Abstract
Autism spectrum disorder (ASD) is a highly prevalent neurodevelopmental disorder, and the exact causal mechanism is unknown. Dysregulated allele-specific expression (ASE) has been identified in persons with ASD; however, a comprehensive analysis of ASE has not been conducted in a family quartet with ASD. To fill this gap, we analyzed ASE using genomic DNA from parent and offspring and RNA from offspring’s postmortem prefrontal cortex (PFC); one of the two offspring had been diagnosed with ASD. DNA- and RNA-sequencing revealed distinct ASE patterns from the PFC of both offspring. However, only the PFC of the offspring with ASD exhibited a mono-to-biallelic switch for LRP2BP and ZNF407. We also identified a novel site of RNA-editing in KMT2C in addition to new monoallelically-expressed genes and miRNAs. Our results demonstrate the prevalence of ASE in human PFC and ASE abnormalities in the PFC of a person with ASD. Taken together, these findings may provide mechanistic insights into the pathogenesis of ASD.
Similar content being viewed by others
Introduction
Autism spectrum disorder (ASD) is a heritable neurodevelopmental disorder characterized by social difficulties, communication challenges, and repetitive behaviors1. ASD has been identified in 1 out of 68 children2, which reveals its high prevalence and indicates its importance as a public health issue. There is no definitive biomarker for ASD and no widely accepted treatment. Although medications can provide an improvement in behaviors for persons with ASD, they cannot reliably ameliorate all of the core symptoms of this disorder3,4. This limitation of therapeutic efficacy is due to the heterogeneous and multifactorial causes of ASD, which include genetic components, environmental insults, and gene-environment interactions5,6. Altered gene expression in the brain has been consistently identified in persons with ASD through genome-wide analysis7,8,9,10. In addition, phenotypic and genetic variations are entangled, which further hinders deciphering the etiology of ASD11,12. Although a single ultimate neuropathological feature in the brains of individuals with ASD may be impossible to define, we now know that common variant of small effect and rare de novo variants of large effect can combine to influence the risk for ASD. Apart from de novo mutations, copy number variations13,14,15, and aberrant microRNA profiles16, epigenetic mechanisms such as variations in DNA methylation on differential gene expression have been proposed to play a pivotal role in ASD17,18,19. Although Fragile X syndrome is the most well-known single-gene disorder, it only accounts for approximately 5% of all ASD cases20; the cause of most cases of ASD are unclear. Moreover, the challenge of investigating the causes is further complicated by the complex nature of ASD.
Allele-specific expression (ASE) is an unequal expression of alleles. One extreme example is mono-allelic expression (MAE), in which one allele is expressed, while the other is inactive. MAE consists of several different mechanisms, such as genomic imprinting21, X chromosome-inactivation22, expression quantitative trait loci (eQTL)23,24, and autosomal random MAE25. Genomic imprinting is essential for neurodevelopment and viability of the organism26. Mono-allelically expressed genes play important roles during development, and they are predisposed to loss of their function through mutations, thus contributing to diseases27,28. Indeed, random or stochastic monoallelically expressed genes are enriched for candidate genes for neurodevelopmental disorders29. Furthermore, MAE affects epigenetic processes in brains of individuals with ASD30 and dysregulated genomic imprinting has been identified in persons with ASD31,32.
The quality of genetic information can be improved by analyzing complete genome sequences from family members, which was demonstrated by the pioneering work of Roach et al. (2010) using a family quartet (two siblings and their parents)33. Moreover, information regarding single nucleotide polymorphisms (SNPs) from the parent is critical for parent-of-origin allele-specific expression analysis, although haplotype phasing in an outbred human population is complicated34. Therefore, it is necessary to acquire samples from a family quartet to determine the parental source of transcripts in offspring. Despite the relevance of MAE to ASD, comprehensive analysis of MAE in a family quartet with ASD has not been conducted due to limited access to samples.
To address this critical knowledge gap, we investigated the role of ASE in the pathogenesis of ASD using special human parent-child quartet samples. Genomic DNA from parent and offspring and RNA from offspring’s postmortem prefrontal cortex (PFC) of the brain were analyzed. One offspring had been diagnosed with ASD. We observed distinct ASE patterns of genes and microRNAs (miRNAs) in the PFC of both offspring. Importantly, we found a mono-to-biallelic switch for LRP2BP (LRP2 binding protein) and ZNF407 (Zinc finger protein 407) in the offspring diagnosed with ASD. We also found a novel site of RNA editing in KMT2C (Lysine (K) methyltransferase 2 C); a novel development stage- and brain-specific maternally-expressed gene, DUSP22 (Dual specificity phosphatase 22); and a novel development stage-specific paternally-expressed miRNA, miR-335, in the PFC of both offspring. KMT2C, DUSP22 and miR-335 are autism susceptibility genes and miRNAs. Our results provide further evidence that ASE could contribute to ASD.
Results
Quality and quantity of DNA and RNA of a parent-child quartet with ASD met the requirements for deep sequencing
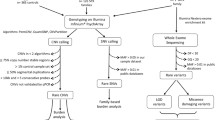
We analyzed ASE on a genome-wide scale and investigated whether dysregulated ASE occurs in persons with ASD using samples from a family quartet with ASD consisting of genomic DNA from the parent and offspring and RNA extracted from postmortem PFC of the offspring (Fig. 1a). Sequencing of parental genomic DNA provided information regarding single nucleotide polymorphisms (SNPs), which is essential for determining the parental source of offspring’s transcripts. In this family quartet, both offspring were female, and one had been diagnosed with ASD (Fig. 1a). Supplementary Fig. 1 provides a more detailed pedigree of this family, which shows epilepsy and deafness co-occurred in both the affected and unaffected offspring in addition to other familial health conditions. Next, we performed DNA sequencing of parents and offspring; RNA-sequencing and follow-up transcriptomic and ASE analysis was performed on the offspring. The detailed demographic and deep sequencing information is presented in Supplementary Table 1. The quantity and quality of genomic DNA from parents and offspring met the requirements for DNA-sequencing (Fig. S2a); in addition, the quantity and quality of the offspring’s RNA met the requirements for RNA-sequencing (Fig. S2b). Total read number and mappability of the offspring’s RNA-Seq data met the requirements for further statistical analysis (Supplementary Table 1). Taken together, the DNA and RNA from the offspring qualified for deep sequencing.
