Abstract
Megalopas of 15 brachyuran crab species collected in the open sea plankton, and unknown until now, were identified using DNA barcodes (COI and 16S rRNA). Specimens belonging to the families Portunidae, Pseudorhombilidae and Xanthidae (Crustacea, Decapoda, Brachyura), and corresponding to the species Achelous floridanus, Arenaeus mexicanus, Callinectes amnicola, C. arcuatus, C. ornatus, C. toxones, Charybdis (Charybdis) hellerii, Portunus hastatus, Thalamita admete, Scopolius nuttingi, Etisus odhneri, Liomera cinctimanus, Neoliomera cerasinus, Pseudoliomera variolosa, and Williamstimpsonia stimpsoni, are described and illustrated, and compared with other congeneric species previously described. We also provide a new geographical record for N. cerasinus and the most remarkable features for each species.
Similar content being viewed by others
Introduction
One of the most relevant and influential scientific method in the last decade is DNA barcoding. It is considered an effective tool for species identification in different animal groups1,2 and is becoming increasingly common in biodiversity and conservation science3,4. Since its introduction 17 years ago, DNA barcoding has been widely applied by taxonomists as indicated by hundreds of published taxonomic studies5,6,7,8.
In this context, crustaceans, that represent one of the most diverse metazoan groups from a morphological and ecological point of view9 with more than 67,000 described species so far10, are an interesting target taxon for DNA barcoding because they are not always easy to identify by traditional approaches and usually require the help of highly trained taxonomists11. One of the biggest problems is to identify the larval stages of this group, because the larvae are distinguishable but not easily matched with the correct adult form12,13. Therefore, this problem causes obstacles in studies such as plankton ecology or population connectivity14,15.
This is the case of crabs. Most brachyuran crabs pass through a planktonic larval period with two phases, zoea and megalopa, which are remarkably different from each other and from the adult form16,17. The megalopa is a planktonic phase characterized by the existence of functional pleonal swimming appendages, the pleopods, while the anterior thoracic appendages (the maxillipeds) assume functions as mouthparts18,19. This stage usually looks for structurally complex habitats, which can provide refuge and food20 and many studies refer to the megalopa as settle and recruitment phase21,22.
Particularly, the identification of megalopas from plankton samples based on morphological characters is a difficult task and in many cases is not possible do it at genus or species level19,23. In this sense, DNA barcoding provide rapid and accurate identifications of plankton specimens24,25,26,27, being the only limitation the need of previous knowledge of DNA markers for the species in accessible databases as Genbank or BOLD.
In this study DNA barcoding was used to identify the megalopa stage of brachyuran crabs from open sea plankton across the world, collected in the context of the MALASPINA and MAF research projects. In this work, we focus on describing unknown megalopas belonging to the families Portunidae Rafinesque, 1815, Pseudorhombilidae Alcock, 1900 and Xanthidae MacLeay, 1838. Portunidae are among the most diverse and species rich groups of brachyuran crabs with a worldwide distribution, including many taxa which are of high ecological and economical significance28. Portunids larvae identification is particularly difficult29,30 because the larvae of different species are so similar, that it is difficult to tell species apart other than by examination of minute characteristics31,32. On the other hand, representatives of the Xanthidae present a circumtropical distribution while Pseudorhombilidae are known almost exclusively from waters of the Americas. Commonly known as mud, pebble, rubble, or blackfingered crabs33, are familiar forms in many marine settings although many species remain poorly described and lack detailed illustrations34.
Once identified by DNA barcoding, in the present work a morphological description and illustrations are carried out for the megalopas of 15 species, namely the portunids: Achelous floridanus (Rathbun, 1930), Arenaeus mexicanus (Gerstaecker, 1856), Callinectes amnicola (de Rochebrune, 1883), Callinectes arcuatus Ordway, 1863, Callinectes ornatus Ordway, 1863, Callinectes toxones Ordway, 1863, Charybdis (Charybdis) hellerii (A. Milne-Edwards, 1867), Portunus hastatus (Linnaeus, 1767), and Thalamita admete (Herbst, 1803), the xanthids: Etisus odhneri Takeda, 1971, Liomera cinctimanus (White, 1847), Neoliomera cerasinus Ng, 2002, Pseudoliomera variolosa (Borradaile, 1902), and Williamstimpsonia stimpsoni (A. Milne-Edwards, 1879), and the pseudorhombilid: Scopolius nuttingi (Rathbun, 1898).
Results
A total of 462 megalopas were collected in the course of two different projects, 375 in the MALASPINA Expedition, and 87 in the MAF cruise.
These megalopas were initially sorted according to their general external morphology in main morphotypes groups and, from each of these; representatives were selected for DNA barcoding. Partial mitochondrial COI and/or 16S rRNA gene sequences were obtained for 139 larvae, leading to the identification of 67 megalopas from 34 species.
DNA barcode identification
Among a total of 139 megalopas analysed by DNA barcoding, 72 could not be identified to species level only based on morphological features, since their DNA barcodes did not allow for accurate identification, and therefore cannot be described. The other 67 were identified as belonging to 34 different species of the families Calappidae De Haan, 1833 [in De Haan, 1833–1850] (4), Cryptochiridae Paulson, 1875 (3), Dromiidae De Haan, 1833 [in De Haan, 1833–1850] (1), Eriphiidae MacLeay, 1838 (1), Grapsidae MacLeay, 1838 (6), Homolidae De Haan, 1839 [in De Haan, 1833–1850] (1), Ocypodidae Rafinesque, 1815 (1), Panopeidae Ortmann, 1893 (1), Parthenopidae MacLeay, 1838 (2), Portunidae (9), Pseudorhombilidae (1), and Xanthidae (5).
Of these 34 species, only 3, Menippe nodifrons Stimpson, 1859, Eurypanopeus abbreviatus (Stimpson, 1860), and Homola barbata (Fabricius, 1793), have its megalopa previously described35,36,37, and when they were compared no significant differences were found, for this reason do not need to be redescribed. Their sequences have been deposited in Genbank: M. nodifrons (16S: MW264136, COI: MW264437), E. abbreviatus (16S: MW264137, COI: MW264438), and H. barbata (COI: MW264436). The present work focuses on the 31 megalopas of Portunidae, Pseudorhombilidae and Xanthidae assigned to 15 species (Table 1).
While 16S sequences were obtained for all 31 megalopas analyzed, COI could not be reared for five species, specially xanthoids (see Table 1). Using 16S sequences eight species were identified fitting 100% with sequences in Genbank, but only four with COI in Genbank and BOLD, because it is a marker with more intraspecific variability.
The results of the BLAST search show some inaccuracies. In the case of the sequences of the megalopa of a Thalamita Latreille, 1829 species we obtained two sequences fitting 100% in 518 bp and 531 bp for the 16S sequence. The first one corresponded to Thalamita admete, specimen ULLZ 4382 from South Africa, obtained by Mantelatto et al.38, the second one to Thalamita gatavakensis Nobili, 1906, specimen UF: 17,469 collected in Lizard Island (Australia), obtained by Evans39. In the case of COI sequence, only one sequence fitted 99% (5 mutations in 657 bp) and belong to UF: 17,486, another specimen of Thalamita gatavakensis from Lizard Island, also obtained by Evans39. Taken into account that the megalopa was collected close to South Africa we have considered T. admete as the right identification, but future studies are needed to clarify the relationship between T. admete and T. gatavakensis. Similarly, for one megalopa its 16S sequence fit 99% (only 2 mutations in 462 bp) with one sequence of Liomera cinctimanus (as Liomera cinctimana) obtained by Lai et al.40. However, other two sequences are deposited in Genbank as belonging to L. cinctimanus, with 19 mutations in 518 bp and 21 mutations in 519 bp, both obtained by Wetzer et al.41. In all cases, the specimens were collected in Guam, but they clearly do not belong to the same species according to the differences found, higher than intraspecific variability. The megalopa was collected close to South Africa, therefore so far from Guam, but within of the wide distribution of this species. Scopolius nuttingi, represent a third similar case, since two sequences of 16S were found fitting 99% (only 1 mutation) with that of the megalopa. The two sequences are identical, but one is identified as Micropanope nuttingi (MF490190 obtained by Mantelatto et al.8), and another one as M. scultipes (KT279707 obtained by Faria et al.42). However, a third sequence (GU144437), shorter and with only one mutation in the same position, is identified as M. nuttingi by Felder and Thoma43 and the COI sequence of the megalopa fit 100% with two sequences of M. nuttingi. For these reasons we have considered Scopolius nuttingi as the correct identification.
