Abstract
Unravelling the biosynthetic pathway of quinolizidine alkaloids (QAs), regarded as antinutritional compounds of narrow-leafed lupin (NLL) seeds, is fundamental to best exploit NLL as food or feed. We investigated 12 candidate genes connected to QA biosynthesis, selecting them by transcriptomic and genomic approaches, from the landscape of genes differentially expressed in leaves of the high- and low-alkaloid NLL accessions. Linkage analysis enabled the assessment of the location of the candidate genes in relation to iucundus, a major locus of unknown identity, that confers reduced QA content in seeds. The key finding was the identification of APETALA2/ethylene response transcription factor, RAP2-7, cosegregating with the iucundus locus and located within a region with highly significant QTLs that affect QA composition. We additionally identified a 4-hydroxy-tetrahydrodipicolinate synthase (DHDPS) gene involved in L-lysine biosynthesis as being closely linked to iucundus. The distributed location of other remaining candidates (including previously known QA genes) across different linkage groups, also indirectly supports the transcription factor as a possible regulator of lupin alkaloid biosynthesis. Our findings provide crucial insight into QA biosynthesis in NLL. Additionally, we evaluated and selected appropriate reference genes for qRT-PCRs to analyse the expression levels of QA genes in NLL.
Similar content being viewed by others
Introduction
Quinolizidine alkaloids (QAs) are lysine-derived secondary metabolites that are distributed mainly in family Leguminosae and predominantly in genus Lupinus1,2,3. QAs protect plants against herbivores and pathogens, and because of their bitter taste and toxic effects on human and animals they are considered as antinutritional factors of food and feed1.
Previous studies showed that QAs were synthesized in aerial parts of plants and accumulated in maturing seeds with the highest synthesis levels at the flowering and fruit formation stages1,3,4,5. Results are still divided on whether QAs are exclusively transported or partially synthesised in situ in seeds3. The first step in QA biosynthesis is the decarboxylation of lysine by lysine decarboxylase (LDC)6 to give cadaverine amine. Cadaverine is then oxidized by amine oxidase (LaCAO)7, leading to a spontaneous intramolecular Schiff base formation, thus causing ring closure4,5,8,9. The addition of different functional groups and modifications gives the end products of QA synthesis, and their transport/storage forms4,6,7,8,10,11,12. QA ester formation is catalysed by an acyltransferase tigloyl-CoA:(−)-13α-hydroxymultiflorine/(+)-13α-hydroxylupanine O-tigloyltransferase (HMT/HLT; EC 2.3.1.93)10,11. An acyltransferase-like gene (LaAT) was also reported to be involved in the formation of QA esters or N-acylated polyamine conjugates2. To date, only a few studies have aimed to identify genes and characterize enzymes involved in QA biosynthesis in lupins2,6,7,10,11,13,14,15 and the regulatory mechanism is still to be explored.
Due to its great sampling depth, transcriptome sequencing (RNA-Seq) provides a nearly complete picture of transcriptional events, including rare transcripts, at a certain biological time point. Therefore, it has been widely used to characterize and annotate the transcriptomes of nonmodel plants, where the full gene repertoire is not known16,17. RNA-Seq technology has also been commonly employed in the comparative transcriptomic studies that elucidate the response of the plant to different treatment/conditions, and thus allowing the identification of candidate genes underlying particular traits as well as gene expression studies16,18. Lately, RNA-Seq has been used to identify candidate genes involved in various metabolic pathways and in the stress response in legumes (e.g., Rezaei, et al.19, Li, et al.20) and the biosynthesis of different alkaloid classes has been elucidated based on RNA-Seq data (e.g., Rai, et al.21, Cardenas, et al.22).
Recently, lupins have gained much attention as a valuable source of proteins (up to 44%), lipids (up to 14%), and fibre (up to 40% of seed mass), and could be an alternative to soybean in animal feed provided the antinutritional compounds in their seeds are reduced23,24. Consequently, many current lupin breeding programs aim to produce cultivars with strictly reduced alkaloid levels in their seeds. The accepted industry threshold for alkaloid levels is currently 0.02% of the seed dry weight8. In the Polish Lupinus Gene Bank (Poznan Plant Breeders Ltd., Poland), we previously found accessions of narrow-leafed lupin (Lupinus angustifolius L.) and white lupin (Lupinus albus L.) with much lower alkaloid content in their seeds (0.0005% and 0.016% of the seed dry weight, respectively)24,25. This demonstrates the improvement of breeding, and shows the high potential of collection as the source of low alkaloid material.
The major alkaloids (>1% of total alkaloids) detected in narrow-leafed lupin (NLL) are lupanine, 13-hydroxylupanine, angustifoline, and isolupanine1,25. Several single recessive genes related to reduced alkaloid content have been discovered in the seeds of plants with low-alkaloid NLL phenotypes26,27. Among them, the iucundus allele has been the most widely used in NLL breeding, although its identity is unknown28. A mapping population of low-alkaloid (sweet) 83A:476 and high-alkaloid (bitter) P27255 accessions (83 A:476 × P27255) was used to build high-density genetic maps that have provided markers tagging key agronomic traits and a background for synteny analysis with model legumes. The iucundus locus has been integrated into the first and all succeeding versions of the molecular genetic map29,30,31,32,33,34. This has provided an opportunity for the identification of markers linked to the alkaloid-content-related iucundus locus that may be suitable for marker-assisted selection (MAS) with the first attempt reported by Li, et al.35. Recently, a draft L. angustifolius cv. Tanjil transcriptome32 and a comprehensive draft NLL genome sequence34 have been reported (hereafter referred to as Tanjil transcriptome and draft NLL genome, respectively). Furthermore, genetic and genomic analyses of NLL were integrated using molecular cytogenetic tools, which provided insights into the cytomolecular organization and evolution of the NLL genome36,37,38,39,40.
The QA biosynthesis pathway is still obscure in lupins compared with other plants that produce alkaloids (e.g. steroidal glycoalakaloids in Solanaceae species22 and indole alkaloids in Catharanthus roseus L.41). Elucidation of the QA biosynthesis is fundamental to facilitate breeding efforts in providing valuable lupin sources that are used for animal feeding and human consumption. In this study, we sought to identify genes involved in QA biosynthesis by comparative transcriptome analysis of leaf tissue, derived from NLL accessions with contrasting seed alkaloid content. The availability of high resolution NLL linkage maps provided a valuable platform to identify the genetic positions of QA candidate genes as well as to detect quantitative trait loci (QTLs) associated with total QA content and relative abundance of individual QAs in seeds. Our research focused especially on those differentially expressed genes (DEGs) that were also cosegregating or closely linked to the major alkaloid iucundus locus and major QTLs underlying alkaloid composition. The results of this inquiry provide novel insight into the complex molecular mechanisms surrounding QA biosynthesis in NLL, and broaden our knowledge associated with the alkaloid metabolism in plants.
Results
RNA-Seq and P27255 transcriptome assembly, merging, and annotation
When leaf-derived cDNA libraries of two low-alkaloid (sweet) and two high-alkaloid (bitter) NLL accessions were sequenced (Table 1) and low quality and adaptor sequences were filtered, 632,431,153 high-quality reads remained (90.9% of processed reads). A total of 167,062,967 high-confidence short RNA-Seq reads for the reference P27255 transcriptome (pooled from two replications) were incorporated into the de novo assembly. The high-alkaloid accession P27255 was chosen to assemble the reference transcriptome, as we expected that genes involved in QA biosynthesis are upregulated in bitter accessions.
After merging redundant transcripts from individual assemblies using the tr2aacds pipeline, we obtained 147,912 transcripts (average length 978.33 bp, N50 1750) and 65,835 candidate loci. The resultant merged sequence is hereafter referred to as the P27255 transcriptome. The average GC content of the P27255 transcriptome was 40.09%. Assessment of ortholog completeness (BUSCO v3 with the Embryophyta-specific set of single-copy orthologs) showed that the merged assembly captured more single-copy orthologs than any of the four separate assemblies and contained a low percentage of duplicated data (Fig. 1). A total of 28,769 candidate loci (43.7% of the total loci) were annotated using BLAST2GO, and gene ontology (GO) terms were assigned to 22,703 candidate loci (78.9% of the annotated loci).