Differential gene and miRNA expression patterns in the postmortem prefrontal cortex (PFC) of a person without and with ASD. (a) Schematic diagram of the family tree from a family quartet with ASD. Square indicates male and circle indicates female. Black solid circle indicates the offspring with ASD; open circle indicates the offspring without ASD. (b) Heatmap analyses showing gene expression profiles for the offspring without and with ASD. (c) Gene ontology (GO) enrichment analysis was performed for affected genes from the offspring with ASD. (d) The top genes that were up-regulated (top) and down-regulated (bottom) were validated by qPCR; expression levels of genes for the offspring with ASD (black bars) compared with levels for the offspring without ASD (open bars). (e) Heatmap analyses showing miRNA expression profiles for the offspring without and with ASD. (f) Gene ontology (GO) enrichment analysis was performed for affected miRNAs from the offspring with ASD. (g) The top miRNAs that were up-regulated (top) and down-regulated (bottom) were validated by qPCR; expression levels of miRNA in the offspring with ASD (black bars) compared with levels in the offspring without ASD (open bars). NA, the offspring without ASD; A, the offspring with ASD. The data points above the red dashed line represent P values less than 0.05. Euclidean distance was used to generate the heatmap plots.
Gene and miRNA expression are altered in the PFC of the offspring with ASD
To determine whether dysregulated gene expression was present in the postmortem PFC of offspring with ASD, we compared gene expression patterns of the offspring with and without ASD on a genome-wide scale. We focused on genes with expression levels of more than 0.3 FPKM in the offspring without ASD and a fold change between the offspring with and without ASD larger than 0.5 or less than −0.5, on a base-10 logarithmic scale (Fig. 1b and Supplementary Table 2). Rather than use the typical two-fold change, we used a fold change of 3.16 (=10^(0.5)), which is a more stringent threshold. When compared to the offspring without ASD, we detected 2293 up-regulated and 5980 down-regulated gene isoforms in the PFC of the offspring with ASD. We then performed gene ontology (GO) enrichment analysis on the affected gene isoforms to further investigate which functions, processes, and components were affected. Focusing on gene isoforms involved in brain-related functions, we observed that genes related to the development of dendrites, axons and the olfactory bulb were affected (Fig. 1c and Supplementary Table 3). None of the GO terms survived after false discovery rate (FDR) correction. The functions of the genes related to olfactory bulb development (ID2, AGTPBP1 and SEMA3A) are related to overall brain development35,36,37. If we applied more stringent criteria (log2 (ASD/non-ASD > 2 or −2 and q value < 0.05), we only detected 67 genes with altered expression in the offspring with ASD (Supplementary Table 4). Due to the small size of the genes, GO analysis could not be further pursued. To validate the accuracy of our heatmap results, we confirmed the expression levels of the top genes that were up-regulated (such as RNF220, SEMA3E, and SEPT5) and down-regulated (such as HLA-A, NPAS4, and TNFRSF8) with qPCR (Fig. 1d). These genes were chosen based on the availability of reliable primers, high expression levels, and a difference in expression level between ASD and non-ASD samples. To examine if the altered genes could be observed in other ASD cohorts, we compared our altered genes with RNA-Seq data from Dr. Weinberger’s group7 for the dorsolateral prefrontal cortex from three persons with ASD and three matched controls. We observed that expression of 2406 out of 8273 genes was also altered in this ASD cohort (Fig. S3a and Supplementary Table 5). For analysis of Dr. Weinberger’s RNA-Seq data, we used the same criteria of altered gene expression as we used for our ASD cohort. To investigate if the altered genes could be observed in a larger ASD cohort, we compared our altered genes with RNA-Seq data from a study by Dr. Geschwind’s group10 of the cortex from persons with ASD and matched controls. We found that expression of 387 out of 1087 genes was also altered in this ASD cohort (Supplementary Table 6).
Small RNAs, such as miRNAs, affect gene expression, but whether dysregulated miRNAs contribute to the etiology of ASD has not been well studied. To address this question, we compared the expression levels of miRNAs from the PFC of the two offspring on a genome-wide scale. We focused on miRNAs with a fold change between the offspring with and without ASD larger than 0.5 or less than −0.5, on a base-10 logarithmic scale, which resulted in a threshold of fold change of 3.16 (=10^(0.5)). When PFC miRNAs from the offspring with ASD were compared with those of the offspring without ASD, we found 105 up-regulated and 125 down-regulated miRNAs in the offspring diagnosed with ASD (Fig. 1e and Supplementary Table 7). We then performed GO analysis on the affected miRNAs (Fig. 1f and Supplementary Table 8) and examined genes targeted by altered miRNAs (Supplementary Table 9). None of GO terms survived after FDR correction. To validate the accuracy of our heatmap results, we performed qPCR to confirm the expression levels of the top up-regulated miRNAs (such as miR365a-3p, miR-2277-3p, and miR-184-3p) and down-regulated miRNAs (such as miR-21-5p, miR-136-3p, and miR-127-3p) (Fig. 1g). These miRNAs were chosen based on the availability of reliable primers, high expression levels, and a difference in expression level between ASD and non-ASD samples. To determine whether those altered miRNAs could be detected in other ASD cohorts, we compared our altered miRNAs with RNA-Seq data from Dr. Weinberger’s group7 for the dorsolateral prefrontal cortex of three persons with ASD and the three matched controls, using same criteria for analysis as we used in our ASD cohort. We observed that expression of 12 out of 210 miRNAs was also altered in this ASD cohort (Fig. S3b and Supplementary Table 5). To investigate if the altered miRNAs could be detected in a larger ASD cohort, we compared our altered miRNAs with RNA-Seq data from a study by Dr. Geschwind’s group16 of the cortex from persons with ASD and matched controls. We found that expression of 1 out of 58 miRNAs was also altered in this ASD cohort (shaded genes, Supplementary Table 6). Taken together, our results showed the PFC of the offspring with ASD contained genes and miRNAs with altered expression.
