability of B. pseudomallei to resist digestion by free-living amoebae may preadapt the bacterial pathogen to life as an intracellular pathogen.
Materials and methods
Ethics statement
Burkholderia pseudomallei H777 (from Melioidosis Research Center, Khon Kaen University) had been collected as a part of the study of the epidemiology of B. pseudomallei approved by the Khon Kaen University Ethics Committee for Human Research (HE490324). Patient cannot be identified as the isolates de-identified when we received them. All procedures were conducted following the appropriate guidelines and regulations.
Bacterial strains
Burkholderia pseudomallei H777, M10 and C17 isolates18,49 (Table 1) from glycerol stock at -80 ºC were cultured on Ashdown’s agar at 37 °C for 48 h. A single colony was cultured in 5 mL Luria Bertani (LB) broth at 37 °C, 200 rpm for 16–18 h before dilution to an optical density (OD600) 0.1 (≈ 1 × 107 CFU/mL) for 2% inoculum in fresh LB broth for 8 h to reach log phase. Subsequently, the bacterial cells were harvested and washed twice with sterile Page’s modified-Neff’s amoeba saline (PAS)50 at 3000 × g for 5 min and adjusted to OD600 = 0.1 for the co-cultivation experiment with amoebae. Burkholderia pseudomallei in LB at OD600 = 0.8–0.9 was used as the starter inoculum for biofilm establishment18,51.
Escherichia coli grown in LB broth at 37 °C, 200 rpm for 16–18 h were harvested and washed twice with PAS and were used to feed Acanthamoeba sp. to maintain the trophozoite stage27.
Cultivation of amoebae
Acanthamoeba sp. previously isolated from a B. pseudomallei-positive soil sample in Khon Kaen, Thailand27 was used in this study. The amoebae from soil stock were cultured on a non-nutrient agar plate with the addition of 0.03% trypticase soy broth (TSB) and E. coli as food. Plates were observed daily under a stereo microscope until the amoebae cells reached 70% confluence. The cells were then harvested and washed with PAS for further investigation.
Acanthamoeba sp. cells were grown in a gradually increased kanamycin concentration (from 30 to 300 µg/mL) administered via daily changes of PAS for 10 days27 to induce tolerance to 300 µg/mL kanamycin. The kanamycin pre-treated amoebae were used in co-cultivation experiments.
Monitoring adhesion and intracellular survival of two B. pseudomallei biofilm phenotypes
To monitor adhesion, the first step in the process of bacterial internalization of B. pseudomallei biofilm phenotype, amoeba cell suspensions in PAS (1 × 103 cells/well) were seeded in a 24-well tissue-culture plate and allowed to attach to the bottom of the culture plate for 15 min. Co-culture of non-encapsulated biofilm cells of B. pseudomallei H777 (wild-type strain) or M10 (biofilm-defect strain) with Acanthamoeba sp. was performed by adding mid-log suspensions of B. pseudomallei at a multiplicity of infection (MOI) of 100 and incubation for 1 h at 30 °C. Non-adherent bacteria were then removed by five gentle washes using PAS. Subsequently, amoeba cells were then lysed with 0.1% (v/v) Triton X-100 in PBS pH 7.4 for 20 s to liberate adherent bacteria. The percentage of adhered bacteria was calculated from the number of colony-forming units (CFUs) after incubated for 48 h at 37 °C on Ashdown’s agar using a drop plate technique, compared to the number of CFUs of the inoculum.
Microscopic observations of the adhesion experiments were also performed using a sterile coverslip placed in a 24-well plate before Acanthamoeba sp. and B. pseudomallei were co-cultured for 1 h. After washing with PAS, both amoeba and bacterial cells adhering to the coverslip were fixed with 1.25% (v/v) glutaraldehyde (EM grade; Electron Microscopy Sciences, Hatfield, PA) and stained with 0.1% crystal violet for 3 min. After washing with PBS buffer and air-drying at room temperature for 60 min, the coverslip was then mounted onto a glass slide and examined under a bright field microscope (Nikon, Eclipse Ni, Japan) at 100 × oil-immersion objective magnification.