Comparison of individual and merged assemblies for P27255 accession based on single-copy ortholog completeness by BUSCO v3. The individual assemblies obtained using four methods (CLC Genomics Workbench, SOAPdenovo-TRANS, Trinity, VELVET + OASES) are shown. The entire merged tr2aacds transcriptome as well as the non-redundant version (using the predicted isoform with highest coding potential (longest coding sequence) for each of the predicted loci) are also shown. Our assessments of the previously available Tanjil and P27255 transcriptomes32 as well as recently available draft NLL proteome34 are included for comparison. Single indicates full-length orthologs; Fragment indicates fragmented orthologs; and Duplicated indicates duplicated orthologs. The numbers under the bars indicate the percentages of each ortholog and the summed total (∑) is also indicated.
Differential gene expression, QA candidate gene selection, and genetic mapping
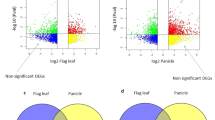
We detected 1489 differentially expressed transcripts (probability of equal expression <0.0001) in the bitter versus sweet accessions of NLL (2 accession in 2 replicates for both bitter and sweet); 1092 were significantly upregulated and 397 were downregulated in bitter accessions (posterior probability of differential expression ≥0.9999). We selected a subset of 962 transcripts with more than two-fold higher expression in bitter accessions (estimated posterior fold change (postFC) >2) for analysis (Supplementary Table S1) with further restriction to a set of 550 transcripts present across both bitter and sweet accessions.
To identify QA candidate genes, we focused mainly on the P27255 transcriptome. Moreover we used the draft NLL genome sequence34 that was released while we were analysing our results, to directly investigate the iucundus region in pseudochromosome NLL-07. A total of 12 candidate transcripts were selected; 11 chosen based on the functional annotation (differentially expressed representatives of transcription factor (TF) families and enzymes associated with legume secondary metabolism) and one that we identified as adjacent to the iucundus locus in the draft NLL genome (associated with primary metabolism) (Table 2). The candidate sequences were used in linkage analysis to determine their genetic positions in relation to the major alkaloid iucundus locus. The candidates were mapped to genome positions using the draft NLL genome (Table 2). Following analyses were focused on candidate QA genes closely linked to the iucundus locus.
Segregation analysis in 93 recombinant inbred lines (RILs) generated new transcriptome-derived markers that were mapped to the NLL linkage groups (LGs) of the Hane, et al.34 map to produce an enriched linkage map (Supplementary Table S2). Among the 12 markers, 10 fitted the expected Mendelian segregation and two showed significant segregation distortions (P < 0.05) that favoured the wild-type parent (Supplementary Table S3). These two markers were located in the iucundus region of NLL-07. The total map length with new markers incorporated was increased by 9.4 cM to 2447.6 cM, but the average interval size remained unchanged (0.82 cM).
We found two transcriptome-derived markers that were closely linked to the iucundus locus and one of them, RAP2-7, represented by P27255_008724 candidate locus (postFC 98.8), co-segregated with iucundus (Supplementary Table S2, Fig. 2). RAP2-7 is located in scaffold_162_1 that was not assigned to a pseudochromosome in the assembly v1.0. A BLASTN search of the P27255_008724 sequence against the annotated coding sequences set (cds v1.0) of the draft NLL genome34 revealed that its longest isoform corresponded to TanjilG_07628 (LOC109342033), which is annotated as APETALA2/ethylene response (AP2/ERF) TF (Table 2). Gene structure analysis revealed that TanjilG_07628 contained eight exons (Fig. 3). Investigation of differences between TanjilG_07628 and the longest coding isoform of P27255_08724 with WebScipio revealed two substitutions, M48V and S196R (the numbering is according to the XP_019435537.1 RefSeq protein for locus TanjilG_07628).
Reliable QTLs mapped to linkage groups NLL-07 and NLL-08 in the reference NLL genetic map34 enriched with new transcriptome-derived markers. For clarity, only a part of each linkage group is shown (NLL-07, 55–100 cM; NLL-08, 70–100 cM). Vertical bars on the left of each linkage group show the positions of the QTLs, with an inner block representing 1-LOD support interval and the end points determining 2-LOD support interval for each QTL location.
Gene models predicted by WebScipio71 (a) RAP2-7 based on sequence of longest coding isoform of candidate locus P27255_08724 (corresponding to TanjilG_07628), (b) DHDPS based on P27255_011214 candidate locus within the merged transcriptome. Dark grey bars and light grey bars mark exons and introns, respectively. Blue lines in the exon denote indel (compared to draft Tanjil genome), red lines denote mismatches (single nucleotide substitutions).
We confirmed the genetic position of DHDPS (P27255_011214 candidate locus with much lower postFC 4.88) by analysing the iucundus region in pseudochromosome NLL-07. DHDPS, which encodes chloroplast 4-hydroxy-tetrahydrodipicolinate synthase, was mapped 0.5 cM from the iucundus (Supplementary Table S2, Fig. 2). The DHDPS sequence is present in the current draft NLL genome, but the NCBI Gnomon pipeline reannotation marked TanjilG_17658 (LOC109352463) as a pseudogene. We detected a single mismatch and a single frameshift-causing deletion when the P27255_011214 transcript and TanjilG_017658 in the draft pseudochromosome NLL-07 sequence (NCBI:NC_032015.1) were compared using WebScipio (Fig. 3). We considered the frameshift-causing deletion of guanine may be one of the factors that distinguish sweet from bitter NLL.
Other transcriptome-derived markers mapped to LGs other than NLL-07 (Table 2 and Supplementary Table S2). Notably, some of the candidate genes identified in our analysis encode enzymes already known to be involved in QA biosynthesis, namely LDC6 (P27255_008143, NLL-15), LaAT 2 (P27255_007730, NLL-16), HMT/HLT11 (P27255_007723, NLL-04), and LaCAO7 (P27255_002184, NLL-15).
Validation of RNA-Seq data by qRT-PCR
To assess the accuracy of the RNA-Seq data, candidate QA transcripts were incorporated into qRT-PCR analysis. The eleven DEGs were found to be significantly differentially expressed (P < 0.0001) in the qRT-PCR analysis, whereas HMT/HLT was not (P = 0.003, 2.06-fold change). In the bitter accessions, LaCAO and LDC had the highest mean fold change (1666-fold and 1450-fold upregulation accordingly), followed by CCR (415-fold), LaAT (358-fold), CES1L (91-fold), DFR1 (64-fold), RAP2-7 (63-fold), F3H (61-fold), MYB (53-fold), LDOX (42-fold), and DHDPS (4-fold) (Fig. 4). The expression levels of QA candidate genes were normalized using the reference genes, alpha tubulin (TUBA), ATP synthase (ATPsyn), and alcohol dehydrogenase class-3 (ADH3) (see Supplementary Data S1 for details).
Expression profiles of 12 selected candidate QA genes by qRT-PCR. Bars indicate mean fold change expression in the bitter vs. sweet NLL accessions. Relative quantification was determined by qRT-PCR analyses normalized to three reference genes (alpha tubulin, ATP synthase, and alcohol dehydrogenase class-3).
QTL mapping and statistical analysis of bitter/sweet phenotypes
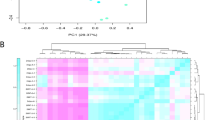
The principal component biplot for two subgroups of RILs established based on their iucundus alleles shows the distribution of total QA content (estimated as % of seed dry weight) and relative abundance of individual QAs (% of total QAs) (Fig. 5). The low-alkaloid subgroup showed greater relative abundance of isolupanine, whereas the high-alkaloid subgroup had higher relative abundance of angustifoline, as well as the expected higher total QA content. The relative abundances of 13-hydroxylupanine and lupanine were similar in the two subgroups. Mean values for total QA content and relative abundance of individual QAs for the two RIL subgroups are given in Supplementary Table S4.