Autism susceptibility genes were altered in the PFC of the offspring with ASD
To investigate whether autism susceptibility genes were preferentially altered, we compared expression levels of autism susceptibility genes in the postmortem PFC of the two offspring. The source of autism susceptibility genes was from the Simons Foundation Autism Research Initiative (SFARI, September 2016; Supplementary Table 10). We focused on genes with expression levels of more than 0.3 FPKM in the offspring without ASD and a fold change between the offspring with and without ASD larger than 0.5 or less than −0.5, on a base-10 logarithmic scale and as a result, the threshold for fold change used was 3.16 (=10^(0.5)). Compared with the offspring without ASD, the PFC of the offspring with ASD exhibited 142 up-regulated and 312 down-regulated autism susceptibility gene isoforms (Fig. 2a and Supplementary Table 11). We performed a hypergeometric test to assess whether the above overlaps were more than one would expect by chance and found significant overlaps (p < 0.001). To validate the heatmap results regarding differences in regulation of these gene isoforms in the offspring with ASD, we compared the offspring’s top up-regulated autism susceptibility genes (such as AGAP1, EFR3A and KAT6A, Fig. 2b) and down-regulated autism susceptibility genes (such as NRXN2, SERPINE1, and BBS4, Fig. 2c). These genes were chosen based on the availability of reliable primers, high expression levels, and a difference in expression level between ASD and non-ASD samples. To determine whether the altered ASD susceptibility genes could be observed in other ASD cohorts, we compared our altered ASD susceptibility genes with RNA-Seq data from Dr. Weinberger’s group7 for the dorsolateral prefrontal cortex of three persons with ASD and three matched controls. We observed altered expression in 51 out of 455 genes in this ASD cohort (Fig. S3c and Supplementary Table 5). For analysis of the data in the cohort from Dr. Weinberger’s data, we used the same criteria for the ASD susceptibility genes we used in our ASD cohort. To investigate if the altered genes could be observed in a larger ASD cohort, we compared our altered genes with RNA-Seq data from a study by Dr. Geschwind’s group10 of the cortex from persons with ASD and matched controls. We found that expression of 29 out of 1087 genes was also altered in this ASD cohort (shaded genes, Supplementary Table 6).
Distinct expression patterns of autism susceptibility genes and miRNAs from postmortem prefrontal cortex (PFC) of the offspring diagnosed with ASD. (a) Heatmap analyses showing different gene expression profiles of autism susceptibility genes from the PFC of the offspring without and with ASD. The top genes that were up-regulated (b) and down-regulated (c) were validated by qPCR; expression levels in the offspring with ASD (black bars) compared with levels in the offspring without ASD (open bars). (d) Autism susceptibility miRNAs were determined with qPCR for the offspring without ASD (open bars) and the offspring with ASD (black bars). NA, the offspring without ASD; A, the offspring with ASD. Euclidean distance was used to generate the heatmap plots.
When compared with the offspring without ASD, the following autism susceptibility miRNAs16,38 were altered in the offspring with ASD: miR-619-5p, miR-23a-3p, miR-103a-3p, miR-106b-5p, miR-143a-3p, miR-146a-5p, and miR-204-3p (Fig. 2d and Supplementary Table 12). A hypergeometric test showed the above overlaps were not significant (p = 0.92). Since miRNAs regulate gene expression, we also examined genes targeted by the affected miRNAs (Supplementary Table 12). We then performed gene ontology (GO) enrichment analysis on the affected miRNA target genes to further investigate which functions and pathways were affected. However, we did not observe any significant results. Our data showed altered expression of autism susceptibility genes from SFARI in the PFC from the offspring with ASD.
Allele-specific gene expression was altered in the postmortem PFC of the offspring with ASD
To examine allele-specific gene expression and to determine whether dysregulated allele-specific gene expression occurred in the offspring with ASD, the parents’ and offspring’s genomic DNA was analyzed with DNA-Seq, and the offspring’s postmortem PFC RNA was analyzed with RNA-Seq followed by ASE analysis. First, we observed a distinct allele-specific gene expression pattern, which contained a diagonal line with one paternally-dominant cohort and one maternally-dominant cohort in both offspring (Fig. 3a and Supplementary Table 13). Genes within the diagonal line indicate both of their alleles were expressed equally, which represented the majority of genes; genes above or below the diagonal line indicate genes that were expressed predominantly from either maternal or paternal allele. This pattern has been observed consistently in different brain regions related to the mouse visual system39 as well as in cell types of the mouse visual cortex40. In contrast, when we compared the ASE patterns of all of SFARI’s autism susceptibility genes, several genes differed between the offspring with and without ASD (Fig. 3b and Supplementary Table 14). A hypergeometric test demonstrated these differences in genes between siblings were not by chance (p < 0.001). We validated allele-specific expression for our candidate genes with Sanger sequencing and determined LRP2BP and ZNF407 were both mono-allelically expressed in the offspring without ASD, but bi-allelically expressed in the offspring with ASD (Fig. 3c,d, top). LRP2BP was also monoallelically expressed in other non-ASD brain samples (Fig. S4a). This mono-to-biallelic switch reflects their expression levels (Fig. 3c,d, bottom). Because there is an overlap between autism susceptibility genes from SFARI (Supplementary Table 10, September 2016) and known human imprinted genes from the Geneimprint website (Supplementary Table 15, September 2016) (Fig. 3e), we validated seven of the 19 overlapped genes with Sanger sequencing (Fig. 3f,g). Due to the lack of availability of SNPs and low gene expression levels, the remaining overlapped genes could not be validated. For genes that could be validated, Sanger sequencing showed ATP10A, CTNNA3, DLGAP2, GABRB3, and HTR2A were not imprinted in either of the offspring (Fig. 3f), whereas MAGEL2 and SNRPN were imprinted in both (Fig. 3g). The imprinting status of ATP10A, DLGAP2, and HTR2A was further confirmed in other brain samples (Fig. S4b). Taken together, our data suggest that allele-specific gene expression occurs in human PFC and dysregulated allele-specific gene expression occurred in the PFC of the offspring with ASD.