New record
A single cave-dwelling megalopa of Neoliomera cerasinus was caught on 13 February 2011 in South African coast (35° 08′ 10′′ S, 25° 33′ 47′′ E). This specimen constitutes the first occurrence of Neoliomera cerasinus from the Indian Ocean coast of South Africa. Previous records of this cave-dwelling xanthid crab from the Indian Ocean were from Christmas Island: Thunderdome Cave [topotypical locality] and West White Cave44, and in the Pacific Ocean in Kumejima Island, Ryukyu Islands, Japan45 and Okinawa Island and Shimoji Island, Ryukyu Islands46, expanding widely the distribution of this species to the opposite extreme of the Indian Ocean (see Fig. 1).
Megalopas descriptions
Family Portunidae Rafinesque, 1815
Genus Achelous De Haan, 1833
Achelous floridanus (Rathbun, 1930)
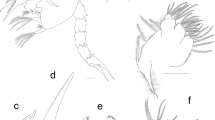
Appendages of Achelous floridanus megalopa: (a) antennule; (b) antenna; (c) mandible; (d) maxillule; (e) maxilla; (f) first maxilliped; (g) second maxilliped; (h) third maxilliped; (i) cheliped (j) second pereiopod; (k) fifth pereiopod; (l) second pleopod; (m) uropod. Scale bars: (c–e), 0.1 mm; (a, b, h, j–m) 500 µm; (i) 1 mm.
Lateral view of megalopa: (a) Achelous floridanus; (b) Arenaeus mexicanus; (c) Portunus hastatus; (d) Charybdis (Charybdis) hellerii; (e) Thalamita admete; (f) Callinectes toxotes. Rostrum dorsal view: (g) Achelous floridanus. Antenna: (h) Arenaeus mexicanus; (i) Portunus hastatus; (j) Charybdis (Charybdis) hellerii; (k) Thalamita admete; (l) Callinectes toxotes; (m) Callinectes arcuatus; (n) Callinectes ornatus; (o) Callinectes amnicola. Cheliped: (p) Portunus hastatus; (q) Callinectes amnicola. Fifth pereiopod: (r) Callinectes amnicola. Scale bars: (a–f, p–r, I) 1 mm; (h–e, m–n, o) 0.5 mm.
Size: CL: 3.86 ± 0.15 mm; CW: 2.84 ± 0.16 mm; n = 5.
Cephalothorax (Figs. 2a, 4a, g) Longer than broad, with long, thin and slightly curved upwards spine rostral with ventrally minute setae; orbital region with 7 plumose setae; a pair of lobes on the mesobranchial regions with hepatic regions moderately inflated; setation as drawn; dorsal organ present; eyes stalked.
Antennule (Fig. 3a) Peduncle 3-segmented with 16 plumose + 10 simple setae around the first segment, 4 plumose + 1 simple setae on second segment and 2 long plumose + 1 simple setae on the distal segment; primary flagellum with 5 annuli, with 0, 0, 1, 3, 1 simple + 1 sparsely setae and 0, 18, 16, 12, 6 aesthetascs respectively; accessory flagellum without annuli with 1 medial + 1 subterminal + 4 terminal simple setae.
Antenna (Fig. 3b) Peduncle 3-segmented with 5 simple + 1 plumodenticulate, 2 plumose + 1 simple, 1 simple + 5 plumose setae; flagellum 8-segmented with 0, 2, 4, 2, 3 + 2 (long serrated), 3, 4, 4–5 simple setae respectively.
Mandible (Fig. 3c) Palp 3-segmented with 22–24 plumodenticulate marginal setae on distal segment.
Maxillule (Fig. 3d) Coxal endite with 22–23 setae; basial endite with 6 small setae + 2 long sparsely plumose lateral setae and 22–24 cuspidate setae plus other one on the dorsolateral margin; endopod 2-segmented with 4 + 1 long plumose setae on proximal segment and 2 terminal simple setae on distal segment; long exopodal sparsely plumose seta present.
Maxilla (Fig. 3e) Coxal endite bilobed with 12 + 7 terminal plumose setae; basial endite bilobed with 10–12 + 15–16 sparsely plumodenticulate setae; endopod unsegmented with 3 short plumodenticulate setae on dorsal margin; exopod (scaphognathite) with 105–108 marginal plumose setae plus 19–20 setae on lateral surface.
First maxilliped (Fig. 3f) Epipod triangular shaped with 9 proximal plumodenticulate and 29–30 distal long simple setae; coxal endite with 20–24 plumose setae; basial endite with 37–42 sparsely plumodenticulate setae; endopod unsegmented with 2 + 4 simple setae; exopod 3-segmented with 4 plumose distal setae on proximal segment and 5 terminal plumose setae on distal segment.
Second maxilliped (Fig. 3g) Epipod reduced without setae; coxa with 2 + 2 + 4 simple setae; endopod 5-segmented with 2 simple, 2 simple, 4 long simple, 12 plumodenticulate and 11 (6 cuspidate and 5 plumodenticulate) setae, respectively; exopod 3-segmented with 1 medial simple seta on proximal segment and 5 terminal plumose setae on distal segment.
Third maxilliped (Fig. 3h) Epipod with 5 plumodenticulate + 20–21 long simple setae; protopod with 16–20 plumodenticulate setae; endopod 5-segmented with 41–45, 22–27, 17–19, 18–20, 13–15 sparsely plumose setae respectively; exopod 3-segmented with 4 marginal simple setae on proximal segment and 6 terminal plumose setae on distal segment.
Pereiopod (Figs. 3i–k, 8a) Cheliped setation as drawn; pereiopod II with hook coxal and pereiopod III with small tubercle on coxal segment; pereiopods II-IV with propodial setae present; pereiopods II-V thin and setose, inner margin of dactyl with 12–13 stout ventral spines; dactylus of pereiopod V with 3 long setae (feelers) plus 5 shorts + 3 long small setae like feelers.
Sternum (Fig. 8a) Maxilliped sternites completely fused with 10 simple setae plus one central pair of setae, cheliped sternites with 7 simple setae each, pereiopod sternites 2–3 with small tubercle with 7 simple setae each; pereiopod sternite IV with a long pointed posterolateral spine with 4 setae; sternal sutures are interrupted medially.
Pleon (Fig. 2a) Six pleonites; pleonite I without setae; setation of pleonites II-VI as shown; pleonite VI reduced.
Pleopods (Fig. 3l, m) Present on pleonites II-VI; endopods unsegmented with 4–5 cincinuli; exopod unsegmented with 36–37, 36–38, 32–34, 27–29 long plumose natatory setae; uropod 2-segmented, proximal segment with 1 seta, distal segment with 17–18 terminal plumose natatory setae.
Telson (Fig. 3m) Reduced, subquadrate, with 1 pair of dorsal setae and 7–8 setae on posterior margin.
Distinctive morphological features for Arenaeus mexicanus, Portunus hastatus, Charybdis (Charybdis) hellerii, Thalamita admete, Callinectes amnicola, Callinectes arcuatus, Callinectes ornatus, and Callinectes toxotes, are listed in the Table 2 and illustrated in the Figs. 2b–I, 4b–o, 8b–i.
Superfamily Xanthoidea MacLeay, 1838
Family Xanthidae MacLeay, 1838
Genus Neoliomera Odhner, 1925
Neoliomera cerasinus Ng, 2002
Appendages of Neoliomera cerasinus: (a) antennule; (b) antenna; (c) mandible; (d) maxillule; (e) maxilla; (f) first maxilliped; (g) second maxilliped; (h) third maxilliped; (i) cheliped; (j) dactylus of second pereiopod; (j,1) detail of dactylus of second pereiopod; (k), dactylus of fifth pereiopod; (l) second pleopod; (m) uropod. Scale bars: 0.1 mm.