The distribution for total QA content was bimodal with apparent separation of low- and high-alkaloid RILs, whereas the distribution for relative abundance of isolupanine was left-skewed across the 5 years (Supplementary Fig. S1). The normality test showed that for some phenotypes the data did not follow a normal distribution (Supplementary Table S5). In the assessment of variance components, higher variance was observed for lines compared with years (Supplementary Table S6). The high broad-sense heritability justifies the conditions for QTL analysis (Supplementary Table S6).
QTL analysis performed to confirm the involvement of the iucundus region and detect other candidate genomic regions that may contribute to QA biosynthesis in NLL, identified 47 loci underlying the traits of total QA content and relative abundance of individual QAs across the 5 years (Supplementary Table S7). QTLs for the same and/or different traits were found to be colocalised on the LGs (overlapping map intervals) across years.
Colocalisation of reliable QTLs for different QAs was identified mainly in the iucundus region of NLL-07 (Fig. 2), confirming it as a major region involved in the QA biosynthesis pathway. The colocalising QTLs included those detected each year for total QA content, which individually accounted for 18.9–70.7% of the phenotypic variance (LOD values 7.8–27.8), as well as QTLs for relative abundance of angustifoline detected in four years (2008, 2010, 2011, and 2015) that were individually responsible for 12.6–56.4% of the phenotypic variance (LOD values 3.9–21.7). In both cases, QTLs originating from alleles of the paternal line (P27255) were responsible for increased total QA content and relative abundance of angustifoline in seeds. Colocalising QTLs for relative abundance of isolupanine detected in the iucundus region in four years (2008, 2010, 2013, and 2015) explained 11.4–20.3% of the trait variation (LOD values 3.6–6.9). These QTLs originated from alleles of the maternal line (83 A:476) and were responsible for increased abundance of isolupanine in seeds.
Colocalising QTLs for relative abundance of isolupanine across 5 years were also detected in the 79–90.2 cM map interval of NLL-08, which contains the carboxylesterase 1 (CES1L) gene that was mapped in this study (Fig. 2). These stable QTLs were individually responsible for 25.2–34.2% of the phenotypic variance (LOD values 7.2–9.9), originated from alleles of the paternal line P27255, and were responsible for increased abundance of isolupanine.
No consistent distribution of QTLs across years was detected for relative abundance of lupanine and 13-hydroxylupanine.
Phylogeny of RAP2-7 homologs in Viridiplantae genomes
A maximum-likelihood phylogenetic tree was constructed to reveal the evolutionary relationship between the candidate AP2/ERF TF gene RAP2-7 (P27255_008724) and related genes in available Viridiplantae genomes (Supplementary Fig. S2). The NLL sequences clustered with other AP2/ERF sequences that were excluded in previous studies, namely At2g39250 (SCHNARCHZAPFEN, SNZ), At3g54990 (SCHAFLMÜTZE, SMZ), and At5g60120 (TARGET OF EARLY ACTIVATION TAGGED 2, TOE2)42,43. These genes were considered to encode a AP2/ERF domain, but were distinct from the typical ERF-type and more closely related to the AP2-type. A cluster of 65 AP2/ERF amino acid sequences, including three lupin sequences (TanjilG_07628 (LOC109342033), TanjilG_14185 (LOC109346040), and TanjilG_11298 (LOC109351048)), was formed in our phylogenetic tree. TanjilG_07628 was located in scaffold_162_1, TanjilG_14185 in NLL-04, and TanjilG_11298 in NLL-06 of the lupin genome. Differential expression analysis of these genes against sequences from the draft NLL genome34 showed that TanjilG_07628 and TanjilG_14185 were upregulated in the bitter accessions (postFC 65 and 500 respectively), whereas TanjilG_11298 was not significantly upregulated (postFC 1.8). Cross-referencing the draft NLL genome with the P27255 transcriptome revealed that TanjilG_07628 as well as several isoforms derived from TanjilG_14185 were putatively assembled as one candidate locus (RAP2-7, P27255_008724). However, when assessed by RSEM, the expression levels of the TanjilG_07628-derived isoforms were postFC 98.8, whereas TanjilG_11298 was not significantly differentially expressed. It is worth noting that, based on the consensus tree topology, TanjilG_07628 and TanjilG_14185 are recent paralogs (100% support), and are part of a strongly separated monophyletic subtree that includes AP2/ERF homologs from other legumes. Interestingly, the legume clade was identified as sister to another subtree that included the known Brassicaceae AP2/ERFs genes SMZ (At3g54990, Bra007123) and SNZ (At2g39250). We identified TOE2 as a distant homolog of TanjilG_07628. The three genes, SMZ, SNZ and TOE2, are known to repress flowering.
Discussion
Based on the assumption that genes involved in QA biosynthesis are upregulated in leaves of bitter accessions, we assembled a reference transcriptome for high-alkaloid P27255, as the reference genotype very likely to express alkaloid-associated transcripts at consistent levels that would enable the assembly of good-quality, full-length transcripts. Recent works have shown that, typically, no single assembly captures all aspects of a transcriptome equally well, so an approach that combines multiple methods is more likely to produce a well supported assembly (e.g. Nakasugi, et al.44, Chen, et al.45). Thus, we used four individual assemblers that were merged in the EvidentialGene tr2aacds pipeline. The high-quality P27255 transcriptome that we obtained was more comprehensive than the previously reported Tanjil transcriptome32 and rivalled the currently available draft NLL genome34, regarding the ortholog completeness assessment (Fig. 1).
Based on the P27255 transcriptome and the iucundus region in the draft NLL genome34 we selected 12 candidate genes connected to QA biosynthesis and in the subsequent analyses we focused on the candidates most closely linked to iucundus locus and the major QTL underlying alkaloid composition. The linkage analysis of selected candidates confirmed that major QA genes colocalised with the major alkaloid iucundus locus on NLL-07. Mapping of QTLs underlying total QA content and relative abundance of individual QAs in NLL seeds confirmed iucundus as a major QA locus as well as identified other genomic regions involved in QA biosynthesis. The key finding of this study was the identification of RAP2-7 (P27255_008724), an AP2/ERF TF that cosegregated with the iucundus locus, as well as DHDPS (P27255_011214, TanjilG_017658) that was located 0.5 cM from the iucundus. Both genes were located within an interval related to highly significant QTLs on NLL-07.
We demonstrated here, for the first time, that AP2/ERF TF is likely to be involved in the regulation of QA biosynthesis in NLL. The AP2/ERF TF superfamily has been divided into four families, AP2, ERF, RAV, and Soloist, based on the number of AP2 and other DNA-binding domains46, and has been well-studied in model plant systems, such as A. thaliana L. and rice (Oryza sativa L.). The ERF subfamily comprises 12 subgroups (I–XII) defined by Nakano, et al.42, and several members of ERF group IX were later found to regulate the biosynthesis of distinct classes of alkaloids in different plant species. For example, in C. roseus L., CrORCA2 and CrORCA3 regulated the synthesis of terpenoid indole alkaloids47,48; in Nicotiana tabacum L., the NIC2 locus was involved in the synthesis of a pyridine alkaloid of nicotine49; in tomato and potato, GAME9 and JRE4 regulated the expression of steroidal glycoalakaloids22,50; and in Ophiorrhiza pumila Champ. ex Benth, OpERF2 regulated the production of monoterpenoid indole alkaloid camptothecin43. The phylogenetic relationships of NLL RAP2-7 within the AP2/ERF TF superfamily (Supplementary Fig. S2) showed that it was a homolog of a distant subset of atypical A. thaliana AP2/ERF TFs, AT5G60120 (TOE2), AT3G54990 (SMZ), and AT2G39250 (SNZ), which were previously rejected from the Nakano, et al.42 and Udomsom, et al.43 large-scale phylogenies as outliers. Notably, TOE2 and SMZ and its paralog SNZ belonged to an AP2/ERF clade that contained repressors of flowering and were predicted targets of microRNA miR17251,52,53. The downregulation of AP2-like target genes by miR172 resulted in the promotion of flowering51. The involvement of NLL RAP2-7 in flowering regulation has not been reported so far and, to our knowledge, the association of alkaloid content and flowering time has not been investigated for lupins. Thus, despite the apparent analogy in the regulation of alkaloid biosynthesis in NLL and species with distinct alkaloid classes, such as C. roseus, O. pumila, and Solanaceae, a different molecular background may have evolved in lupins. This confirms the previous observation that unlike other groups of secondary metabolites, the biosynthesis and phylogeny of the numerous and structurally diversified classes of alkaloids are seemingly unrelated (or very distantly related)12.