Distinct patterns of allele-specific gene expression in postmortem prefrontal cortex (PFC) of the offspring without and with ASD. (a) Parental expression patterns from the PFC of the offspring without ASD (top) and with ASD (bottom) analyzed on a genome-wide scale with RNA-Seq. (b) Heatmap analyses showing allele-specific gene expression profiles in the offspring without and with ASD. (c) Sanger sequencing validated the allele-specific expression of LRP2BP from the two offspring. Expression levels of LRP2BP from the PFC of the offspring without ASD (open bar) and with ASD (black bar) were quantified by qPCR. (d) Sanger sequencing validated the allele-specific expression of ZNF407 from the two offspring. Expression levels of ZNF407 from the PFC of the offspring without ASD (open bar) and with ASD (black bar) were quantified by qPCR. The offspring with and without ASD each had unique SNPs for ZNF407: G/A for the offspring without ASD and T/A for the offspring with ASD. (e) Venn diagram showing overlapped Simons Foundation Autism Research Initiative (SFARI) genes with known human imprinted genes. (f,g) The imprinting status of seven of the 19 overlapped genes was verified by Sanger sequencing. NA, the offspring without ASD; A, the offspring with ASD; B, bi-allelic; M, maternal; p, paternal. *Gene could not be validated due to lack of availability of SNPs or low gene expression levels. SNP information was shown as (paternal allele/maternal allele). Euclidean distance was used to generate the heatmap plots.
Allele-specific miRNA expression was altered in the PFC of the offspring with ASD
To determine whether allele-specific expression of miRNAs occurs in the PFC of humans and whether the expression is dysregulated in persons with ASD, we profiled allele-specific miRNA expression in the PFC of the offspring with and without ASD. We observed a pattern of ASE for miRNAs, which showed a diagonal line with a paternally-dominant cohort and a maternally-dominantly cohort (Fig. 4a), similar to that seen for genes (Fig. 3a). In addition, this pattern has been observed consistently in different brain regions related to the mouse visual system39. Heatmap clustering compared miRNAs from the two offspring whose fold change on a base-10 logarithmic scale was larger than 0.5 or less than −0.5 (Fig. 4b and Supplementary Table 16), and as a result, the threshold for fold change used was the 3.16 (=10^(0.5)). We validated the allele-specific miRNA expression with Sanger sequencing (Fig. 4c,d). There was no difference in allele-specific miRNA expression between offspring without and with ASD. However, we identified maternally-expressed miRNAs (miR-299 and miR-654) in both offspring (Fig. 4d). In summary, our data show distinct patterns of allele-specific miRNA expression in the PFC of both offspring.
Distinct patterns of allele-specific miRNAs in the postmortem prefrontal cortex (PFC) of offspring without and with ASD. (a) Parental expression pattern of miRNAs was analyzed on a genome-wide scale with RNA-Seq in the PFC of both offspring. (b) Heatmap analyses of allele-specific expression profiles for miRNAs with fold changes of >0.5 or <−0.5 on a base-10 logarithmic scale between the offspring without and with ASD. (c,d) Allele-specific expression was validated for the miRNAs in (b) from the two offspring by Sanger sequencing. SNP information is shown as (paternal allele/maternal allele). Euclidean distance was used to generate the heatmap plots.
Genomic map of parent-of-origin-specific gene and miRNA expression in the human PFC
To map parent-of-origin-specific gene and miRNA expression on a genome-wide scale in human PFC, we compared our data obtained from the PFC of the offspring without ASD for allele-specific gene and miRNA expression with known human imprinted genes reported by the Geneimprint website (Supplementary Table 15, September 2016) and previous literature41. RNA-Seq and follow-up ASE analysis confirmed the presence of 60% of known paternally expressed genes and 60% of known maternally expressed genes (Supplementary Fig. 5 and Table 17). We mapped the confirmed imprinted genes and miRNAs into 23 human chromosomes (Fig. 5). This mapping identifies the parent-of-origin-specific genes and miRNAs expressed on a genome-wide scale in the human PFC. Canonical genomic imprinting involves silencing of the maternal and paternal allele. In contrast, noncanonical genomic imprinting involves maternal or paternal allele expression biases42. We identified a noncanonical imprinted gene, NOS1 (Fig. 6a) in the PFC of both offspring.
Chromosome map of parent-of-origin-specific genes and miRNAs in the PFC of the offspring without ASD. Examination of 23 chromosomes for the presence of paternally- (blue) and maternally- (red) expressed imprinted genes and miRNAs in the PFC from the offspring without ASD. Dotted line separates the p and q arm of each chromosome.
Identification of noncanonical imprinting in the PFC of the offspring without and with ASD and editing of KMT2C transcript in human PFC. (a) Sanger sequencing was used to analyze noncanonical imprinting of NOS1 in the PFC of the offspring without and with ASD. “M*” stands for maternally-biased expression. (b) Schematic diagram of the genomic locus of human KMT2C. Arrows indicate the direction of transcription. (c) Sanger sequencing was performed to analyze RNA editing for KMT2C in the PFC of the offspring without and with ASD, fetal PFC, and blood from parents of the fetus. (d) Schematic diagram of the genomic locus of mouse Kmt2c. Arrows indicate the direction of transcription. (e) Sanger sequencing was performed to analyze RNA editing for Kmt2c in PFC and blood from postnatal day 28 (P28) mice and PFC from embryonic day 15.5 (E15.5) mice. SNP information is shown as (paternal allele/maternal allele).