Lateral view of megalopa: (a) Neoliomera cerasinus; (b) Liomera cinctimanus; (c) Pseudoliomera variolosa; (d) Williamstimpsonia stimpsoni; (e) Scopolius nuttingi; (f) Etisus odhneri. Antenna: (g) Liomera cinctimana; (h) Pseudoliomera variolosa; (i) Williamstimpsonian stimpsoni; (j) Scopolius nuttingi; (k) Etisus odhneri. Cheliped: (l) Williamstimpsonian stimpsoni; (p) Etisus odhneri. Fifth pereiopod: (m) Williamstimpsonian stimpsoni; (o) Pseudoliomera variolosa. Dactylus of second pereiopod: (n) Pseudoliomera variolosa. Scale bars: 0.5 mm.
Sternum of megalopa: (a) Achelous floridanus; (b) Arenaeus mexicanus; (c) Portunus hastatus; (d) Charybdis (Charybdis) hellerii; (e) Thalamita admete; (f) Callinectes toxotes; (g) Callinectes arcuatus; (h) Callinectes ornatus; (i) Callinectes amnicola; (j) Neoliomera cerasinus; (k) Liomera cinctimana; (l) Pseudoliomera variolosa; (m) Williamstimpsonian stimpsoni; (n) Scopolius nuttingi; (o) Etisus odhneri. Scale bars: 0.5 mm.
Size: CL: 1.34 mm; CW: 1.05 mm; n = 1.
Cephalothorax (Figs. 5a, 7a) Longer than broad; frontal margin with a pair of frontal submedian horns with a rostrum directed obliquely downwards; prominent tubercle on mesogastric, and a pair of lobes on protogastric and mesobranchial region; small tubercle on cardiac region; hepatic region inflated; dorsal organ present; eyes stalked.
Antennule (Fig. 6a) Peduncle 3-segmented with 5, 2, 2 simple setae respectively; primary flagellum with 4 annuli, with 0, 0, 2, 1 + 1 simple setae and with 0, 9, 7, 4 aesthetascs respectively; accessory flagellum without annuli, with 1 medial + 1 subterminal + 3 terminal simple setae.
Antenna (Fig. 6b) Peduncle 3-segmented with 2, 2, 2 simple setae; proximal segment with stout process; flagellum 8-segmented with 0, 0, 3 simple, 0, 2 long plumodenticulate + 3 short simple, 0, 4 simple, 4 simple setae respectively.
Mandible (Fig. 6c) Palp 2-segmented with 8 plumodenticulate terminal setae on distal segment.
Maxillule (Fig. 6d) Epipodal sparsely plumodenticulate seta present; coxal endite with 17 setae; basial endite with 4 small ventrolateral setae + 10 cuspidate and 9 plumodenticulate terminal setae + 3 long plumose setae; endopod 2-segmented with 2 simple setae on proximal segment and 1 long simple seta + 2 short terminal simple setae on distal segment; 2 long exopodal setae.
Maxilla (Fig. 6e) Coxal endite bilobed with 10 + 6 terminal plumose setae; basial endite bilobed with 6 + 10 sparsely plumodenticulate setae; endopod unsegmented with 3 short plumodenticulate setae on dorsal margin; exopod (scaphognathite) with 50–51 marginal plumose setae plus 2 + 2 setae on lateral surface.
First maxilliped (Fig. 6f) Epipod triangular shaped with 1 proximal and 7 distal long simple setae; coxal endite with 6 long plumose + 5 long simple setae; basial endite with 17 sparsely plumodenticulate setae; endopod unsegmented with 3 simple setae; exopod 2-segmented with 2 distal plumose setae on proximal segment and 4 terminal plumose setae on distal segment.
Second maxilliped (Fig. 6g) Epipod reduced with 2 setae; coxa without setae; endopod 5-segmented with 1 simple, 3 simple + 1 long plumodenticulate, 5 plumodenticulate and 9 (7 cuspidate and 2 plumodenticulate) setae, respectively; exopod 2-segmented with 1 medial simple seta on proximal segment and 5 terminal plumose setae on distal segment.
Third maxilliped (Fig. 6h) Epipod with 5 proximal plumose and 13 terminal long simple setae; protopod with 10 plumodenticulate setae; endopod 5-segmented, ventral margin of the proximal segment denticulate, and 16, 10, 7–8, 7, 7 setae respectively; exopod 2-segmented with 3 distal simple setae on proximal segment and 5 terminal plumose setae on distal segment.
Pereiopods (Fig. 6i–k, j1) Cheliped sparsely setose as shown, with prominent ischial curved hook; pereiopods II–V thin and setose, inner margin of dactyl with 3 stout spines and 2 shorter lateral spines; pereiopod II with a tubercle on ischial segment; dactylus of pereiopod V with 3 long setae (feelers).
Sternum (Fig. 8j) Maxilliped sternites completely fused with 4 simple setae, cheliped sternites with 2 simple setae each, pereiopod sternites 2–5 without setae; sternal sutures are interrupted medially.
Pleon (Fig. 5a) Six pleonites; setation of pleonites II-VI as shown; pleonite VI reduced.
Pleopods (Fig. 6l, m) Present on pleonites II-VI; endopods unsegmented with 3 cincinuli; exopod unsegmented with 16, 14, 14, 14 long plumose natatory setae; uropod 2-segmented, proximal segment with 1 seta, distal segment with 9 terminal plumose natatory setae.
Telson (Fig. 5a) Reduced, with 1 pair of dorsal setae.
Distinctive morphological features for Liomera cinctimanus, Pseudoliomera variolosa, Williamstimpsonia stimpsoni, Scopolius nuttingi and Etisus odhneri are listed in the Table 3 and illustrated in the Figs. 5b–f, 7b–p, 8k–o.
Remarks
Family Portunidae Rafinesque, 1815
The megalopas of Portunidae can be distinguished from those of other brachyurans for the following diagnostic combination of characters: presence of rostral spine projecting almost horizontally, the dactyl of the 5th pereiopods paddle-like, pair of spines projecting posteriorly on 4th sternite, and 5th segment of pleon with lateral spines. Therefore, such features can also be observed in the megalopas of the following genera analyzed in this study: Achelous De Haan, 1833, Arenaeus Dana, 1851, Callinectes Stimpson, 1860; Charybdis De Haan, 1833; Portunus Weber, 1795 and Thalamita Latreille, 1829. Nevertheless, as different authors argue29,30,47,48 the identification of the megalopa stage of portunids at specific level is a difficult task, because to the close similarity of its morphologies that makes all larvae remarkably similar. This task is more complicated when the larvae are from planktonic samples. Next, we highlight the most distinctive morphological features for each studied species.
Achelous floridanus (Figs. 2a, 3a–m, 4a, g, 8a)
Mantelatto et al.38 proposed, and recently corroborated by Mantelatto et al.8, the resurrection of genus Achelous De Haan 1833, a reassignment of nine American species and eleven (of twelve) eastern Pacific species respectively, formerly treated as Portunus Weber, 1795. Achelous now contains a total of 21 American species, between them: Achelous spinicarpus Stimpson, 1871 and Achelous spinimanus (Latreille, 1819), the only ones whose megalopa have been described (Bookhout and Costlow31, as Portunus spinicarpus; and Negreiros-Fransozo et al.49, as Portunus spinimanus, respectively).
The megalopa morphology of A. spinicarpus and A. spinimanus is similar but both are different to A. floridanus. A. spinicarpus and A. spinimanus have 2 carpal spines on cheliped while they are absent in A. floridanus; the pair of spines projecting posteriorly on 4th sternite on the sternum is much longer in A. floridanus, reaching the 3rd pleonite but it is smaller in both remaining species; A. floridanus shows a characteristic rostrum curved upwards with numerous minute setae (Fig. 2a), and the orbital region presents 7 plumose setae (Fig. 4g), both characters are not present in A. spinicarpus and A. spinimanus. Finally, other important character that usually remains constant within the genus, such as the number of antennal segments and setation of endopod of the maxillule, is different between these species: A. floridanus shows an antenna 8-segmented while 7-segmented in A. spinicarpus and A. spinimanus.