We found that RAP2-7 was located in an unplaced scaffold_162_1 that was characterized by long strings of unknown bases as well as transposon sequences, which may explain why it was not assigned to pseudochromosome NLL-07 in the currently available draft NLL genome34. Remarkably, the candidate DHDPS gene was identified in the iucundus region of pseudochromosome NLL-07. The structures of both scaffold_162_1 and the genomic region surrounding the iucundus locus imply some underlying assembly issues (the pseudochromosome region contains 48 gaps of unknown length, including a 77-kbp gap upstream of DHDPS), which could be indicative of the activity of repetitive elements during evolution. Significant segregation distortions for genetic markers neighbouring the iucundus locus have been reported by Boersma, et al.33, and were also apparent for the RAP2-7 and DHDPS markers mapped in this study.
DHDPS homologs are typically involved in the L-lysine biosynthesis subpathway, a known precursor of QAs. The close location of DHDPS to iucundus and RAP2-7 suggests but does not yet confirm the role of this NLL genomic region in QA biosynthesis. The other QA candidate genes investigated in our study were assigned to LGs other than NLL-07 (Table 2), and so were the known structural QA genes also identified in our analysis, namely LDC (NLL-15)6, LaCAO (NLL-15)7, LaAT (NLL-16)2, and HMT/HLT (NLL-04)2. Therefore, our results suggest that QA genes are not arranged in a cluster centered on the iucundus locus, or that the cluster is not very big. However, additional genes involved in QA biosynthesis and regulation may be recognized in the iucundus region when NLL genomic resources are enhanced. Indeed, clustering of nonhomologous biosynthetic genes of specific alkaloid pathways was reported previously in plants54. The GAME genes in the core steroidal glycoalkaloid biosynthesis pathway were found to be organized in metabolic gene clusters in the potato and tomato genomes55. Similarly, a cluster of genes encoding enzymes involved in noscapine biosynthesis was found in opium poppy56. Additionally, ERF-controlled gene clusters have been found in some plant genomes, including N. tabacum and other Solanaceae representatives22,49. The refinement of genetic maps, such as the very recently published high density map of Zhou, et al.57, may increase the identification of novel QA genes clustered in the iucundus region.
Despite the identification of several QA biosynthesis-related genes, the QA biosynthetic pathway in NLL is still far from being understood. Based on RNA-Seq data all the QA candidate genes were present in NLL genotypes with different alkaloid accumulation levels. Differential expression in the bitter vs. sweet accessions was also tested by qRT-PCR for twelve candidate QA genes (Fig. 4). Notably, in recent work of Yang et al.7 the researchers have identified and validated enzymatic function of a copper amine oxidase (LaCAO) found to be co-expressed with LDC. In their RNA-seq data and further qRT-PCR assay LDC and LaCAO expression was the highest in leaves, stems and pedicles. At the same time LDC expression was not detected in large seeds. The expression of characterised QA biosynthetic genes (LDC, LaCAO and LaAT) in bitter NLL accession P27255 and two sweet cultivars: Tanjil and Unicrop was also investigated by Frick et al.15. Significantly higher expression of these genes in leaf tissue points to leaves rather than seeds as the focal point of the alkaloid biosynthesis. Moreover, other, novel candidate QA genes were identified, which exhibited expression patterns similar to LDC and LaCAO in qRT-PCR analyses. These were chiefly annotated as major latex-like proteins (LaMPL1-like, LaMPL2-like and LaMPL4-like,) and were found to be located in NLL-10 (Lup019334) and NLL-06 (Lup015922 and Lup015923) groups in the draft genome15. The low level expression of QA candidates from this and other, recent studies4,7 in sweet accessions updates previously published results where LDC and LaAT expression was detected only in bitter NLL cultivars by gel-based RT-PCR2,6. Thus, the initial step of QA biosynthesis and some subsequent steps were not completely blocked at the transcriptional level in sweet NLL genotypes, as initially proposed by Hirai, et al.14. Furthermore, despite the different quantities, the same end products of QA biosynthesis were identified in both NLL genotypes. The observed downregulation of LDC in sweet NLL genotypes implied that the QA biosynthesis pathway was regulated at the initial step of lysine decarboxylation, whereas qualitative differences in QA composition of sweet and bitter RILs (Table 1) implied additional regulatory steps in the QA biosynthesis pathway downstream of lupanine. Our HMT/HLT gene expression results are in concurrence with previous studies that reported HMT/HLTase activity was similar in both sweet and bitter forms of L. angustifolius, L. albus, and L. luteus, and support the earlier hypothesis that HMT/HLT was regulated independently from genes that determine alkaloid accumulation and its expression was uncorrelated with final concentrations of alkaloids in seeds11,13,14.
As a whole, the growing body of evidence implies interaction of genes underlying QA biosynthesis in a form of gene network. Gene network reconstruction is a powerful approach for understanding the complex biological systems that underlie phenotypes58. The abundance of genomics/transcriptomics data based on next-generation sequencing technology enabled recently deeper investigations of gene networks in many plants, including legume representatives59,60,61. The insights from such datasets (or their lupin counterparts, once these become available), can be expected to allow for future quantitative as well as qualitative description of regulatory principles guiding secondary metabolism, including alkaloid biosynthesis.
Within the present study, we identified, for the first time, individual QTLs underlying the total QA content and relative abundance of major alkaloids in NLL seeds (Supplementary Table S7). Credible QTLs for total QA content as well as relative abundance of angustifoline and isolupanine were consistently detected over a period of 4–5 years, indicating that they were not affected by the environment. We uncovered strong support for the colocalisation of these QTLs in the iucundus region using the transcriptome-derived candidate RAP2-7 and DHDPS loci (Fig. 2). Colocalisation of QTLs for different QAs may indicate that a gene in the colocalisation site determines the key step in the biosynthesis/accumulation of all compounds, possibly through a common regulatory mechanism. Stable QTLs in the iucundus region that accounted for the high phenotypic variability (Supplementary Table S7), confirmed that this region may govern the most decisive steps in QA accumulation. RAP2-7 is likely the gene underlying the major QTL and it may be useful for MAS in breeding programs to lower QA levels in NLL seeds. The major QTL in NLL-08 for relative abundance of isolupanine contained the mapped QA marker CES1L, and its homologs are known to be involved in secondary metabolism in plants, including noscapine biosynthesis in the opium poppy62. No stable QTLs for relative abundance of lupanine and 13-hydroxylupanine were detected in the NLL genome. Similar tendencies were observed for the bitter and sweet RIL subgroups (Fig. 5).
In conclusion, our findings provide crucial insights into the QA biosynthesis pathway and its transcriptional regulation in NLL. The results strongly suggest that QA biosynthesis in this species is controlled mostly at the level of transcriptional regulation. We suggest that our characterization of RAP2-7 is a worthwhile starting point for better disentangling the signalling and transcriptional regulatory networks associated with QA biosynthesis in this important legume crop. Moreover, the identified and characterized DEGs are a valuable resource to further explore the mechanisms underlying QA biosynthesis, for functional studies, and for the development of markers that are useful in MAS.
Materials and Methods
Plant material
Seeds of the investigated NLL accessions deposited in the Polish Lupinus Gene Bank and breeding programs were provided by Poznan Plant Breeders Ltd., Wiatrowo Branch (Poland). Seeds of the parental lines (83A:476 and P27255) and 93 RILs were provided by the Department of Agriculture and Food, Western Australia (DAFWA, Australia). All seeds were grown in early spring (March/April) with natural vernalisation in a field experiment in Wiatrowo, Poland. Mechanical weed control treatment as well as P2O5 (60 kg/h) and K2O (90 kg/h) fertilisers were applied.