To examine the intracellular survival of B. pseudomallei, bacteria were co-cultivated with Acanthamoeba sp. at MOI 100 for 1 h. After the non-adherent bacteria were removed, the amoebae were washed 3 times with PAS followed by the kanamycin protection assay to eradicate the extracellular bacteria using kanamycin at 300 µg/mL for 30 min. Subsequently, the initial internalized bacteria at 1.5 h post-infection (p.i.) and the intracellular survival after 3 h further incubation (4.5 h p.i.) were liberated using Triton X-100 and counted. The percentage of the internalized B. pseudomallei were reported compared to the inoculum.
To investigate the interaction between B. pseudomallei and Acanthamoeba sp., time-lapse video recording was performed. The amoeba cells were co-cultured with mid log-phase B. pseudomallei at MOI of 100 for 1 h at 30 °C. The interactions were then observed under an inverted-light microscope for another 10 min, 600 × magnification. The time-lapse video was displayed at 32 × speed.
Burkholderia pseudomallei biofilm formation versus Acanthamoeba sp. grazing
The liquid–air interface of B. pseudomallei biofilm was established using a 1 mL inoculum on a sterile glass coverslip in a 24-well plate with an Amsterdam Active Attachment (AAA) model at 37 °C for 24 h and 48 h51. The 24-h and 48-h pre-formed biofilms on the glass lid were washed once with sterile PBS before approximately 1 × 103 Acanthamoeba sp. cells/well in PAS were inoculated and incubated at 30 °C for an additional 24 h. The 48-h and 72-h biofilms challenged with the amoeba and controls on the glass coverslip were stained with 50 µg/mL FITC-ConA (Sigma, Saint Louis, Missouri, USA) for 20 min, fixed with 2.5% glutaraldehyde in PBS at room temperature for 3 h, washed three times with PBS and mounted with 80% glycerol. The biofilm structure was examined under confocal laser scanning microscope (CLSM, LSM 800, Carl Zeiss, Jena, Germany). The biofilm intensity and biomass of adherent cells were analysed using ZEN (version 2.1 blue edition) and COMSTAT software (version 2.1).
To assess the quantity of amoeba cells after their co-cultivation with pre-formed B. pseudomallei biofilms, the suspension from each well was collected and the number of amoeba cells was counted using a haemocytometer.
Sample preparation for LC–MS metabolite profiling
In our previous study, we observed that B. pseudomallei survived for up to 3 h post-infection but complete eradicated by 6 h within Acanthamoeba sp.27. We hypothesized that during this time frame, internalized B. pseudomallei might express certain metabolites crucial for its survival. The internalised B. pseudomallei H777 in Acanthamoeba sp. at 4.5 h p.i. were therefore liberated and grown on Ashdown’s agar at 37 °C for 48 h. A single colony was then taken and grown in 10 mL LB broth at 37 °C, 200 rpm for 6 h to achieve log-phase growth. The bacteria were again co-cultured with amoebae at MOI 100 for two additional cycles. After the third co-cultivation, the internalised bacterial cells were liberated, grown on Ashdown’s agar, and recovered in 10 mL LB broth and harvested for metabolomic analysis. Cells were washed three times with cold PBS pH 7.4 and centrifuged at 3,000 × g for 5 min at 4 °C before cell density was adjusted to OD600 0.5–0.6 (≈ 1 × 108 CFU/mL). The bacterial suspension was then centrifuged at 10,000 × g at 4 °C, for 10 min to obtain bacterial pellets. In parallel, B. pseudomallei H777 was grown on Ashdown’s agar and in LB broth for 3 rounds before harvested by centrifugation as an untreated control. The bacterial pellets were kept at − 80 °C for further metabolite extraction.