Arenaeus mexicanus (Figs. 2b, 4b, h, 8b)
The genus Arenaeus encompasses only two species: A. cribrarius (Lamark, 1818) and A. mexicanus. Stuck and Truesdale50 describe the larval development of A. cribanarius and the diagnosis characters of the genus are summarized in an antennal flagellum 8-segmented, exopod of uropods with 1,12–13 setae, carpal spine and ischial hook on cheliped, coxal spine on 2nd pereiopod and coxal tubercle on 3rd pereiopod, and tubercles on 2nd–3rd sternites.
The megalopa of A. mexicanus can be easy separated from that of A. cribrarius by the setation of antennal flagellum: 0,0,4,2,1 + 5,0,3,3 for A. mexicanus and 0,0,4,2,4,2,4,4 for A. cribrarius.
Portunus hastatus (Figs. 2c, 4c, p, 8c)
The genus Portunus includes about 100 species51 but the species with known megalopas only are P. trituberculatus (Miers, 1876), P. pelagicus (Linnaeus, 1758), and P. gibbesii (Stimpson, 1859) described by Kurata29, Yatsuzuka and Sakai52 and Negreiros-Fransozo et al.49, respectively.
Similar to the Achelous genus, the species included in Portunus show a high variability in certain morphological characters that have been considered key to characterize the genus. Therefore, there is not a set of morphological characters common to all species of the genus Portunus that allow distinguishing its megalopas from those of other genera of portunids. However, there are some characters that can be used to distinguish the megalopas already described, such as: presence of carpal spine in P. hastatus, P. pelagicus and P. gibbesii and absence in P. trituberculatus; ischial hook on cheliped present in P. pelagicus and P. trituberculatus, but absence in P. hastatus and P. gibbesi; and the 8-segmentation of the antennal segment in P. hastatus and P. pelagicus while it is 7-segmented in P. gibbesii.
Charybdis (Charybdis) hellerii (Figs. 2d, 4d, j, 8d).
There are published megalopas descriptions for seven species of Charybdis: C. japonica (A. Milne Edwards, 1861) by Kurata and Nishina53, C. acuta (A. Milne Edwards, 1869) by Kurata and Omi54, C. callianassa (Herbst, 1789) by Greenwood and Fielder47, C. truncata (Fabricius, 1798) by Greenwood and Fielder55, C. feriata (Linnaeus, 1758) by Fielder et al.56, C. bimaculata (Miers, 1886) by Hwang and Kim57 and C. natator (Herbst, 1794) by Islma et al.58.
Dineen et al.59 described the larval development of C. hellerii but no provided detailed morphological description of the megalopa stage, only one photograph. Kurata and Nishina52 described the megalopas of C. acuta and C. japonica, collected in Japan, but they are too brief to allow detailed comparison with other species of Charybdis (see Table 4).
Although, the earlier published larval descriptions lack enough detail, it is noted that these species share the same external characters, such as: antennal flagellum 8-segmented, absent of carpal spine and ischial hook on cheliped (absent in C. feriata), and a coxal spine on 2nd pereiopod (absent in C. feriata).
Thalamita admete (Figs. 2e, 4e, k, 8e)
In the genus Thalamita, descriptions of the megalopas are available for T. danae Stimpson, 1858 by Fielder and Greenwood60, T. crenata Rüppell, 1830 by Krishnan and Kannupandi61, and T. pelsarti Montgomery, 1931 by Islam et al.62.
We compared them and found that all species share the absence of spines on cheliped and pereiopods. These characters combined with an 8-segmented antennal flagellum and setation of uropods (1,11) characterize the genus. Thalamita admete can be differentiated from the three species by antennal setation (Table 4).
Callinectes spp. (Figs. 2f–I, 4f, l–o, q, r, 8f–i)
The megalopa stage known of Callinectes are: C. sapidus Rathbun, 1896 by Costlow and Bookhout63, and C. similis Williams, 1966 by Bookhout and Costlow64. The authors differenced the megalopa stage of these two species by size and examination of minute characteristics, but larval descriptions lack enough detail.
In this study megalopas of four species of the Callinectes are described: C. amnicola, C. arcuatus, C. ornatus, and C. toxotes as shown in Fig. 2f–i. The megalopas of Callinectes genus are strongly similar, being difficult to differentiate them only based on external characters. These six species share the most important external characters like the antennal flagellum 8-segmented, carpal spine on cheliped absent, ischial hook on cheliped present and coxal spine on 2nd pereiopod absent. Even the carapace with tubercle on protogastric region with a row of 8–10 minute setae is the same for the four Callinectes species of this study.
The four species can be differentiated only by a thorough examination of the mouthparts (see Tables 2 and 4).
Family Xanthidae MacLeay, 1838
We have identified the megalopa stage of 5 species of xanthids. Xanthidae is a large and heterogeneous family containing about 124 genera and around 640 species47,58. The intergeneric variation in xanthids megalopas appears to be too significant to find constants group characters, as in adults40,65.
In this paper is presented for the first time morphological features of the megalopa stage for the genera Neoliomera Odhner, 1925, Liomera Dana, 1851 (Figs. 5b, 7b, g, 8k), Pseudoliomera Odhner, 1925 (Figs. 5c, 7c, h, o, n, 8l), and Williamstimpsonia Števčić, 2011 (Figs. 5d, 7d, i, l, m, 8m) (Table 3). The lack of previous descriptions of the megalopas of other species belonging to these genera does not allow a review of morphological characteristics.
Etisus odhneri (Figs. 5f, 7f, k, m, 8o)
In this genus, the only larval development known is for Etisus laevimanus Randall, 1840 by Suzuki66. We compared them and found several minute differences, all related with the number of setae in antennule, antenna, and uropods. Antennule accessory flagellum in E. odhneri presents 1 + 1 + 3 setae and in E. laevimanus 1 + 3 setae; antennal flagellum setation in E. odhneri is 0,0,3,0,3,0,4,4 and 0,0,2–3,0,4,0,4,4 in E. laevimanus; and uropod in E. odhneri presents 0,10 setae and 1, 10–11 in E. laevimanus. Both species show ischial spine on cheliped.
Family Pseudorhombilidae Alcock, 1900
Pseudorhombilidae includes 19 genera and 50 species67, but larval data are only known for 3 species. In the present study only one megalopa of the monospecific genus Scopolius have been identified, S. nuttingi (Figs. 5e, 7e, j, 8n), that lack of previous larval descriptions.
Discussion
DNA barcoding is a useful tool for the identification of crustaceans by assigning indeterminate specimens to known species11,68,69 and faster method for the descriptions of brachyuran larvae13. As more research uses these genetic markers begin to be addressed questions relating to the biodiversity, ecology, and evolution of natural systems70,71.
But, although this molecular technique is widely used and popular, identifying unknown specimens through DNA barcodes requires a reference library containing morphologically-identified barcoded specimens against which unknowns can be compared72, highlighting the reliability of the database with adequate validation and detection of erroneous sequences73,74. While the use of DNA barcode databases for the identification of many marine species is an increasingly used technique in taxonomy, there are still large numbers of unexplored taxa, with little or no DNA barcode coverage or, sometimes, several species lack of sequence in the database is correctly assigned to higher taxa75,76. Thus, the results provide useful information to estimate species numbers, regardless of their formal taxonomic state, distribution and ecology as well as a framework for future taxonomic work77,78.
The availability of this information, especially for the family Portunidae, is of great importance not only to understand the life history of these species that are of commercial and therefore economic interest, but also because they have invasive potential. Species belonging to this family has a high dispersal potential because the adults are swimmers, have long larval periods and all stages of the life cycle can actively migrate long distances and, therefore, are dispersal agents both within the region of origin and to new environments79, which can turn them into invasive species, as is the case of the blue crab, Callinectes sapidus, in the Mediterranean.
In the present work, Charybdis hellerii is the only one portunid that has been reported as an invasive crab80 where it was collected, and this species continues to expand its range81 in Caribbean Sea. However, although all the other Portunidae megalopas were collected within their known range, it is interesting to note that they were collected several miles from the coast, in the open sea, which highlights the potential to expand their range of distribution.This work focused mainly on the taxonomic applications of DNA barcoding to increase the knowledge of unknown brachyuran megalopa stages. This study provides a valuable larval morphology information about 9 portunids, 5 xanthids and 1 pseudorhombilid that will support future systematic, ecological, and biological studies about these families.