Accessions used for RNA-Seq and qRT-PCR were grown in 2014 (Table 1). Young leaves were collected at flowering time and immediately frozen at −80 °C. Flowering time varied among lines and the sample collection proceeded for each plant when most of the flowers on the main stem inflorescence were open (erect standard petal).
For QTL mapping, the field trials included the RILs of the mapping population and parental lines, which were sown in a completely randomized design with two replicates (5 seeds per plot) in five growing seasons: 2008, 2010, 2011, 2013, and 2015. Seed samples for gas chromatography analyses were harvested from the RILs each year after full maturity.
Assessment of total QA content and relative abundances of QAs in NLL seeds
Quantitative and qualitative QA composition were evaluated by gas chromatography (GC-2014; Shimadzu, Kyoto, Japan) to validate the alkaloid profiles of seeds from the selected accessions (Table 1) and from 80 representative RILs (mixture of seeds from a plot × 2 replicates, each growing seasons). Alkaloid extraction and analyses were conducted as described by Kamel, et al.25. The results were received as mean values from the two gas chromatography replicates for each RIL. Total QA values are the percentage of the sum of the major QAs (lupanine, 13-hydroxylupanine, angustifoline, and isolupanine) of the seed dry weight (% of seed dry weight). Relative abundance of major QAs were assessed as the percentage of total QAs (sum of all QAs = 100%).
RNA and DNA extraction
Total RNA was isolated from 30 mg of ground leaf tissue using a SV Total RNA Isolation System Kit (Promega, Madison, WI). Total RNA concentration and quality were determined using a 2100 Bioanalyzer (Agilent Technologies, Palo Alto, CA) with a minimum RNA integrity number of 8. The extracted RNA was used for RNA-Seq and qRT-PCR analyses.
Genomic DNA was isolated from frozen, young leaves using a DNeasy Plant Mini Kit (Qiagen, Germantown, MD). DNA concentration and quality were determined using a NanoDrop ND-1000 (Thermo Fisher Scientific, Waltham, MA).
cDNA library preparation and transcriptome sequencing (RNA-Seq)
Two low-alkaloid (sweet, 83A:476 and W-225) and two high-alkaloid (bitter, P27255 and Vitigudino-1) NLL accessions were used for RNA-Seq in two biological replicates (Table 1). cDNA library preparation was conducted in the Institute of Bioorganic Chemistry, Polish Academy of Sciences using a TruSeq RNA Sample Preparation Kit v2 (Illumina, San Diego, CA). Library sequencing was performed on an Illumina platform (HiSeq 1500, PE 2 × 75 bp, HiSeq v4 reagent sets) in the Medical University of Warsaw, Poland. The RNA-Seq data have been submitted to GenBank as short-read archives under the following accessions: SRR5723679, SRR5723680, SRR5723681, and SRR5723682 in BioProject PRJNA389154.
Transcriptome de novo assemblies of accession P27255, merging, and annotation
The raw data were filtered to remove low-quality reads (<Q30) and reads with terminal Illumina primer and adaptor sequences using Cutadapt 1.163 with the following parameters: times 3, overlap 10, minimum length 25. Reads with a discarded pair <25 bp were kept as singleton reads.
The pre-processed high-quality paired reads of high-alkaloid accession P27255 were assembled de novo using four methods: Trinity 2.2.0 with glue and k-mer coverage thresholds of 564; SOAPdenovo-TRANS 1.03 with k-mer length values from 31–71 in steps of 1065; VELVET 1.2.10 (with k-mer values from 25–65 in steps of 10) and OASES 0.2.0966,67; and CLC Genomics Workbench 9.0.1 with default settings (https://www.qiagenbioinformatics.com/). Because of memory constraints for the VELVET + OASES assembly, the data were further filtered (digitally normalized) with KHMER 2.068 at a uniform coverage value C of 100. Non-redundant transcripts from the four P27255 assemblies were merged using the EvidentialGene tr2aacds pipeline (http://arthropods.eugenes.org/genes2/about/EvidentialGene_trassembly_pipe.html). We chose P27255 as the reference transcriptome based on the assumption that genes involved in QA biosynthesis will be upregulated in the high-alkaloid accessions.
The individual P27255 assemblies were compared based on single-copy ortholog completeness by BUSCO v369 with the Embryophyta reference core genes (dataset embryophyta_odb9). The ortholog completeness assessment results for the currently available Tanjil transcriptome32 and the draft NLL genome34 were included for comparison. The assembled transcripts were functionally annotated using Blast2GO Pro v.4.1.5 by aligning them against UniProt/SwissProt70. The merged P27255 transcriptome sequence has been deposited in GenBank as a transcriptome shotgun assembly (TSA) under accession GGED00000000.
The draft NLL genome34 was used as the reference to determine the genome positions of the investigated QA candidates (denominated as TanjilG_number in the NLL genome at NCBI and equivalent to Lup_number of Hane, et al.34). The precise exon/intron structures of selected QA candidate genes were assessed based on the predicted protein sequence of the transcript and the draft NLL genome using WebScipio71.
Differential gene expression and selection of QA candidate genes
Differentially expressed genes between bitter and sweet NLL accessions were detected using RSEM 1.2.2872. Twelve candidate genes were selected for thorough investigation (Table 2) using two different approaches:
Analysis of the assembled P27255 transcriptomes
As sweet NLL accessions are known to produce low amount of alkaloids we assumed that they possess genetic determinants required for biosynthesis. Therefore candidate DEGs that were overexpressed in the bitter accessions were filtered based on multiple serially applied criteria. First, only DEGs with a maximum posterior probability of equal expression value of 0.0001 (posterior probability that a gene/transcript is equally expressed ≤0.0001, which corresponds to having used the rsem-control-fdr algorithm with default hard threshold parameter turned on) were selected. Second, from among these DEGs, only those with more than two-fold higher expression than the sweet accessions baseline were selected (RSEM was used to estimate fold changes between groups; PostFC >2, see Supplementary Table S1 for full list of candidate genes.) Third, the selected DEGs were further filtered to retain only those with read counts in both the sweet and bitter accessions. Finally, based on the annotations and GO terms, 10 candidate genes related to plant secondary metabolism and its regulation were selected for an in-depth investigation. QA biosynthesis pathway is not well understood regarding candidate enzymes involved in oxidation/reduction processes leading to ring closure4,5,7,8,9 as well as modifications of the final QA products. Therefore we also considered singular representatives for the oxidoreductive enzyme families and transcription factors known to be involved in other secondary metabolism pathways, both due to possible involvement in QA modification and due to possibility of correlation/cross-talk between different metabolite groups. Additionally, a candidate locus annotated as HMT/HLT was selected for further analyses despite its equal expression in the bitter and sweet accessions RNA-Seq libraries, because of its verified role in QA biosynthesis11.
Analysis of the iucundus region in pseudochromosome NLL-07
We assumed that some candidate DEGs may be present in the draft NLL genome, particularly in the region of the major alkaloid iucundus locus in pseudochromosome NLL-07 (i.e., between markers LaDArT_PAV21233 and LaSSR_025 on the molecular genetic map of Hane, et al.34 corresponding to physical interval NLL-07:14591111–15337202). Two DEGs were found in this region: P27255_011214 (equivalent to TanjilG_017658 by reciprocal BLAST search) predicted to encode 4-hydroxy-tetrahydrodipicolinate synthase (DHDPS), and P27255_002404 (equivalent to TanjilG_007701) annotated as metalloprotease and, hence unlikely to be associated with QA biosynthesis. We selected only P27255_011214 for further analyses.