Metabolite extraction
The frozen B. pseudomallei cell pellets were resuspended in 200 µL of ice-cold methanol: water (1:1, v/v) and transferred to cryotubes containing 0.3 g of 0.5 mm sterile glass beads. Aqueous metabolite extraction was performed using a bead beater (OMNI bead rupture 24, Georgia) at 4.5 m/s for 30 s at 25 °C for 2 cycles followed by centrifugation at 20,000 × g, 4 °C for 10 min. The supernatant was transferred into a new tube before mixed with 200 of ice-cold chloroform, incubated on ice with interval vortex mixing every 3 min. After 20 min, the mixture was centrifuged at 20,000 × g, 4 °C for 10 min. The aqueous phase was dried using a CentriVap concentrator (LABCONCO, Missouri) at 45 °C for 3–4 h. Each sample was stored at -80 °C for further UHPLC-ESI-QTOF-MS/MS analysis. The dried extracts were reconstituted with 100 µL solvent mixture of water: acetonitrile (1:1, v/v), sonicated at room temperature for 10 min for 2 times, and centrifuged twice for 15 min at 20,000 × g, 4 °C47,52. From each sample, 15 µL was collected, pooled, and then used as a quality control (QC) sample.
LC–MS data acquisition
Metabolite profiling of B. pseudomallei samples during interaction with Acanthamoeba sp. was carried out using ultra-high-performance liquid chromatography coupled with electrospray ionization (ESI)-quadruple time-of-flight mass spectrometry (UHPLC ESI-QTOF-MS/MS) (Bruker, Germany) at Khon Kaen University Phenome Centre (KKUPC). In brief, the aqueous phase extracts of samples were analysed on a reverse-phase liquid chromatography platform using a Bruker intensity HPLC C18 (2.1 × 100 mm, 2 µm column). The column temperature was set at 40 ºC and the autosampler temperature was set at 4 °C. Mobile phase A was 100% water with 0.1% formic acid (FA) and mobile phase B was 100% acetonitrile with 0.1% FA. The flow rate was set at 0.35 mL/min and the elution gradient was set as follows: 99% A (0.0–2.0 min, 0.25 mL/min), 1% A (2.0–20.0 min, 0.25 mL/min), 99% A (20.1–28.3 min, 0.35 mL/min), 99% A (28.5–30.0 min, 0.25 mL/min). Two µL of samples were injected for both positive and negative ionisation polarity mode. The MS temperature was set at 220 °C, desolvation gas 8 L/min. Sodium formate solution (2 mM sodium hydroxide, 0.1% FA and 50% isopropanol) was directly injected as an external calibrant with the flow rate of 0.5 µL/min. The capillary voltage in positive and negative ionization polarity modes were 4000 and 4500 V, respectively. The data scan was set to mass range 50–1500 m/z)53.
Data pre-processing, metabolite assignment and multivariate statistical analysis
Raw data were imported to MetaboScape 7.0.1 software (Bruker, Massachusetts, US) for data pre-processing. In MetaboScape, the bucket table parameters were generated by using T-ReX _3D (LC-QTOF) workflow. Detection of molecular features was set 1500 counts of intensity threshold with a minimum peak length of 8 spectra. Assignment of metabolites was performed by comparing the MS/MS fragmentation patterns of detected features against the public database, human metabolome database (HMDB), METLIN, Bruker Metabobase and LipidBlast database. The level of assignment (LoA) included (1) accurate mass matched to database indicating tentative assignment, (2) accurate mass matched to database and tandem MS spectrum matched to in silico fragmentation pattern, (3) tandem MS spectrum matched to database or literature, (4) retention time and the molecular mass matched to standard compound, and (5) MS/MS spectrum matched to standard compound. The multivariate statistical analysis, including principal component analysis (PCA) and orthogonal signal correction-projection to latent structures-discriminant analysis (O-PLS-DA), were conducted using the Pareto scaling method in SIMCA software version 14.1 (Umetrics, Umeå, SE).