Methods
Fieldwork
Megalopas were collected in two research cruises under the support of the MALASPINA 2010–2011 and MAF 2015 research projects (Table 5). The MALASPINA Circumnavigation Expedition was carried out with the general objectives of assessing the impact of global change on the oceans and exploring its biodiversity. The research cruise was conducted between December 2010 and July 2011, involved two oceanographic research vessels, the Hespérides and the Sarmiento de Gamboa, which covering a total of 42,000 nautical miles through the tropical and subtropical regions of the Atlantic, Indian, and Pacific oceans, sampling in a total of 147 stations. MAF research cruise was held to assess the carbon vertical active flux in the open sea due to zooplankton and micronekton and main responsible species. A total of 13 stations were sampled between 3rd and 29th April 2015 on board of the RV Hespérides, which crossed the tropical and subtropical Atlantic regions from Salvador de Bahia, Brazil, to Las Palmas, Canary Islands, Spain. (Fig. 1).
Sample processing
In both research expeditions, megalopas were collected from the superficial layer with a neuston net with a mesh size of 200 microns, hauled from 10 to 15 min at 2–3 knots. Samples were immediately fixed and preserved in 95% ethanol. All brachyuran megalopa stages were counted and sorted from the zooplankton samples using a stereomicroscope. Prior to DNA extraction, all larvae were examined morphologically and sorted into morphotypes according to the external characters.
Megalopas morphological descriptions
For easier observation of larvae structures and setation under microscope, megalopas were first placed for 5–10 min in a watch glass with 2 ml of warm lactic acid before proceeding with the dissection of the body parts82.
Drawings and measurements of megalopa stage were made using a Leica MZ6 and microscope Nikkon Eclipse 90i with integrated camera lucida. All measurements were made using an ocular micrometer. Descriptions were based on all the collected megalopas of each species identified by DNA barcoding (see Table 1). The following measurements were taken for the megalopa: cephalothorax length (CL), measured from the rostrum (tip of rostrum in portunids) to posterior margin of cephalothorax; and cephalothorax width (CW), measured as the cephalothorax maximum width (mesobranchial regions).
For the megalopas dorsal view, only the left pereiopods were drawn since one of the right pereiopods was used for molecular analyzes.
Descriptions were arranged according to the standards proposed by Clark et al.83 and Clark & Cuesta84, and setal terminology follows the classification by Landeira et al.85. A detailed description of Achelous floridanus is provided while the others portunids descriptions are summarized in the Table 2. For Xanthidae and Pseudorhombilidae families, the specie of Neoliomera cerasinus is described in detail and the others xanthoids descriptions are summarized in the Table 3.
Molecular analysis
The identification of the megalopas was based on partial sequences of the 16S rRNA and COI mitochondrial genes. Total genomic DNA of the megalopas from MALASPINA Expedition was extracted from muscle tissue from one pereiopod and incubated for 1–24 h in 300 µl lysis buffer (5 ml of 1 M Tris–HCl (pH 8), 1 ml 0.5 M EDTA, and 5 ml of 10% SDS solution to 400 ml of distilled water) at 65º C. Protein was precipitated by addition of 100 µl of 7.5 M ammonium acetate and subsequent centrifugation, and DNA precipitation was obtained by addition of 300 µl of isopropanol and posterior centrifugation. The resulting pellet was washed with ethanol (70%), dried, and finally resuspended in Milli-Q distilled water82. In the megalopas from MAF Expedition, total genomic DNA was also extracted from muscle tissue from one pereiopod, but the extraction process followed a modified Chelex 10% protocol by Estoup et al.86. Target mitochondrial DNA from the 16S rRNA and COI genes was amplified with the primers and the cycling conditions of the polymerase chain reaction (PCR) listed in Table 6. PCR products were sent to New Biotechnic, CISA-INIA, and Stab Vida companies to be purified and then bidirectionally sequenced.
Sequences were edited using the software Chromas version 2.0. With the obtained final DNA sequences were performed a BLAST (Basic Local Alignment Search Tool) on NCBI (National Center for Biotechnology Information) web facility on GenBank sequences database (http://www.ncbi.nlm.nih.gov/genbank/) to get the best matches for identification. The COI sequences were also searched in the official Barcode of Life database (BOLD) (http://v3.boldsystems.org/index.php/IDS_OpenIdEngine). Identifications were considered as positive when retrieved sequences showed similarity values greater than 99%, only differed in 1–3 or 1–7 mutations in 16S or COI, respectively, a more conservative limit than other previous works identifying decapod larvae considering > 98%96. Larval sequences for both genes are deposited in Genbank (see Table 1).
Ethical approval
This article does not contain any studies with human participants performed by any of the authors. All applicable international, national, and institutional guidelines for the care and use of animals were followed.
References
Nigro, L. M., Angel, M. V., Blachowiak-Samolyk, K., Hopcroft, R. R. & Bucklin, A. Identification, discrimination, and discovery of species of marine planktonic ostracods using DNA barcodes. PLoS ONE 11(1), e0146327. https://doi.org/10.1371/journal.pone.0146327 (2016).
Wildish, D. J., Smith, S. R., Loeza-Quintana, T., Radulovici, A. E. & Adamowicz, S. J. Diversity and dispersal history of the talitrids (Crustacea: Amphipoda: Talitridae) of Bermuda. J. Nat. Hist. 50(29–30), 1911–1933. https://doi.org/10.1080/00222933.2016.1180719 (2016).
Tanabe, A. S. & Toju, H. Two new computational methods for universal DNA barcoding: A benchmark using barcode sequences of bacteria, archaea, animals, fungi, and land plants. PLoS ONE 8(10), e76910. https://doi.org/10.1371/journal.pone.0076910 (2013).
Thomsen, P. F. & Willerslev, E. Environmental DNA: An emerging tool in conservation for monitoring past and present biodiversity. Biol. Conserve. 183, 4–18. https://doi.org/10.1016/j.biocon.2014.11.019 (2015).
Weis, A. et al. Pallenopsis patagonica (Hoek, 1881): A species complex revealed by morphology and DNA barcoding, with description of a new species of Pallenopsis Wilson, 1881. Zool. J. Linn. Soc. 170(1), 110–131. https://doi.org/10.1111/zoj12097 (2014).
Raupach, M. J. et al. The application of DNA barcodes for the identification of marine crustaceans from the North Sea and adjacent regions. PLoS ONE 10(9), e0139421. https://doi.org/10.1371/journal.pone.0139421 (2015).
Palero, F., Genis-Armero, R., Hall, M. R. & Clark, P. F. DNA barcoding the phyllosoma of Scyllarides squammosus (H. Milne Edwards, 1837) (Decapoda: Achelata: Scyllaridae). Zootaxa 4139(4), 481. https://doi.org/10.11646/zootaxa.4139.4.2 (2016).
Mantelatto, F. L., Robles, R., Wehrtmann, I. S., Schubart, C. D. & Felder, D. L. New insights into the molecular phylogeny of the swimming crabs of the genera Portunus Weber, 1795 and Achelous De Haan, 1833 (Brachyura: Portunidae) of the Americas. J. Crustac Biol. 38(2), 190–197 (2018).
Raupach, M. J. & Radulovici, A. E. Looking back on a decade of barcoding crustaceans. ZooKeys 539, 53. https://doi.org/10.3897/zookeys.539.6530 (2015).
Ahyong, S. T. et al. Subphylum Crustacea Brünnich, 1772. In: Zhang Z-Q (Ed.) Animal biodiversity: an outline of higher-level classification and survey of taxonomic richness. Zootaxa 3148, 165–191 (2011).
Radulovici, A. E., Sainte-marie, B. & Dufresne, F. DNA barcoding of marine crustaceans from the Estuary and Gulf of St Lawrence: A regional-scale approach. Mol. Ecol. Resour. 9(s1), 181–187. https://doi.org/10.1111/j.1755-0998.2009.02643.x (2009).