Linkage mapping of candidate QA genes
PCR primers were designed using both the assembled transcripts and pre-processed raw read sequences. Transcriptome-derived SNP markers were designed for the 12 selected candidate QA genes (Table 2). The primer sequences and PCR thermal conditions are listed in Supplementary Table S3. Segregation analysis was performed for 93 RILs. Transcriptome-derived markers were distributed within the linkage groups of the reference genetic map34 limited to framework markers and six trait loci (total 2963 loci) using MapManager QTXb2073. The initial map imported to MapManager software (based on the 2963 loci) was 2438.2 cM, with average interval size 0.82 cM (compared with 2500.8 cM and 0.85 cM, respectively, in the genetic map of Hane, et al.34). For each locus the observed segregations were tested against the expected Mendelian segregation ratio (1:1) using the chi-square test. The LGs was visualized using MapChart74.
Validation of gene expression by qRT-PCR
To confirm the expression levels obtained using the RNA-Seq data, we performed qRT-PCRs for 12 QA candidates, including 11 DEGs and one gene (HMT/HLT) that was expressed constitutively in the sweet and bitter accessions. First, seven housekeeping genes in the NLL transcriptome, namely actin 2/7 (ACT2/7), ADH3, ATPsyn, cyclophilin (CYP), elongation factor 1-beta (ELF1B), glucose-6-phosphate 1-dehydrogenase (G6PD), TUBA were tested as candidate reference genes for the qRT-PCRs. We selected the three most stable reference genes, TUBA, ATPsyn, and ADH3, to normalize the expression levels of the selected QA candidate genes (see Supplementary Data S1 for details).
Seven low-alkaloid and seven high-alkaloid NLL accessions were selected for the qRT-PCR analyses (Table 1). Primer pairs and probes were designed based on DNA sequence of candidate QA and reference genes for the 83A:476 and P27255 accessions. The qRT-PCRs were performed as one-step reactions using LightCycler 480 RNA Master Hydrolysis Probes on a LightCycler480 system (Roche, Mannheim, Germany) according to the manufacturer’s protocol (reaction volume 10 µl). Three biological and two technical replicates of each accession, as well as a negative control, were included in each assay. PCR amplification efficiencies ranged from 0.97 to 1.09 (Supplementary Table S8), as determined using a standard curve derived from a pooled RNA mixture. The primer/probe sequences and reaction conditions for the qRT-PCRs are listed in Supplementary Table S8.
Gene expression was analysed using the delta-delta Ct method with the three reference genes (TUBA, ATPsyn, and ADH3). All Cq values were subjected to efficiency corrections75. Statistical testing of differences in mean Cq values was conducted using the Wilcoxon signed rank test corrected for multiple comparisons (Bonferroni correction; final P-value threshold of 0.0001).
QTL mapping and statistical analysis of the bitter/sweet phenotypes
Phenotypic data distribution was evaluated across 5 years for both total QA content and relative abundance of four major QAs. Variance components for years and lines were estimated by the restricted maximum likelihood method, and broad-sense heritability was computed using the formula of Cullis, et al.76. A principal component biplot was created to visualize the distribution of of total QA content and relative abundance of individual QAs in the sweet and bitter subgroups of RILs established based on their iucundus allele. These analyses were performed in Genstat 1877.
QTL analysis was conducted on mean values for RILs, for each year separately using the composite interval mapping approach in the QTL Cartographer 2.5 software78. The genome was scanned at a walking speed of 1 cM. Significance threshold values of LOD for QTL detection were determined by a permutation test with 1000 replicates (P value < 0.05). The percentage variations explained by each QTL and additive effect were estimated. The QTL confidence intervals were determined as one-LOD and two-LOD support intervals79.
Phylogeny of ethylene-responsive TF (RAP2-7) homologs in Viridiplantae genomes
Available Viridiplantae genomes (Supplementary Table S9) were annotated with known ERFs using iTAK software based on the presence of sequence signatures of associated protein domains and conserved motifs80. The identified AP2/ERF TFs were compared all-against-all (USEARCH/UBLAST) and clustered using the Markov cluster algorithm81 based on expectation values. Clustering quality was tested based on silhouette width as described by Koczyk, et al.82. The optimal parameter to discriminate the cluster containing candidate NLL TFs was set as I = 1.7. For phylogeny reconstruction, all sequences <200 amino acids in length were removed from the cluster, the remaining sequences were subsequently aligned using MAFFT-LINSI83. In particular, several partial sequences derived from Selaginella, Coccomyxa, and Chlamydomonas, as well as one Medicago truncatula gene sequence (Mtr_7g061190) were rejected as too short and/or lacking class-specific conserved residues. Finally, a maximum-likelihood tree was constructed by IQTREE 1.5.5 based on aligned protein sequences84 (ultrafast bootstrap replicates threshold set to 3000). The evolutionary model was selected by the program according to the Bayesian information criterion (JTT + F + I + G4). Chlamydomonas reinhardtii and Volvox carteri sequences represented the outgroup that was used to root the final tree.
Data Availability
Transcriptome Shotgun Assembly project has been deposited in GenBank under the accession GGED00000000. RNA-Seq data have been submitted as short-read archives under the following accessions: SRR5723679, SRR5723680, SRR5723681, and SRR5723682 in BioProject PRJNA389154.
References
Wink, M., Meißner, C. & Witte, L. Patterns of quinolizidine alkaloids in 56 species of the genus Lupinus. Phytochemistry 38, 139–153 (1995).
Bunsupa, S., Okada, T., Saito, K. & Yamazaki, M. An acyltransferase-like gene obtained by differential gene expression profiles of quinolizidine alkaloid-producing and nonproducing cultivars of Lupinus angustifolius. Plant Biotechnol. 28, 89–94 (2011).
Lee, M. J., Pate, J. S., Harris, D. J. & Atkins, C. A. Synthesis, transport and accumulation of quinolizidine alkaloids in Lupinus albus L. and L. angustifolius L. J. Exp. Bot. 58, 935–946 (2007).
Bunsupa, S., Yamazaki, M. & Saito, K. Quinolizidine alkaloid biosynthesis: recent advances and future prospects. Front. Plant Sci. 3, 239 (2012).
Wink, M. & Witte, L. Quinolizidine alkaloids as nitrogen source for lupin seedlings and cell cultures. Zeitschrift für Naturforschung C 40, 11–12 (1985).
Bunsupa, S. et al. Lysine decarboxylase catalyzes the first step of quinolizidine alkaloid biosynthesis and coevolved with alkaloid production in leguminosae. Plant Cell 24, 1202–1216 (2012).
Yang, T. et al. Transcript profiling of a bitter variety of narrow-leafed lupin to discover alkaloid biosynthetic genes. J. Exp. Bot. 68, 5527–5537 (2017).
Frick, K. M., Kamphuis, L. G., Siddique, K. H., Singh, K. B. & Foley, R. C. Quinolizidine alkaloid biosynthesis in lupins and prospects for grain quality improvement. Front. Plant Sci. 8, 87 (2017).
Schafer, H. & Wink, M. Medicinally important secondary metabolites in recombinant microorganisms or plants: progress in alkaloid biosynthesis. Biotechnol J 4, 1684–1703 (2009).
Suzuki, H., Murakoshi, I. & Saito, K. A novel O-tigloyltransferase for alkaloid biosynthesis in plants. Purification, characterization, and distribution in Lupinus plants. J. Biol. Chem. 269, 15853–15860 (1994).
Okada, T., Hirai, M. Y., Suzuki, H., Yamazaki, M. & Saito, K. Molecular Characterization of a Novel Quinolizidine Alkaloid O-Tigloyltransferase: cDNA Cloning, Catalytic Activity of Recombinant Protein and Expression Analysis in Lupinus Plants. Plant Cell Physiol. 46, 233–244 (2005).
Facchini, P. J. & St-Pierre, B. Synthesis and trafficking of alkaloid biosynthetic enzymes. Curr. Opin. Plant Biol. 8, 657–666 (2005).
Saito, K., Koike, Y., Suzuki, H. & Murakoshi, I. Biogenetic implication of lupin alkaloid biosynthesis in bitter and sweet forms of Lupinus luteus and L. albus. Phytochemistry 34, 1041–1044 (1993).
Hirai, M. Y., Suzuki, H., Yamazaki, M. & Saito, K. Biochemical and partial molecular characterization of bitter and sweet forms of Lupinus angustifolius, an experimental model for study of molecular regulation of quinolizidine alkaloid biosynthesis. Chem. Pharm. Bull. 48, 1458–1461 (2000).