Univariate statistical analysis
Statistical analysis was performed in IBM SPSS statistics for Windows version 28 (SPSS Inc., Chicago, USA). Data was analysed from three independent experiments. The data was illustrated as a graph of the mean ± standard deviation (SD), using Graph Pad prism 5 (GraphPad Software Inc., California, USA). One-way ANOVA followed by Tukey post hoc test were used to identify significant differences among groups. Comparisons of metabolite spectra intensities were performed using a non-parametric test, the Mann–Whitney U test. A statistically significant difference was considered at p < 0.05.
Data availability
The datasets used and/or analysed during the current study available from the corresponding author on reasonable request.
References
Inglis, T. J. & Sagripanti, J. L. Environmental factors that affect the survival and persistence of Burkholderia pseudomallei. Appl. Environ. Microbiol. 72, 6865–6875. https://doi.org/10.1128/AEM.01036-06 (2006).
Kamjumphol, W., Chareonsudjai, P., Taweechaisupapong, S. & Chareonsudjai, S. Morphological Alteration and Survival of Burkholderia pseudomallei in Soil Microcosms. Am. J. Trop. Med. Hyg. 93, 1058–1065. https://doi.org/10.4269/ajtmh.15-0177 (2015).
Kamjumphol, W., Chareonsudjai, S., Chareonsudjai, P., Wongratanacheewin, S. & Taweechaisupapong, S. Environmental factors affecting Burkholderia pseudomallei biofilm formation. Southeast Asian. J. Trop. Med. Public Health 44, 72–81 (2013).
Suebrasri, T., Wang-ngarm, S., Chareonsudjai, P., Sermswan, R. W. & Chareonsudjai, S. Seasonal variation of soil environmental characteristics affect the presence of Burkholderia pseudomallei in Khon Kaen, Thailand. Afr. J. Microbiol. Res. 7, 1940–1945 (2013).
Wang-Ngarm, S., Chareonsudjai, S. & Chareonsudjai, P. Physicochemical factors affecting the growth of Burkholderia pseudomallei in soil microcosm. Am. J. Trop. Med. Hyg. 90, 480–485. https://doi.org/10.4269/ajtmh.13-0446 (2014).
Wiersinga, W. J. et al. Melioidosis. Nat. Rev. Dis. Primers 4, 17107. https://doi.org/10.1038/nrdp.2017.107 (2018).
Turner, P. et al. A retrospective analysis of melioidosis in Cambodian children, 2009–2013. BMC Infect. Dis. 16, 688. https://doi.org/10.1186/s12879-016-2034-9 (2016).
Allwood, E. M., Devenish, R. J., Prescott, M., Adler, B. & Boyce, J. D. Strategies for intracellular survival of Burkholderia pseudomallei. Front. Microbiol. 2, 170. https://doi.org/10.3389/fmicb.2011.00170 (2011).
Mariappan, V. et al. Hijacking of the Host’s Immune Surveillance Radars by Burkholderia pseudomallei. Front. Immunol. 12, 718719. https://doi.org/10.3389/fimmu.2021.718719 (2021).
Limmathurotsakul, D. et al. Predicted global distribution of Burkholderia pseudomallei and burden of melioidosis. Nat. Microbiol. 1, 15008. https://doi.org/10.1038/Nmicrobiol.2015.8 (2016).
Savelkoel, J., Dance, D. A. B., Currie, B. J., Limmathurotsakul, D. & Wiersinga, W. J. A call to action: Time to recognise melioidosis as a neglected tropical disease. Lancet Infect Dis 22, e176–e182. https://doi.org/10.1016/S1473-3099(21)00394-7 (2022).