Novel transcriptional processes of autism susceptibility genes were identified in human PFC
Validation of allele-specific expression in the PFC of both offspring with Sanger sequencing identified three novel transcriptional processes of autism susceptibility genes. First, we identified a novel human-specific site of RNA editing in KMT2C (Fig. 6b). When we analyzed RNA editing in human adult and fetal PFC (Fig. 6c), we observed T-to-C RNA editing of KMT2C sense transcript, which was in contrast to A-to-G RNA editing for KMT2C antisense, which could cause a phenylalanine-to-leucine change at the protein level (p.F291L). This pattern of RNA editing was not seen when we analyzed PFC from postnatal day 28 (P28) and embryonic day 15.5 (E15.5) mouse (Fig. 6d,e, respectively), suggesting this RNA editing in KMT2C is human-specific. These differences in RNA editing patterns between human and mouse tissue were also seen when we examined human and mouse blood (Fig. 6c,e). Second, we identified a development stage- and brain-specific maternally-expressed gene, DUSP22 (Fig. 7). We found that DUSP22 was maternally expressed in adult PFC but bi-allelically expressed in fetal PFC and adult blood (Fig. 7b). The imprinting status of DUSP22 has been validated in other fetal tissue (Fig. S4b). Because we have been unable to identify the exonic SNPs in mouse Dusp22, we have not examined the ASE pattern of Dusp22 in mouse PFC and blood. Finally, we identified a development stage-specific paternally-expressed miRNA, miR-335 (Fig. 8), which was paternally-expressed in the adult PFC but paternally-biased in the fetal PFC. Taken together, our results add new information about the dynamic transcriptomic processes in the brain.
DUSP22 is maternally expressed in the PFC of the offspring without and with ASD. (a) Schematic diagram of the genomic locus of DUSP22. Arrows indicate the direction of transcription. (b) Sanger sequencing was performed to analyze allele-specific DUSP22 expression in the PFC of the offspring without and with ASD, fetal PFC, and blood from the parents of the fetus. “M” stands for maternal expression. “B” stands for biallelic expression. SNP information is shown as (paternal allele/maternal allele).
miR-335 is paternally expressed in the PFC of the offspring without and with ASD. Sanger sequencing was performed to analyze allele-specific miR-335 expression in the PFC of the offspring without and with ASD, fetal PFC, and blood from the parents of the fetus. “P” stands for paternal expression. “B” stands for biallelic expression. “P*” stands for paternally-biased expression. SNP information is shown as (paternal allele/maternal allele).
Discussion
Our genome-wide analysis provides new information regarding expression of allele-specific genes and miRNAs in human PFC and persons with ASD. Analysis of the PFC revealed a distinct allele-specific expression pattern, which contained a diagonal line with a maternally dominant cohort and a paternally dominant cohort. Moreover, we identified novel allele-specific genes such as DUSP22 and miRNAs such as miR-335 in both the offspring with and without ASD, as well as a mono-to-biallelic switch for LRP2BP and ZNF407 in the offspring diagnosed with ASD. We also identified a novel human-specific site of RNA editing in KMT2C. Importantly, our study results indicate that a genome-wide ASE map could provide a powerful model for understanding neuropsychiatric disorders through the study of key features of dynamic allele-specific gene and miRNA expression in human PFC, supplemented with the study of the roles of dysregulated allele-specific genes and miRNAs in ASD.
Dual specificity phosphatase 22 (DUSP22) is an enzyme, which activates the JNK signaling pathway43. JNK activation has been shown to play an essential role in organogenesis during mouse development by regulating cell survival, apoptosis, and proliferation44. The physiological role of DUSP22 in the brain is unclear, and a rare DUSP22 deletion was found in a patient with autism and mild intellectual disability45. DUSP22 in the PFC from the offspring without and with ASD consistently showed maternal expression. Our finding further extends a previous finding showing DUSP22 is a monoallelically-expressed gene46. Interestingly, DUSP22 from fetal PFC showed bi-allelical expression, therefore it would be of interest to investigate how a biallelic-to-monoallelic switch is regulated for DUSP22 during prefrontal development. Moreover, LRP2BP and ZNF407 in the PFC from the offspring with ASD consistently showed a mono-to-biallelic switch. Importantly, dysregulation of LRP2BP47 and ZNF40748 has been identified in persons with ASD. However, the roles of LRP2BP and ZNF407 in the brain have not been identified and the function of LRP2BP protein is still unknown. In contrast, the function of ZNF407 protein has been shown to regulate glucose homeostasis49. Therefore, because glucose homeostasis is critical for normal brain function, dysregulation of ZNF407 could affect brain development and function. It would be of interest not only to investigate how a mono-to-biallelic switch is regulated for LRP2BP and ZNF407, which could provide insight into their roles in ASD specifically, but also to study the roles of LRP2BP and ZNF407 more generally during neurodevelopment. The PFC from the offspring without and with ASD consistently showed paternal expression of miR-335, which further extends a previous finding showing mouse miR-335 is paternally-expressed50.
Lysine methyltransferase 2 C (KMT2C) has histone methylation activity and is a transcriptional coactivator. In our studies, we found that T-to-C RNA editing occurred in the KMT2C sense transcript. This editing causes a missense mutation, which converts phenylalanine to leucine at amino acid 291 of KMT2C. Phenylalanine and leucine are both neutral and non-polar amino acids, therefore, this change should have a minor effect on the protein structure of KMT2C. Indeed, when we used SIFT (http://sift.jcvi.org/) to predict whether such amino acid substitution affects protein function, the result predicted that this change is a tolerated substitution. It would be of interest to investigate the physiological significance of the T-to-C editing event in the sense transcript and an A-to-G editing event in the antisense transcript.
It has been reported that parent-of-origin-specific expression is also brain-region specific51. Our results demonstrate expression of parent-of-origin-specific genes and miRNAs in human PFC, which has not previously been reported. Our transcriptomic and ASE analysis of the PFC of the offspring with and without ASD of a family quartet identified dysregulated gene and miRNA expression, and ASE in the offspring with ASD, which could identify a novel set of autism susceptibility genes and miRNAs. It will be important to identify the regulatory mechanisms for dysregulated transcriptomic and allele-specific expression of these genes and miRNAs identified in the offspring with ASD. To this end, it will be essential to determine if stringent identification of allele-specific genes together with the systematic screening of allele-specific chromatin and DNA modifications could unravel markers for further mechanistic validation. It would also be interesting to further investigate whether cis-transcribed non-coding RNAs, intra-nuclear allelic positioning, and chromosomal interactions are associated with the allele-specific expression.