Marco-Herrero, E., González-Gordillo, J. I. & Cuesta, J. A. Larval morphology of the family Parthenopidae, with the description of the megalopa stage of Derilambrus angulifrons (Latreille, 1825) (Decapoda: Brachyura), identified by DNA barcode. J. Mar. Biol. Ass. U. K. 95(03), 513–521. https://doi.org/10.1017/S0025315414001908 (2015).
Di Muzio, G. et al. Morphology of planktonic zoeal stages of Palicus caronii (Decapoda, Brachyura), identified by DNA barcoding, provides novelties to Palicoidea larval systematics. Sci. Rep. 9(1), 1–10. https://doi.org/10.1038/s41598-019-55412-3 (2019).
Villarino, E. et al. Large-scale ocean connectivity and planktonic body size. Nat. Commun. 9(1), 1–13. https://doi.org/10.1038/s41467-017-02535-8 (2018).
Ershova, E. et al. Diversity and distribution of meroplanktonic larvae in the Pacific Arctic and connectivity with adult benthic invertebrate communities. Front. Mar. Sci. 6, 490. https://doi.org/10.3389/fmars.2019.00490 (2019).
Rice, A. L. The megalopa stage in brachyuran crabs. The Podotremata Guinot. J. Nat. Hist. 15(6), 1003–1011. https://doi.org/10.1080/00222938100770751 (1981).
Anger, K. Contributions of larval biology to crustacean research: a review. Invertebr. Reprod. Dev. 49(3), 175–205. https://doi.org/10.1080/07924259.2006.9652207 (2006).
Anger, K. The Biology of Decapod Crustacean Larvae Vol. 14 (AA Balkema Publishers, 2001). https://doi.org/10.0013/epic.15410.d001.
Marco-Herrero, E. Aplicación de técnicas morfológicas y moleculares en la identificación de la megalopa de decápodos braquiuros de la península ibérica. PhD Dissertation, Spain. http://hdl.handle.net/10550/47114 (2015).
Webley, J. A., Connolly, R. M. & Young, R. A. Habitat selectivity of megalopas and juvenile mud crabs (Scylla serrata): Implications for recruitment mechanism. Mar. Biol. 156(5), 891–899. https://doi.org/10.1007/s00227-009-1134-0 (2009).
Little, K. T. & Epifanio, C. E. Mechanism for there-invasion of an estuary by two species of brachyuran megalopas. Mar. Ecol. Prog. Ser. 68, 35–242 (1991).
Olaguer-Feliú, A. O., Flores, A. A. V., Queiroga, H. & González-Gordillo, J. I. Shelf and estuarine transport mechanisms affecting the supply of competent larvae in a suite of brachyuran crabs with different life histories. Mar. Ecol. Prog. Ser. 410, 125–141. https://doi.org/10.3354/meps08604 (2010).
Martin, J. W. Brachyura. In Martin, J. W., Olesen, J. & Høeg J. T. (Eds.) Atlas of Crustacean Larvae 295–310 (Johns Hopkins University Press, 2014) https://doi.org/10.1353/book.31448
Pardo, L. M., Ampuero, D. & Véliz, D. Using morphological and molecular tools to identify megalopa larvae collected in the field: the case of sympatric Cancer crabs. J. Mar. Biol. Ass. U. K. 89, 481–490. https://doi.org/10.1017/S0025315409003233 (2009).
Marco-Herrero, E. et al. The systematic position of Ergasticus (Decapoda, Brachyura) and allied genera, a molecular and morphological approach. Zool. Scr. 42, 427–439. https://doi.org/10.1111/zsc.12012 (2013).
Brandão, M. C., Freire, A. S. & Burton, R. S. Estimating diversity of crabs (Decapoda: Brachyura) in a no-take marine protected area of the SW Atlantic coast through DNA barcoding of larvae. Syst. Biodiver. 14(3), 288–302. https://doi.org/10.1080/14772000.2016.1140245 (2016).
Bezeng, B. S. & van der Bank, H. F. DNA barcoding of southern African crustaceans reveals a mix of invasive species and potential cryptic diversity. PLoS ONE 14(9), e0222047. https://doi.org/10.1371/journal.pone.0222047 (2019).
Spiridonov, V. A., Neretina, T. V. & Schepetov, D. Morphological characterization and molecular phylogeny of Portunoidea Rafinesque, 1815 (Crustacea Brachyura): Implications for understanding evolution of swimming capacity and revision of the family-level classification. Zool. Anz. J. Comp. Zool. 253(5), 404–429. https://doi.org/10.1016/j.jcz.2014.03.003 (2014).
Kurata, H. Larvae of decapoda brachyura of Arasaki, Sagami Bay [Japan], 5: The swimming crabs of subfamily Portuninae. Bull. Nansei Regional Fisheries Res. Lab. 8, 39–65 (1975).
Rice, A. L. & Ingle, R. W. The larval development of Carcinus maenas (L.) and C. mediterraneus Czerniavsky (Crustacea, Brachyura, Portunidae) reared in the laboratory. Bull. Br. Mus. (Nat. Hist.) 28, 103–119 (1975).
Bookhout, C. G. & Costlow, J. D. Larval development of Portunus spinicarpus reared in the laboratory. Bull. Mar. Sci. 24, 20–51 (1974).
Clark, P. F. A comparative study of zoeal morphology in the genus Liocarcinus (Crustacea: Brachyura: Portunidae). Zool. J. Linn. Soc. 82(3), 273–290. https://doi.org/10.1111/j.1096-3642.1984.tb00644.x (1984).
Davie P. J. F. Crustacea: Malacostraca: Eucarida (Part 2): Decapoda – Anomura, Brachyura. In: Wells A, Houston WWK, Eds. Zoological Catalogue of Australia. Volume 19.3B. (CSIRO Publishing, 2002).
Thoma, B. P., Guinot, D. & Felder, D. L. Evolutionary relationships among American mud crabs (Crustacea: Decapoda: Brachyura: Xanthoidea) inferred from nuclear and mitochondrial markers, with comments on adult morphology. Zool. J. Linn. Soc. 170(1), 86–109. https://doi.org/10.1111/zoj12093 (2014).
Scotto, L. E. Larval development of the cuban stone crab, Menippe nodifrons (Brachyura, Xanthidae). Fish. Bull. 77(2), 359 (1979).
Negreidos-Fransozo, M. L. Desenvolvimento pós-embrionário de Eurynome abbreviatus (Sitmpson, 1860) (Crustacea, Decapoda, Xanthidae), em laboratório. Bolm. Zool. S. Paulo 10, 19–39 (1986).
Rice, A. L. The metamorphosis of a species of Homola (Crustacea, Decapoda: Dromiacea). Bull. Mar. Sci. 14(2), 221–238 (1964).
Mantelatto, F. L., Robles, R., Schubart, C. D. & Felder, D. L. Molecular phylogeny of the genus Cronius Stimpson 1860, with reassignment of C. tumidulus and several American species of Portunus to the genus Achelous de Haan, 1833 (Brachyura: Portunidae). Crustacean issues: decapod crustacean phylogenetics, 567–579 (2009).
Evans, N. Molecular phylogenetics of swimming crabs (Portunoidea Rafinesque, 1815) supports a revised family-level classification and suggests a single derived origin of symbiotic taxa. PeerJ 6, e4260. https://doi.org/10.7717/peerj.4260 (2018).
Lai, J. C., Mendoza, J. C. E., Guinot, D., Clark, P. F. & Ng, P. K. Xanthidae MacLeay, 1838 (Decapoda: Brachyura: Xanthoidea) systematics: A multi-gene approach with support from adult and zoeal morphology. Zool. Anz A J. Comp. Zool. 250(4), 407–448. https://doi.org/10.1016/j.jcz.2011.07.002 (2011).
Wetzer, R., Martin, J. W. & Trautwein, S. E. Phylogenetic relationships within the coral crab genus Carpilius (Brachyura, Xanthoidea, Carpiliidae) and of the Carpiliidae to other xanthoid crab families based on molecular sequence data. Mol. Phylog. Evol. 27(3), 410–421. https://doi.org/10.1016/S1055-7903(03)00021-6 (2003).
Faria, S. C. et al. Macroevolution of thermal tolerance in intertidal crabs from Neotropical provinces: A phylogenetic comparative evaluation of critical limits. Ecol. Evol. 7(9), 3167–3176. https://doi.org/10.1002/ece3.2741 (2017).