Pumpuang, A. et al. Survival of Burkholderia pseudomallei in distilled water for 16 years. Trans. R. Soc. Trop. Med. Hyg. 105, 598–600. https://doi.org/10.1016/j.trstmh.2011.06.004 (2011).
Kaestli, M. et al. Out of the ground: aerial and exotic habitats of the melioidosis bacterium Burkholderia pseudomallei in grasses in Australia. Environ. Microbiol. 14, 2058–2070. https://doi.org/10.1111/j.1462-2920.2011.02671.x (2012).
Inglis, T. J. J. et al. Interaction between Burkholderia pseudomallei and Acanthamoeba species results in coiling phagocytosis, endamebic bacterial survival, and escape. Infect. Immun. 68, 1681–1686. https://doi.org/10.1128/IAI.68.3.1681-1686.2000 (2000).
Hall-Stoodley, L., Costerton, J. W. & Stoodley, P. Bacterial biofilms: from the natural environment to infectious diseases. Nat. Rev. Microbiol. 2, 95–108. https://doi.org/10.1038/nrmicro821 (2004).
Matz, C. et al. Biofilm formation and phenotypic variation enhance predation-driven persistence of Vibrio cholerae. 102, 16819-16824, https://doi.org/10.1073/pnas.0505350102 (2005).
Matz, C., Bergfeld, T., Rice, S. A. & Kjelleberg, S. Microcolonies, quorum sensing and cytotoxicity determine the survival of Pseudomonas aeruginosa biofilms exposed to protozoan grazing. Environ. Microbiol. 6, 218–226. https://doi.org/10.1111/j.1462-2920.2004.00556.x (2004).
Kunyanee, C. et al. Burkholderia pseudomallei biofilm promotes adhesion, internalization and stimulates proinflammatory cytokines in human epithelial A549 cells. PLoS One 11, e0160741. https://doi.org/10.1371/journal.pone.0160741 (2016).
Molmeret, M., Horn, M., Wagner, M., Santic, M. & Abu Kwaik, Y. Amoebae as training grounds for intracellular bacterial pathogens. Appl. Environ. Microbiol. 71, 20–28. https://doi.org/10.1128/AEM.71.1.20-28.2005 (2005).
Rayamajhee, B. et al. Acanthamoeba, an environmental phagocyte enhancing survival and transmission of human pathogens. Trends Parasitol. 38, 975–990. https://doi.org/10.1016/j.pt.2022.08.007 (2022).
Di Meo, S., Reed, T. T., Venditti, P. & Victor, V. M. Role of ROS and RNS sources in physiological and pathological conditions. Oxid. Med. Cell. Longev. 2016 (2016).
Zhang, X. & Soldati, T. Detecting, visualizing and quantitating the generation of reactive oxygen species in an amoeba model system. J. Visualized Exp. 81, e50717 (2013).
Espinoza-Vergara, G., Hoque, M. M., McDougald, D. & Noorian, P. The impact of protozoan predation on the pathogenicity of Vibrio cholerae. Front. Microbiol. https://doi.org/10.3389/fmicb.2020.00017 (2020).
Imlay, J. A. Cellular defenses against superoxide and hydrogen peroxide. J Annu. Rev. Biochem. 77, 755–776 (2008).
Weitere, M., Bergfeld, T., Rice, S. A., Matz, C. & Kjelleberg, S. Grazing resistance of Pseudomonas aeruginosa biofilms depends on type of protective mechanism, developmental stage and protozoan feeding mode. Environ. Microbiol. 7, 1593–1601. https://doi.org/10.1111/j.1462-2920.2005.00851.x (2005).
Matz, C. et al. Pseudomonas aeruginosa uses type III secretion system to kill biofilm-associated amoebae. ISME J. 2, 843–852. https://doi.org/10.1038/ismej.2008.47 (2008).