Several methods are available for seeking the allele-specific expressed genetic locus on a genome-wide scale, which include in silico prediction pipelines52, SNP genotyping arrays53, gene expression arrays54 and transcriptome sequencing approach55. The transcriptomic approach is based on detecting allelic expression with RNA-Seq reads that map heterozygous SNPs, where the identity of the base is used to distinguish allelic origin and a reciprocal cross is used to discriminate parent-of-origin specificity from strain-specific or random biases. The transcriptomic approach for allele-specific gene expression is a paradigm shift in comparison to previous methods. However, recent literature has indicated a high FDR could explain the majority of novel imprinted genes in an RNA-Seq approach resulting from the contribution of several factors. First, systematic errors in technical and biological replicates include priming, fragmentation, and PCR biases that arise during library construction and sequencing. One can adopt a mock reciprocal cross as a negative control to set a false discovery cutoff for systemic errors. Second, the strain-specific effect could be due to cis-eQTL and tissue-specific effect could be due to trans-eQTL. Third, in comparison to the inbred mouse, the human is an outbred species and the complexity of haplotype phasing is much higher, which might lead to wrong read alignment (mapping bias). Since we cannot acquire a large sample size of SNPs calling data from the same population as the sampled genome, the only way to perform haplotype calling is to make inference via an established reference haploid genome from the same population. If the reference haploid genome distinctly differs from the sampled genome, then FDR inevitably rises. Recently, a Bayesian approach for analysis of ASE using a personal diploid genome as a reference sequence has been established and shows less biased alignments and more consistent ASE56. Fourth, small sample size and confounding underlying diseases (e.g. epilepsy, congenital deafness, or intellectual disability) may contribute to false detection. Fifth, in our study, we only had tissue from the prefrontal cortex of the offspring. However, other brain regions, such as the cerebellum, are also reported to be related to the anatomical and neuropathological causes of ASD57 and have different miRNA expression patterns16 in comparison with the prefrontal area.
Bipolar disorder was also present in the offspring with ASD. Because the age of onset for autism (2 to 3 years of age) is much earlier than that of bipolar disorder (25 years) and the offspring with ASD died at age 29 years, the impact of autism should be higher than that of bipolar disorder. In addition, many psychiatric disorders share genetic roots58. For example, there is an overlap between rare genetic variations linked to bipolar disorder and those implicated in autism59. Moreover, 83% of persons with ASD have been shown to have been diagnosed with at least one comorbid non-ASD developmental disorder60, therefore, it would be difficult to acquire cases of persons with ASD only. The results obtained here are more likely due to ASD rather than bipolar disorder.
Due to tissue availability, we sampled different regions of the prefrontal cortex (BA10 for the offspring without ASD and BA8 for the offspring with ASD) to conduct the analysis. The differential expression analysis may be confounded by the tissue specificity of gene expression in BA8 and BA10. To explore the impact of this potential confounding factor and verify the similarity of these two regions, we first compared the expression patterns of different regions of the central nervous system with principle component analysis (PCA) (Fig. S6a,b). PCA demonstrated similarities between BA8 and BA10 in comparison with the cerebellum and frontal cortex by PC2. It also showed a strong difference of BA8/10 from the amygdala, caudate nucleus, and spinal cord by PC1. Heatmap analysis again showed similarities between BA8 and BA10 in comparison to other regions of the central nervous system (Fig. S6c). To further confirm whether the differential gene expression in Figs 1b and 2a was due to brain region, we used heatmap and volcano analysis to examine those differentially expressed genes and observed that less than 7% of those genes were significantly affected (Fig. S6d to g and Supplementary Table 18). These findings show the two regions are similar, which supports our use of BA8 and BA10 for the comparison of gene expression.
In spite of these potential limitations and confounding factors, our results provide valuable clues for identifying biomarkers and biological signatures of ASD, and increase our understanding of potential underlying mechanisms that contribute to the pathogenesis of the disorder. Our findings not only advance our knowledge of allele-specific gene and miRNA expression in ASD, but also provide the first genomic map for allele-specific gene and miRNA expression in human PFC. These results could also provide clues to the evolutionary development of allele-specific expression. In addition, therapeutic targets and strategies for brain disorders such as ASD, as well as those involving ASE, could be determined. This will require development and validation of a plausible pipeline for ASE analysis in order to identify new genetic candidates for epigenetic mechanisms related to neuropathological characteristics of ASD, which could serve as targets for therapies of ASE-linked neurological disorders.
Methods
Subject material
We studied one family quartet comprised of parents and two offspring with and without ASD. In addition, one family trio, comprised of parents and one fetal offspring, was examined. Additional post-mortem samples included one fetal frontal cortex and one adult frontal cortex and cerebellum, shown in Supplementary Fig. 4. We assessed prefrontal cortex tissue from the dorsorostral pole of the frontal lobe corresponding to Brodmann’s area (BA) 10 for the offspring without ASD, and BA 8 for the offspring with ASD. Areas were based on tissue availability. Supplementary Fig. 1 and Supplementary Table 1 provides a more detailed pedigree of the family quartet showing epilepsy and deafness co-occurred in both the affected and unaffected offspring in addition to other familial health conditions. Moreover, we assessed prefrontal cortex tissue from the fetus and blood from the parents of the fetus. The detailed information of this family trio is shown in Supplementary Table 1. The detailed information of samples used for Supplementary Fig. 4 is shown in Supplementary Table 1. The institutional review boards (IRBs) of the participating institutions approved all experimental protocols. All experiments were carried out in accordance with the approved guidelines of the IRBs of the participating institutions. Written informed consent was obtained from both parents. Human quartet samples were obtained from the University of Utah Autism Research Program. The IRBs of the University of Utah, Icahn School of Medicine and National Taiwan University approved the analyses of samples and data in this study. Human trio samples and samples used for Supplementary Fig. 4 were obtained from National Taiwan University Hospital. The IRBs of the National Taiwan University approved the analyses of samples and data in this study. All tissue was fresh-frozen and stored at −80 degrees.