Felder, D. L. & Thoma, B. P. Description of Etisus guinotae n. sp., and discussion of its recent discovery in the Gulf of Mexico (Brachyura, Decapoda, Xanthidae). In Studies on Brachyura: An Homage to Danièle Guinot 117–138 Brill. (2010).
Hui, T. H., Naruse, T., Fujita, Y. & Kiat, T. S. Observations on the fauna from submarine and associated anchialine caves in Christmas Island, Indian Ocean Territory, Australia. Raffles Bull. Zool. 30, 406–418 (2014).
Ng, P. K. On a new species of cavernicolous Neoliomera (Crustacea: Decapoda: Brachyura: Xanthidae) from Christmas Island and Ryukyus, Japan. Raffles Bull. Zool. 50(1), 95–100 (2002).
Fujita, Y., Naruse, T. & Yamada, Y. Two submarine cavernicolous crabs, Atoportunus gustavi Ng & Takeda, 2003, and Neoliomera cerasinus Ng, 2002 (Crustacea: Decapoda: Brachyura: Portunidae and Xanthidae), from Shimojijima Island, Miyako Group, Ryukyu Islands, Japan. Fauna Ryukyuana 1, 1–9 (2013).
Lebour, M. V. The larval stages of the Plymouth Brachyura. Proc. Zool. Soc. Lond. 98(2), 47–560. https://doi.org/10.1111/j.1469-7998.1928.tb07158.x (1928).
Greenwood, J. G. & Fielder, D. R. The zoeal stages and megalopa of Charybdis callianassa (Herbst) (Decapoda: Portunidae), reared in the laboratory. Proc. R. Soc. Queensl. 91, 61–76 (1980).
Negreidos-Fransozo, M. L., Meyers, N., Fransozo, V. & Thorton-de Victor, S. The megalopa stage of Portunus spinimanus Latreille, 1819 and Portunus gibbesii (Stimpson, 1859) (Decapoda, Brachyura, Portunidae) from the southeastern Atlantic coast of the United States. Zootaxa 1638, 21–37 (2007).
Stuck, K. C. & Truesdale, F. M. Larval development of the speckled swimming crab, Arenaeus cribrarius (Decapoda: Brachyura: Portunidae) reared in the laboratory. Bull. Mar. Sci. 42(1), 101–132 (1988).
Ng, P. K., Guinot, D. & Davie, P. J. Systema Brachyurorum: Part I. An annotated checklist of extant brachyuran crabs of the world. Raffles Bull. Zool. 17(1), 1–286 (2008).
Yatsuzuka, K. & Sakai, K. The larvae and juvenile crabs of Japanese Portunidae (Crustacea, Brachyura), 1: Portunus (Portunus) pelagicus (Linne). Rep. Usa Mar. Biol. Institute-Kochi University 2, 25–41 (1980).
Kurata, H. & Nishina, S. The zoeal stages of the swimming crabs, Charybdis japonica and Portunus hastatoides reared in the laboratory. Bull. Nansei Regional Fisheries Res. Lab. 8, 21–27 (1975).
Kurata, H. & Omi, H. The larval stages of a swimming crab, Charybdis acuta. Bull. Tokai Regional Fisheries Res. Lab. 57, 129–136 (1969).
Greenwood, J. G. & Fielder, D. R. Description of the later zoeal stages and megalopa of Charybdis truncata (Fabricius, 1798) (Crustacea: Portunidae). J. Nat. His. 17(1), 15–30. https://doi.org/10.1080/00222938300770021 (1983).
Fielder, D. R., Greenwood, J. G. & Campbell, G. The megalopa of Charybdis feriata (Linnaeus) with additions to the zoeal larvae descriptions (Decapoda, Portunidae). Crustaceana 46(2), 160–165. https://doi.org/10.1163/156854084X00667 (1984).
Hwang, S. G. & Kim, C. H. Complete larval development of the swimming crab, Charybdis bimaculata (Miers, 1886) (Crustacea, Brachyura, Portunidae), reared in the laboratory. Korean J. Zool. 38, 465–482 (1995).
Islam, M. S., Shokita, S. & Higa, T. Larval development of the swimming crab Charybdis natator (Crustacea: Brachyura: Portunidae) reared in the laboratory. Species Divers 5(4), 329–349. https://doi.org/10.12782/specdiv.5.329 (2000).
Dineen, J. F., Clark, P. F., Hines, A. H., Reed, S. A. & Walton, H. P. Life history, larval description, and natural history of Charybdis hellerii (Decapoda, Brachyura, Portunidae), an invasive crab in the western Atlantic. J. Crustac. Biol. 21(3), 774–805. https://doi.org/10.1163/20021975-99990173 (2001).
Fielder, D. R. & Greenwood, J. G. Larval development of the swimming crab Thalamita danae Stimpson, 1858 (Decapoda, Portunidae), reared in the laboratory. Proc. R. Soc. Qld. 90, 13–20 (1979).
Krishnan, T. & Kannupandi, T. Laboratory cultured zoeae, megalopa and first crab of the estuarine crab Thalamita crenata (Latr.) A. Milne Edwards 1861 (Brachyura: Portunidae). Mahasagar 23(2), 139–152 (1990).
Islam, M. S., Machiko, K. & Shokita, S. Larval development of the swimming crab Thalamita pelsarti Montgomery, 1931 (Crustacea: Brachyura: Portunidae) reared in the laboratory. Russian J. Mar. Biol. 31(2), 78–90. https://doi.org/10.1007/s11179-005-0048-z (2005).
Costlow, J. D. & Bookhout, C. G. The larval development of Callinectes sapidus Rathbun reared in the laboratory. Biol. Bull. 116(3), 373–396. https://doi.org/10.2307/1538947 (1959).
Bookhout, C. G. & Costlow, J. D. Larval development of Callinectes similis reared in the laboratory. Bull. Mar. Sci. 27(4), 704–728 (1977).
Martin, J. W. Phylogenetic significance of the brachyuran megalopa: evidence from the Xanthidae. Symp. Zool. Soc. Lon. 59, 69–102 (1988).
Suzuki, H. The larval development of Etisus laevimanus Randall (Crustacea, Brachyura, Xanthidae). La Mer Bull. Soc. Franco-Japonaise Oceanogr. 16, 176–187 (1978).
Davie, P.J.F., Guinot, D. & Ng, P.K.L. Systematics and classification of Brachyura. In Treatise on Zoology-Anatomy, Taxonomy, Biology. The Crustacea, Volume 9 Part C (2 vols) 1049–1130 (Brill, 2015) https://doi.org/10.1163/9789004190832 (2015).
Bucklin, A., Steinke, D. & Blanco-Bercial, L. DNA barcoding of marine metazoa. Ann. Rev. Mar. Sci. 3, 471–508. https://doi.org/10.1146/annurev-marine-120308-080950 (2011).
Marco-Herrero, E., Drake, P. & Cuesta, J. A. Larval morphology and DNA barcodes as valuable tools in early detection of marine invaders: a new pea crab found in European waters. J. Mar. Biol. Ass. U. K. 98(7), 1675–1683. https://doi.org/10.1017/S0025315417000996 (2018).
Kress, W. J., García-Robledo, C., Uriarte, M. & Erickson, D. L. DNA barcodes for ecology, evolution, and conservation. Trends Ecol. Evol. 30(1), 25–35. https://doi.org/10.1016/j.tree.2014.10.008 (2015).
Da Silva, J. M. & Willows-Munro, S. A review of over a decade of DNA barcoding in South Africa: A faunal perspective. Afr. Zool. 51(1), 1–12. https://doi.org/10.1080/15627020.2016.1151377 (2016).
Collins, R. A. et al. Known knowns, known unknowns, unknown unknowns and unknown knowns in DNA barcoding: a comment on Dowton et al.. Syst. Biol. 63(6), 1005–1009. https://doi.org/10.1093/sysbio/syu060 (2014).
Cognato, A. I. et al. The essential role of taxonomic expertise in the creation of DNA databases for the identification and delimitation of Southeast Asian ambrosia beetle species (Curculionidae: Scolytinae: Xyleborini). Front. Ecol. Evol. 8, 27. https://doi.org/10.3389/fevo.2020.00027 (2020).