Noinarin, P., Chareonsudjai, P., Wangsomnuk, P., Wongratanacheewin, S. & Chareonsudjai, S. Environmental free-living amoebae isolated from soil in Khon Kaen, Thailand, antagonize Burkholderia pseudomallei. PLoS One 11, e0167355. https://doi.org/10.1371/journal.pone.0167355 (2016).
Huws, S. A., McBain, A. J. & Gilbert, P. Protozoan grazing and its impact upon population dynamics in biofilm communities. J. Appl. Microbiol. 98, 238–244. https://doi.org/10.1111/j.1365-2672.2004.02449.x (2005).
Martin, K. H., Borlee, G. I., Wheat, W. H., Jackson, M. & Borlee, B. R. Busting biofilms: Free-living amoebae disrupt preformed methicillin-resistant Staphylococcus aureus (MRSA) and Mycobacterium bovis biofilms. Microbiol. (Reading) 166, 695–706. https://doi.org/10.1099/mic.0.000933 (2020).
Siddiqui, R. & Khan, N. A. Biology and pathogenesis of Acanthamoeba. Parasit. Vectors 5, 6. https://doi.org/10.1186/1756-3305-5-6 (2012).
Mangalea, M. R., Borlee, G. I. & Borlee, B. R. The current status of extracellular polymeric substances produced by Burkholderia pseudomallei. Curr. Tropical Med. Rep. 4, 117–126. https://doi.org/10.1007/s40475-017-0118-2 (2017).
Shu, L., Zhang, B., Queller, D. C. & Strassmann, J. E. Burkholderia bacteria use chemotaxis to find social amoeba Dictyostelium discoideum hosts. ISME J. 12, 1977–1993. https://doi.org/10.1038/s41396-018-0147-4 (2018).
Greub, G. & Raoult, D. Microorganisms resistant to free-living amoebae. Clin. Microbiol. Rev. 17, 413–433. https://doi.org/10.1128/CMR.17.2.413-433.2004 (2004).
Sun, S., Noorian, P. & McDougald, D. Dual role of mechanisms involved in resistance to predation by protozoa and virulence to humans. Front. Microbiol. 9, 1017. https://doi.org/10.3389/fmicb.2018.01017 (2018).
Strassmann, J. E. & Shu, L. Ancient bacteria-amoeba relationships and pathogenic animal bacteria. PLoS Biol. 15, e2002460. https://doi.org/10.1371/journal.pbio.2002460 (2017).
Erken, M., Lutz, C. & McDougald, D. The rise of pathogens: Predation as a factor driving the evolution of human pathogens in the environment. Microb. Ecol. 65, 860–868. https://doi.org/10.1007/s00248-013-0189-0 (2013).
Vanaporn, M. et al. Superoxide dismutase C is required for intracellular survival and virulence of Burkholderia pseudomallei. Microbiology 157, 2392–2400. https://doi.org/10.1099/mic.0.050823-0 (2011).
Adl, S. M. et al. The revised classification of eukaryotes. J. Eukaryot. Microbiol. 59, 429–493. https://doi.org/10.1111/j.1550-7408.2012.00644.x (2012).
Garcia-Conesa, M. T., Plumb, G. W., Waldron, K. W., Ralph, J. & Williamson, G. Ferulic acid dehydrodimers from wheat bran: Isolation, purification and antioxidant properties of 8-O-4-diferulic acid. Redox Rep. 3, 319–323. https://doi.org/10.1080/13510002.1997.11747129 (1997).
Andreasen, M. F., Kroon, P. A., Williamson, G. & Garcia-Conesa, M.-T. Intestinal release and uptake of phenolic antioxidant diferulic acids. Free Radical Biol. Med. 31, 304–314. https://doi.org/10.1016/S0891-5849(01)00585-8 (2001).
Cano, A., Arnao, M. B., Williamson, G. & Garcia-Conesa, M. T. Superoxide scavenging by polyphenols: Effect of conjugation and dimerization. Redox Rep. 7, 379–383. https://doi.org/10.1179/135100002125001153 (2002).