RNA extraction and RNA sequencing (RNA-Seq)
Total RNA was extracted from postmortem PFC tissue using a NucleoSpin miRNA kit (Macherey-Nagel, 740971) according to the manufacturer’s instructions. The RNA was quantified with an ND-1000 spectrophotometer (Nanodrop Technology). The quality of RNA was based on the RNA integrity number (RIN) measured with a Bioanalyzer 2100 (Agilent Technology) and an RNA 6000 LabChip Kit (Agilent Technology). Ribosomal RNA was removed from the purified RNA with the Ribo-Zero Magnetic Gold Kit (Epicentre, MRZG126). Purified RNA was then amplified and prepared for sequencing with a SureSelect Strand-Specific RNA Library Prep Kit (Agilent Technologies, G9691A). Libraries were sequenced using sequencing-by-synthesis technology (TruSeq SBS Kit v3-HS, Illumina, FC-401-3001) on an Illumina HiSeq. 2000 (100 base pairs paired-end reads) at the Welgene Biotech Company, which generated 6 Gb reads of data per sample. The detailed information of RNA-Seq is shown in Supplementary Table 1.
Detection of variants
DNA from the parents and offspring was extracted and sequenced according to standard whole-genome sequencing protocol. The sequencing reads were trimmed with Trimmomatic to obtain the qualified reads. The reads were then aligned to the human reference genome GRCh38 using BWA and processed with SAMtools. Picard (http://broadinstitute.github.io/picard/) was implemented to mark the duplicate reads and exclude them from downstream analyses. The read alignments were further refined with GATK for local realignment of reads around known insertions and deletions (indels) and recalibration of base quality. GATK was also applied to call single-nucleotide variants (SNVs) and short indels. Data from SNVs were consequently used to construct the haplotype scaffolds as described below. The RNA-Seq and DNA-Seq data discussed in this publication have been deposited in NCBI’s Gene Expression Omnibus and are accessible through GEO Series accession number GSE98581.
Allele-specific expression analysis
The SNV data from the parents and offspring were processed with VCFtools. SHAPEIT was then used to phase the SNV data with the family pedigree for building phased haplotype scaffolds. To improve phasing accuracy, the information of recombination rates between SNPs was provided via a genetic map retrieved from The 1000 Genomes Project Phase 3. The reference panel of phased haplotypes belonging to Utah residents with Northern and Western European ancestry (CEU) from The 1000 Genomes Project Phase 3 was also applied to align SNPs between the dataset and the panel for assisting in reliable phasing. The information of phased haplotypes was subsequently analyzed with in-house scripts to create haploid genomic sequences for the parents and offspring based on the human reference genome GRCh38. For RNA-Seq analysis, the sequences generated were filtered to obtain qualified reads. ConDeTri was implemented to trim or remove the reads according to the quality score. Qualified reads after filtering low-quality data were analyzed using TopHat/Cufflinks for gene expression estimation. The gene expression level was calculated as FPKM (Fragments Per Kilobase of transcript per Million mapped reads). A comprehensive analysis of the tissue using mammalian transcriptome data sets suggests that a lower cutoff of FPKM = 0.3 is often justifiable therefore we applied this cutoff for all analysis of mammalian transcriptomes. For differential expression analysis, CummeRbund was employed to perform statistical analyses of gene expression profiles. For allele-specific expression analysis, MMSEQ was then implemented to estimate allelic imbalance and deconvolve the alignment of reads to diploid transcripts derived from diploid genomic sequences and Ensembl gene annotation 74 following the mapping of RNA-Seq reads with Bowtie. SNPs for confirming imprinted miRNAs are within primary miRNA sequence. We defined the sequence for primary miRNAs as the genomic locus from 500 bp upstream to 500 bp downstream of the mature miRNA sequence. The FPKM value is much less than from regular read analysis because only the read with SNPs can be used for allele-specific expression analysis.
Reverse transcription quantitative PCR (RT-qPCR)
Total RNA was extracted from the postmortem PFC of the offspring using a NucleoSpin® miRNA kit (Macherey-Nagel, 740971). Total RNA (10 ng) was converted to cDNA and amplified by One Step SYBR® PrimeScriptTMRT-PCR Kit II (Takara, PR086A). Quantitative real-time PCR was performed with a StepOnePlus Real Time PCR System (Applied Biosystems). Ct values were generated using StepOne Software version 2.2.2. The expression level of each gene was normalized to B2M. All primer sequences of candidate genes were designed by Primer3 software (http://bioinfo.ut.ee/primer3-0.4.0/) and are shown in Supplementary Table 19.
miRNA quantification
We extracted miRNA using a NucleoSpin® miRNA kit (Macherey-Nagel, 740971). Because the length of miRNA is too short to perform normal qRT-PCR, the miRNA was lengthened with a Poly(A) tail (Poly(A) Tailing Kit; Ambion, AM1350). The poly(A) tailed miRNA was reverse transcribed into cDNA with a poly(T) anchor adaptor. The miRNA was amplified and quantitated by qPCR using a specific miRNA forward primer and a universal adaptor primer. Information of primer sequence is shown in Supplementary Table 20.
Similarly, the qPCR product was too small for Sanger sequencing. Therefore, to determine the sequence, the amplified PCR product was inserted into a plasmid vector, and the vector was transformed into bacteria with the TOPO TA Cloning Kit (Invitrogen, 450071), cloned, and cultured according to the manufacturer’s directions. Plasmid DNA containing the inserted qPCR product was extracted with the Presto Mini Plasmid kit (Geneaid, PHD100). Sanger sequencing of the purified plasmid used M13-tailed primers, which can yield sequences up to approximately 200 bp.