Frick, K. M. et al. Characterization of the genetic factors affecting quinolizidine alkaloid biosynthesis and its response to abiotic stress in narrow-leafed lupin (Lupinus angustifolius L.). Plant Cell Environ 23, 13172 (2018).
Strickler, S. R., Bombarely, A. & Mueller, L. A. Designing a transcriptome next-generation sequencing project for a nonmodel plant species. Am. J. Bot. 99, 257–266 (2012).
Martin, L. B., Fei, Z., Giovannoni, J. J. & Rose, J. K. Catalyzing plant science research with RNA-seq. Front. Plant Sci. 4, 66 (2013).
Garg, R. & Jain, M. Transcriptome Analyses in Legumes: A Resource for Functional Genomics. Plant Genome 6, 3 (2013).
Rezaei, M. K., Deokar, A. & Tar’an, B. Identification and expression analysis of candidate genes involved in carotenoid biosynthesis in chickpea seeds. Front. Plant Sci. 7, 1867 (2016).
Li, C. et al. Identification of candidate genes for resistance to Soybean mosaic virus strain SC3 by using fine mapping and transcriptome analyses. Crop Pasture Sci. 68, 156–166 (2017).
Rai, M. et al. De Novo RNA Sequencing and Expression Analysis of Aconitum carmichaelii to Analyze Key Genes Involved in the Biosynthesis of Diterpene Alkaloids. Molecules 22, 12 (2017).
Cardenas, P. D. et al. GAME9 regulates the biosynthesis of steroidal alkaloids and upstream isoprenoids in the plant mevalonate pathway. Nat. Commun. 7, 10654 (2016).
Lucas, M. M. et al. The future of lupin as a protein crop in Europe. Front. Plant Sci. 6, 705 (2015).
Kroc, M. et al. Quantitative and qualitative analysis of alkaloids composition in the seeds of a white lupin (Lupinus albus L.) collection. Genet. Resour. Crop Evol. 64, 1853–1860 (2017).
Kamel, K. A., Święcicki, W., Kaczmarek, Z. & Barzyk, P. Quantitative and qualitative content of alkaloids in seeds of a narrow-leafed lupin (Lupinus angustifolius L.) collection. Genet. Resour. Crop Evol. 63, 711–719 (2016).
Cowling, W. A., Buirchell, B. J. & Tapia, M. E. Lupin. Lupinus L. Promoting the conservation and use of underutilized and neglected crops. 23., (International Board for Plant Genetic Resources (IBPGR), 1998).
Hondelmann, W. The lupin - ancient and modern crop plant. Theor. Appl. Genet. 68, 1–9 (1984).
Gladstones, J. S. Lupins as crop plants. Field Crop Abstracts 23, 123–148 (1970).
Nelson, M. N. et al. Aligning a new reference genetic map of Lupinus angustifolius with the genome sequence of the model legume, Lotus japonicus. DNA Res. 17, 73–83 (2010).
Nelson, M. N. et al. The first gene-based map of Lupinus angustifolius L.-location of domestication genes and conserved synteny with Medicago truncatula. Theor. Appl. Genet. 113, 225–238 (2006).
Kroc, M., Koczyk, G., Święcicki, W., Kilian, A. & Nelson, M. New evidence of ancestral polyploidy in the Genistoid legume Lupinus angustifolius L. (narrow-leafed lupin). Theor. Appl. Genet. 127, 1237–1249 (2014).
Kamphuis, L. G. et al. Transcriptome sequencing of different narrow-leafed lupin tissue types provides a comprehensive uni-gene assembly and extensive gene-based molecular markers. Plant Biotechnol. J. 13, 14–25 (2015).
Boersma, J. G. et al. Construction of a genetic linkage map using MFLP and identification of molecular markers linked to domestication genes in narrow-leafed lupin (Lupinus angustifolius L.). Cell. Mol. Biol. Lett. 10, 331–344 (2005).
Hane, J. K. et al. A comprehensive draft genome sequence for lupin (Lupinus angustifolius), an emerging health food: insights into plant–microbe interactions and legume evolution. Plant Biotechnol. J. 15, 318–330 (2017).
Li, X., Yang, H., Buirchell, B. & Yan, G. Development of a DNA marker tightly linked to low-alkaloid gene iucundus in narrow-leafed lupin (Lupinus angustifolius L.) for marker-assisted selection. Crop Pasture Sci. 62, 218–224 (2011).
Lesniewska, K. et al. Assignment of 3 genetic linkage groups to 3 chromosomes of narrow-leafed lupin. J. Hered. 102, 228–236 (2011).
Ksiazkiewicz, M. et al. Comparative genomics of Lupinus angustifolius gene-rich regions: BAC library exploration, genetic mapping and cytogenetics. BMC Genomics 14, 79 (2013).
Ksiazkiewicz, M. et al. Remnants of the legume ancestral genome preserved in gene-rich regions: insights from physical, genetic, and comparative mapping. Plant Mol. Biol. Rep. 33, 84–101 (2015).
Wyrwa, K. et al. Integration of Lupinus angustifolius L. (narrow-leafed lupin) genome maps and comparative mapping within legumes. Chromosome Res. 24, 355–378 (2016).
Susek, K., Bielski, W. K., Hasterok, R., Naganowska, B. & Wolko, B. A first glimpse of wild lupin karyotype variation as revealed by comparative cytogenetic mapping. Front. Plant Sci. 7, 1152 (2016).
Pan, Q., Mustafa, N. R., Tang, K., Choi, Y. H. & Verpoorte, R. Monoterpenoid indole alkaloids biosynthesis and its regulation in Catharanthus roseus: a literature review from genes to metabolites. Phytochem. Rev. 15, 221–250 (2016).
Nakano, T., Suzuki, K., Fujimura, T. & Shinshi, H. Genome-wide analysis of the ERF gene family in Arabidopsis and Rice. Plant Physiol. 140, 411 (2006).
Udomsom, N. et al. Function of AP2/ERF transcription factors involved in the regulation of specialized metabolism in Ophiorrhiza pumila revealed by transcriptomics and metabolomics. Front. Plant Sci. 7, 1861 (2016).
Nakasugi, K., Crowhurst, R., Bally, J. & Waterhouse, P. Combining transcriptome assemblies from multiple de novo assemblers in the allo-tetraploid plant Nicotiana benthamiana. PLoS One 9, e91776 (2014).
Chen, S., McElroy, J. S. & Dane, F. & Goertzen, L. R. Transcriptome Assembly and Comparison of an Allotetraploid Weed Species, Annual Bluegrass, with its Two Diploid Progenitor Species, Schrad and Kunth. Plant Genome 9, 1 (2016).
Licausi, F., Ohme-Takagi, M. & Perata, P. APETALA2/Ethylene Responsive Factor (AP2/ERF) transcription factors: mediators of stress responses and developmental programs. New Phytol. 199, 639–649 (2013).
Menke, F. L., Champion, A., Kijne, J. W. & Memelink, J. A novel jasmonate- and elicitor-responsive element in the periwinkle secondary metabolite biosynthetic gene Str interacts with a jasmonate- and elicitor-inducible AP2-domain transcription factor, ORCA2. EMBO J. 18, 4455–4463 (1999).
van der Fits, L. & Memelink, J. ORCA3, a jasmonate-responsive transcriptional regulator of plant primary and secondary metabolism. Science 289, 295–297 (2000).
Shoji, T., Kajikawa, M. & Hashimoto, T. Clustered transcription factor genes regulate nicotine biosynthesis in tobacco. Plant Cell 22, 3390–3409 (2010).
Thagun, C. et al. Jasmonate-responsive ERF transcription factors regulate steroidal glycoalkaloid biosynthesis in tomato. Plant Cell Physiol. 57, 961–975 (2016).
Aukerman, M. J. & Sakai, H. Regulation of flowering time and floral organ identity by a MicroRNA and its APETALA2-like target genes. Plant Cell 15, 2730–2741 (2003).