Wang, H. et al. Catalases promote resistance of oxidative stress in Vibrio cholerae. PloS One 7, e53383 (2012).
Noorian, P. et al. Pyomelanin produced by Vibrio cholerae confers resistance to predation by Acanthamoeba castellanii. FEMS Microbiol. Ecol. 93, 147. https://doi.org/10.1093/femsec/fix147 (2017).
Van der Henst, C., Scrignari, T., Maclachlan, C. & Blokesch, M. An intracellular replication niche for Vibrio cholerae in the amoeba Acanthamoeba castellanii. ISME J. 10, 897–910. https://doi.org/10.1038/ismej.2015.165 (2016).
Gjersing, E. L., Herberg, J. L., Horn, J., Schaldach, C. M. & Maxwell, R. S. NMR metabolomics of planktonic and biofilm modes of growth in Pseudomonas aeruginosa. Anal. Chem. 79, 8037–8045. https://doi.org/10.1021/ac070800t (2007).
Yeom, J., Shin, J. H., Yang, J. Y., Kim, J. & Hwang, G. S. 1H NMR-based metabolite profiling of planktonic and biofilm cells in Acinetobacter baumannii 1656–2. PLoS One 8, e57730. https://doi.org/10.1371/journal.pone.0057730 (2013).
Wong, E. H. J. et al. Metabolomic analysis of low and high biofilm-forming Helicobacter pylori strains. Sci. Rep. 8, 1409. https://doi.org/10.1038/s41598-018-19697-0 (2018).
Biggins, J. B., Liu, X., Feng, Z. & Brady, S. F. Metabolites from the induced expression of cryptic single operons found in the genome of Burkholderia pseudomallei. J. Am. Chem. Soc. 133, 1638–1641. https://doi.org/10.1021/ja1087369 (2011).
Taweechaisupapong, S. et al. Virulence of Burkholderia pseudomallei does not correlate with biofilm formation. Microb. Pathog. 39, 77–85. https://doi.org/10.1016/j.micpath.2005.06.001 (2005).
Page, F. A New Key to Freshwater and Soil Amoebae (Freshwater Biological Association, Scientific Publication, 1988).
Pakkulnan, R. et al. Extracellular DNA facilitates bacterial adhesion during Burkholderia pseudomallei biofilm formation. PLoS One 14, e0213288. https://doi.org/10.1371/journal.pone.0213288 (2019).
Vorkas, P. A. et al. Untargeted UPLC-MS profiling pipeline to expand tissue metabolome coverage: Application to cardiovascular disease. Anal. Chem. 87, 4184–4193. https://doi.org/10.1021/ac503775m (2015).
Phukhum, P. et al. The impact of hypoxia and oxidative stress on proteo-metabolomic alterations of 3D cholangiocarcinoma models. Sci. Rep. 13, 3072. https://doi.org/10.1038/s41598-023-30204-y (2023).
Acknowledgements
We would like to acknowledge Prof. David Blair for editing the MS via Publication Clinic KKU, Thailand.
Funding
This work was supported by Development and Promotion of Science and Technology Talents Project, Royal Thai Government, Bangkok, Thailand (Scholarship #592008) to C.B.
Author information
Authors and Affiliations
Contributions
Conceptualization, S.C., C.B. and J.P.; investigation, CB and PN; interpretation, S.C., C.B. and J.P.; writing-original draft preparation, C.B. and S.C.; writing-review and editing, S.C., C.B. and J.P.; funding acquisition, S.C. and C.B.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Bunma, C., Noinarin, P., Phetcharaburanin, J. et al. Burkholderia pseudomallei biofilm resists Acanthamoeba sp. grazing and produces 8-O-4′-diferulic acid, a superoxide scavenging metabolite after passage through the amoeba. Sci Rep 13, 16578 (2023). https://doi.org/10.1038/s41598-023-43824-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-023-43824-1
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.