Salvi, D., Berrilli, E., D’Alessandro, P. & Biondi, M. Sharpening the DNA barcoding tool through a posteriori taxonomic validation: The case of Longitarsus flea beetles (Coleoptera: Chrysomelidae). PLoS ONE 15(5), e0233573. https://doi.org/10.1371/journal.pone.0233573 (2020).
Blanco-Bercial, L., Cornils, A., Copley, N. & Bucklin, A. DNA barcoding of marine copepods: assessment of analytical approaches to species identification. PLoS Curr. 6(1), 1–21. https://doi.org/10.1371/currents.tol.cdf8b74881f87e3b01d56b43791626d2 (2014).
Wilson, J. J. et al. When species matches are unavailable are DNA barcodes correctly assigned to higher taxa? An assessment using sphingid moths. BMC Ecol. 11(1), 1. https://doi.org/10.1186/1472-6785-11-18 (2011).
Mantelatto, F. L. et al. DNA sequence database as a tool to identify decapod crustaceans on the São Paulo coastline. Mitochondrial DNA Part A 29(5), 805–815. https://doi.org/10.1080/24701394.2017.1365848 (2018).
Palecanda, S., Feller, K. D. & Porter, M. L. Using larval barcoding to estimate stomatopod species richness at Lizard Island, Australia for conservation monitoring. Sci. Rep. 10(1), 1–11. https://doi.org/10.1038/s41598-020-67696-x (2020).
Anger, K., Queiroga, H., & Calado, R. Larval development and behaviour strategies in Brachyura in Treatise on Zoology-Anatomy, Taxonomy, Biology. The Crustacea 9C (ed. Castro, P.) 317–374, (Brill, 2015) https://doi.org/10.1163/9789004190832_008.
Bolaños, J. A., Baeza, J. A., Hernandez, J. E., Lira, C. & López, R. Population dynamics and reproductive output of the non-indigenous crab Charybdis hellerii in the south-eastern Caribbean Sea. J. Mar. Biol. Assoc. U.K. 92(3), 469–474. https://doi.org/10.1017/S002531541100052X (2012).
Ferry, R., Buske, Y., Poupin, J. & Smith-Ravin, J. First record of the invasive swimming crab Charybdis hellerii (A. Milne Edwards, 1867) (Crustacea, Portunidae) off Martinique, French Lesser Antilles. BioInvasions Rec. https://doi.org/10.3391/bir.2017.6.3.09 (2017).
Marco-Herrero, E., González-Gordillo, J. I. & Cuesta, J. A. Morphology of the megalopa of the mud crab, Rhithropanopeus harrisii (Gould, 1841) (Decapoda, Brachyura, Panopeidae), identified by DNA barcode. Helgol. Mar. Res. 68, 201–208. https://doi.org/10.1007/s10152-014-0381-8 (2014).
Clark, P. F., Calazans, D. K. & Pohle, G. W. Accuracy and standardization of brachyuran larval descriptions. Inver. Reprod. Dev. 33, 127–144. https://doi.org/10.1080/07924259.1998.9652627 (1998).
Clark, P. F. & Cuesta, J. A. Larval systematics of Brachyura in Treatise on Zoology-Anatomy, Taxonomy, Biology. The Crustacea 9C (ed. Castro, P.), 981–1048. (Brill, 2015) https://doi.org/10.1163/9789004190832_020.
Landeira, J. M., Lozano-Soldevilla, F. & González-Gordillo, J. I. Morphology of first seven larval stages of the striped soldier shrimp, Plesionika edwardsii (Brandt, 1851) (Crustacea, Decapoda, Pandalidae) from laboratory reared material. Zootaxa 1986, 51–66. https://doi.org/10.11646/zootaxa.1986.1.2 (2009).
Estoup, A. C. R. L., Largiader, C. R., Perrot, E. & Chourrout, D. Rapid one-tube DNA extraction for reliable PCR detection of fish polymorphic markers and transgenes. Mol. Mar. Biol. Biotech. 5(4), 295–298 (1996).
Crandall, K. A. & Fitzpatrick, J. F. J. Crayfish molecular systematics: Using a combination of procedures to estimate phylogeny. Syst. Biol. 45, 1–26. https://doi.org/10.1093/sysbio/45.1.1 (1996).
Schubart, C. D., Cuesta, J. A. & Felder, D. L. Glyptograpsidae, a new brachyuran family from Central America: Larval and adult morphology, and a molecular phylogeny of the Grapsoidea. J. Crustac. Biol. 22, 28–44 (2002).
Schubart, C. D. & Huber, M. G. J. Genetic comparisons of German populations of the stone crayfish, Austropotamobius torrentium (Crustacea: Astacidae). Bull. Fr. Pêche Piscic. 380–381, 1019–1028. https://doi.org/10.1051/kmae:2006008 (2006).
Mantelatto, F. L., Robles, R. & Felder, D. L. Molecular phylogeny of the western Atlantic species of the genus Portunus (Crustacea, Brachyura, Portunidae). Zool. J. Linn. Soc. 150, 211–220. https://doi.org/10.1111/j.1096-3642.2007.00298.x (2007).
Windsor, A. M., Moore, M. K., Warner, K. A., Stadig, S. R. & Deeds, J. R. Evaluation of variation within the barcode region of Cytochrome c oxydase I (COI) for the detection of commercial Callinectes sapidus Rathbun, 1896 (blue crab) products of non-US origen. PeerJ 7, e7827. https://doi.org/10.7717/peerj.7827 (2019).
Robles, R. et al. Molecular phylogeny of the American Callinectes Stimpson, 1860 (Brachyura: Portunidae), based on two partial mitochondrial genes. Mar. Biol. 150(6), 1265–1274. https://doi.org/10.1007/s00227-006-0437-7 (2007).
Negri, M., Schubart, C. D. & Mantelatto, F. L. Tracing the introduction history of the invasive swimming crab Charybdis hellerii (A Milne-Edwards, 1867) in the Western Atlantic: Evidences of high genetic diversity and multiple introductions. Biol. Invasions 20(7), 1771–1798. https://doi.org/10.1093/icb/42.5.941 (2018).
Schubart, C. D. & Reuschel, S. Proposal for a new classification of Portunoidea and Cancroidea (Brachyura: Heterotremata) based on two independent molecular phylogenies. Crustacean issues: decapod crustacean phylogenetics, 533–549 (2009).
Mantelatto, F. L. et al. DNA sequence database as a tool to identify decapod crustaceans on the São Paulo coastline. Mitochondrial DNA 29(5), 805–815. https://doi.org/10.1080/24701394.2017.1365848 (2018).
Brandão, M. C., Freire, A. S. & Burton, R. S. Estimating diversity of crabs (Decapoda: Brachyura) in a no-take marine protected area of the SW Atlantic coast through DNA barcoding of larvae. Syst. Biodivers. 14(3), 288–302. https://doi.org/10.1080/14772000.2016.1140245 (2016).
Acknowledgements
The Ministry of Economy and Competitiveness of Spain (MINECO) and the European Fund for Economic and Regional Development (FEDER) through the MALASPINA 2010 Expedition (Consolider-Ingenio 2010, CSD2008-00077) and MAF Research Project (National Research Program, CTM2012-39587-C04-02) funded this study. We thank the captains and crews of RV Hespérides in 2010-11 and 2015 for the support during the cruises, and Carlos Sánchez (ICMAN-CSIC) by laboratory work. Thanks are also due to the editor Claudio Oliveira, and to Dimitry Schepetov and two anonymous reviewers for their comments, critiques and corrections that clearly improved the manuscript.
Author information
Authors and Affiliations
Contributions
J.I.G.G. and J.A.C designed the study and obtained the funds, E.M.H. carried out larval descriptions and morphological analysis, J.A.C. performed molecular analysis. E.M.H, J.I.G.G. and J.A.C wrote and reviewed the main manuscript text.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Marco-Herrero, E., Cuesta, J.A. & González-Gordillo, J.I. DNA barcoding allows identification of undescribed crab megalopas from the open sea. Sci Rep 11, 20573 (2021). https://doi.org/10.1038/s41598-021-99486-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-021-99486-4
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.