Chen, X. A microRNA as a translational repressor of APETALA2 in Arabidopsis flower development. Science 303, 2022–2025 (2004).
Mathieu, J., Yant, L. J., Mürdter, F., Küttner, F. & Schmid, M. Repression of Flowering by the miR172 Target SMZ. PLoS Biol. 7, e1000148 (2009).
Boycheva, S., Daviet, L., Wolfender, J.-L. & Fitzpatrick, T. B. The rise of operon-like gene clusters in plants. Trends Plant Sci. 19, 447–459 (2014).
Itkin, M. et al. Biosynthesis of antinutritional alkaloids in solanaceous crops is mediated by clustered genes. Science 341, 175–179 (2013).
Winzer, T. et al. A Papaver somniferum 10-gene cluster for synthesis of the anticancer alkaloid noscapine. Science 336, 1704–1708 (2012).
Zhou, G. et al. Construction of an ultra-high density consensus genetic map, and enhancement of the physical map from genome sequencing in Lupinus angustifolius. Theor. Appl. Genet. 131, 209–223 (2018).
Li, Y., Pearl, S. A. & Jackson, S. A. Gene Networks in Plant Biology: Approaches in Reconstruction and Analysis. Trends Plant Sci. 20, 664–675 (2015).
Henry, K. F. & Goldberg, R. B. Using giant scarlet runner bean embryos to uncover regulatory networks controlling suspensor gene activity. Front. Plant Sci. 6, 44 (2015).
Li, Y. & Jackson, S. A. Crowdsourcing the nodulation gene network discovery environment. BMC Bioinformatics 17, 223 (2016).
Kim, E., Hwang, S. & Lee, I. SoyNet: a database of co-functional networks for soybean Glycine max. Nucleic Acids Res. 45, D1082–D1089 (2017).
Li, Y. & Smolke, C. D. Engineering biosynthesis of the anticancer alkaloid noscapine in yeast. Nat. Commun. 7, 12137 (2016).
Martin, M. Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet.journal 17, 1 (2011).
Haas, B. et al. De novo transcript sequence reconstruction from RNA-Seq using the Trinity platform for reference generation and analysis. Nat. Protoc. 8, 1494–1512 (2013).
Luo, R. et al. SOAPdenovo2: an empirically improved memory-efficient short-read de novo assembler. Gigascience 1, 1–18 (2012).
Zerbino, D. R. & Birney, E. Velvet: algorithms for de novo short read assembly using de Bruijn graphs. Genome Res. 18, 821–829 (2008).
Schulz, M. H., Zerbino, D. R., Vingron, M. & Birney, E. Oases: robust de novo RNA-seq assembly across the dynamic range of expression levels. Bioinformatics 28, 1086–1092 (2012).
Crusoe, M. R. et al. The khmer software package: enabling efficient nucleotide sequence analysis. F1000Res 4 (2015).
Simão, F. A., Waterhouse, R. M., Ioannidis, P., Kriventseva, E. V. & Zdobnov, E. M. BUSCO: assessing genome assembly and annotation completeness with single-copy orthologs. Bioinformatics 31, 3210–3212 (2015).
Conesa, A. & Gotz, S. Blast2GO: A comprehensive suite for functional analysis in plant genomics. Int. J. Plant Genomics 2008, 619832 (2008).
Hatje, K. et al. Cross-species protein sequence and gene structure prediction with fine-tuned Webscipio 2.0 and Scipio. BMC Res. Notes 4, 265 (2011).
Li, B. & Dewey, C. N. RSEM: accurate transcript quantification from RNA-Seq data with or without a reference genome. BMC Bioinformatics 12, 323 (2011).
Manly, K. F., Cudmore, R. H. Jr. & Meer, J. M. Map Manager QTX, cross-platform software for genetic mapping. Mamm. Genome 12, 930–932 (2001).
Voorrips, R. E. MapChart: software for the graphical presentation of linkage maps and QTLs. J. Hered. 93, 77–78 (2002).
Pfaffl, M. W. A new mathematical model for relative quantification in real-time RT-PCR. Nucleic Acids Res. 29, e45 (2001).
Cullis, B. R., Smith, A. B. & Coombes, N. E. On the design of early generation variety trials with correlated data. J. Agr. Biol. Envir. St. 11, 381 (2006).
Genstat for Windows, 18th Edition. Hemel Hempstead, UK. (VSN International 2015).
Wang, S., Basten, C. J. & Zeng, Z. B. Windows QTL Cartographer 2.5. Department of Statistics, North Carolina State University, Raleigh, NC. (http://statgen.ncsu.edu/qtlcart/WQTLCart.htm). (2012).
Xu, Y. Molecular Plant Breeding. (CAB International, UK 2010).
Zheng, Y. et al. iTAK: a program for genome-wide prediction and classification of plant transcription factors, transcriptional regulators, and protein kinases. Mol. Plant 5, 1667–1670 (2016).
Enright, A. J., Van Dongen, S. & Ouzounis, C. A. An efficient algorithm for large-scale detection of protein families. Nucleic Acids Res. 30, 1575–1584 (2002).
Koczyk, G., Dawidziuk, A. & Popiel, D. The distant siblings -a phylogenomic roadmap illuminates the origins of extant diversity in fungal aromatic polyketide biosynthesis. Genome Biol. Evol. 7, 3132–3154 (2015).
Katoh, K. & Standley, D. M. MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Mol. Biol. Evol. 30, 772–780 (2013).
Minh, B. Q., Nguyen, M. A. & von Haeseler, A. Ultrafast approximation for phylogenetic bootstrap. Mol. Biol. Evol. 30, 1188–1195 (2013).
Acknowledgements
We thank Poznan Plant Breeders Ltd. for conducting field experiments. MK gratefully acknowledges Dr. Karolina Susek (Institute of Plant Genetics, Polish Academy of Sciences, Poznan) and Prof. Frederic Stoddard (University of Helsinki) for their constructive feedback. This work was supported by the Polish Ministry of Agriculture and Rural Development within the Multiannual Program (RM-111-222-15). Computations for de novo transcriptome assembly and analysis were partially carried out in Poznan Supercomputing and Networking Centre.
Author information
Authors and Affiliations
Contributions
M.K. and G.K. conceived the study and participated in QA candidate genes selection. M.K. performed genetic and QTL mapping and drafted the manuscript. M.K. and K.K. developed transcriptome-derived SNP markers and participated in the assessment of candidate reference genes expression stability. P.K. performed statistical analyses. G.K. performed transcriptome de-novo assembly, DE and phylogenetic analysis. K.C., K.K. and O.F. performed qRT-PCR analyses. W.S. contributed to the concept of the study and acquired of funding. J.K. performed transcriptome sequencing. J.P. carried out cDNA libraries construction. P.W. carried out gas chromatography analyses. M.K., G.K. and P.K. participated in production of figures and tables. All authors revised the manuscript critically and approved its final version.
Corresponding author
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Kroc, M., Koczyk, G., Kamel, K.A. et al. Transcriptome-derived investigation of biosynthesis of quinolizidine alkaloids in narrow-leafed lupin (Lupinus angustifolius L.) highlights candidate genes linked to iucundus locus. Sci Rep 9, 2231 (2019). https://doi.org/10.1038/s41598-018-37701-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-018-37701-5
This article is cited by
-
Integrative analysis of the steroidal alkaloids distribution and biosynthesis of bulbs Fritillariae Cirrhosae through metabolome and transcriptome analyses
BMC Genomics (2022)
-
Development and application of a virus-induced gene silencing protocol for the study of gene function in narrow-leafed lupin
Plant Methods (2021)
-
Integrated transcriptome and small RNA sequencing analyses reveal a drought stress response network in Sophora tonkinensis
BMC Plant Biology (2021)
-
Phenotypic characterisation and linkage mapping of domestication syndrome traits in yellow lupin (Lupinus luteus L.)
Theoretical and Applied Genetics (2020)
-
Development of gene-based molecular markers tagging low alkaloid pauper locus in white lupin (Lupinus albus L.)
Journal of Applied Genetics (2019)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.