Graphic representation and statistical analysis
Heatmaps were generated with Pretty Heatmaps software (pheatmap package in R 3.3.2). The hierarchical clustering of heatmaps and the supplementary tables were measured in Euclidean distance. Micro-RNA target prediction was performed via DIANA microT-CDS web-based program. Pedegree was generated with Genial Pedigree Draw (www.pedigreedraw.com). Gene ontology (GO) analysis were performed via web-based Gorilla program and miEAA program for gene set enrichment analysis (GSEA) adapted for miRNA. Statistical analysis and graphic illustrations were performed using R 3.3.2 and Sigmaplot 13.0. All values with technical triplicates are expressed as the mean ± standard error of mean (s.e.m.).
Data availability statement
All data are available in this manuscript.
References
Lord, C. & Bishop, S. L. Recent advances in autism research as reflected in DSM-5 criteria for autism spectrum disorder. Annu Rev Clin Psychol 11, 53–70, https://doi.org/10.1146/annurev-clinpsy-032814-112745 (2015).
Christensen, D. L. et al. Prevalence and Characteristics of Autism Spectrum Disorder Among Children Aged 8 Years - Autism and Developmental Disabilities Monitoring Network, 11 Sites, United States, 2012. MMWR Surveill Summ 65, 1–23, https://doi.org/10.15585/mmwr.ss6503a1 (2016).
Ebrahimi-Fakhari, D. & Sahin, M. Autism and the synapse: emerging mechanisms and mechanism-based therapies. Current opinion in neurology 28, 91–102, https://doi.org/10.1097/WCO.0000000000000186 (2015).
Yatawara, C. J., Einfeld, S. L., Hickie, I. B., Davenport, T. A. & Guastella, A. J. The effect of oxytocin nasal spray on social interaction deficits observed in young children with autism: a randomized clinical crossover trial. Mol Psychiatry 21, 1225–1231, https://doi.org/10.1038/mp.2015.162 (2016).
Sandin, S. et al. The familial risk of autism. JAMA 311, 1770–1777, https://doi.org/10.1001/jama.2014.4144 (2014).
Kim, Y. S. & Leventhal, B. L. Genetic epidemiology and insights into interactive genetic and environmental effects in autism spectrum disorders. Biol Psychiatry 77, 66–74, https://doi.org/10.1016/j.biopsych.2014.11.001 (2015).
Wright, C. et al. Altered expression of histamine signaling genes in autism spectrum disorder. Translational psychiatry 7, e1126, https://doi.org/10.1038/tp.2017.87 (2017).
Gupta, S. et al. Transcriptome analysis reveals dysregulation of innate immune response genes and neuronal activity-dependent genes in autism. Nat Commun 5, 5748, https://doi.org/10.1038/ncomms6748 (2014).
Voineagu, I. et al. Transcriptomic analysis of autistic brain reveals convergent molecular pathology. Nature 474, 380–384, https://doi.org/10.1038/nature10110 (2011).
Parikshak, N. N. et al. Genome-wide changes in lncRNA, splicing, and regional gene expression patterns in autism. Nature 540, 423–427, https://doi.org/10.1038/nature20612 (2016).
Chaste, P. et al. A genome-wide association study of autism using the Simons Simplex Collection: Does reducing phenotypic heterogeneity in autism increase genetic homogeneity? Biol Psychiatry 77, 775–784, https://doi.org/10.1016/j.biopsych.2014.09.017 (2015).
Yuen, R. K. et al. Whole-genome sequencing of quartet families with autism spectrum disorder. Nat Med 21, 185–191, https://doi.org/10.1038/nm.3792 (2015).
Yuen, R. K. et al. Genome-wide characteristics of de novo mutations in autism. NPJ Genom Med 1, 160271–1602710, https://doi.org/10.1038/npjgenmed.2016.27 (2016).
Sanders, S. J. et al. Insights into Autism Spectrum Disorder Genomic Architecture and Biology from 71 Risk Loci. Neuron 87, 1215–1233, https://doi.org/10.1016/j.neuron.2015.09.016 (2015).
Pinto, D. et al. Functional impact of global rare copy number variation in autism spectrum disorders. Nature 466, 368–372, https://doi.org/10.1038/nature09146 (2010).
Wu, Y. E., Parikshak, N. N., Belgard, T. G. & Geschwind, D. H. Genome-wide, integrative analysis implicates microRNA dysregulation in autism spectrum disorder. Nature neuroscience 19, 1463–1476, https://doi.org/10.1038/nn.4373 (2016).
Tsang, S. Y. et al. Variation of global DNA methylation levels with age and in autistic children. Hum Genomics 10, 31, https://doi.org/10.1186/s40246-016-0086-y (2016).
Wong, C. C. et al. Methylomic analysis of monozygotic twins discordant for autism spectrum disorder and related behavioural traits. Mol Psychiatry 19, 495–503, https://doi.org/10.1038/mp.2013.41 (2014).
Ladd-Acosta, C. et al. Common DNA methylation alterations in multiple brain regions in autism. Mol Psychiatry 19, 862–871, https://doi.org/10.1038/mp.2013.114 (2014).
McCary, L. M. & Roberts, J. E. Early identification of autism in fragile X syndrome: a review. J Intellect Disabil Res 57, 803–814, https://doi.org/10.1111/j.1365-2788.2012.01609.x (2013).
Wilkinson, L. S., Davies, W. & Isles, A. R. Genomic imprinting effects on brain development and function. Nature reviews. Neuroscience 8, 832–843, https://doi.org/10.1038/nrn2235 (2007).
Lyon, M. F. Gene action in the X-chromosome of the mouse (Mus musculus L.). Nature 190, 372–373 (1961).
Nica, A. C. & Dermitzakis, E. T. Expression quantitative trait loci: present and future. Philos Trans R Soc Lond B Biol Sci 368, 20120362, https://doi.org/10.1098/rstb.2012.0362 (2013).
Lappalainen, T. et al. Transcriptome and genome sequencing uncovers functional variation in humans. Nature 501, 506–511, https://doi.org/10.1038/nature12531 (2013).
Reinius, B. & Sandberg, R. Random monoallelic expression of autosomal genes: stochastic transcription and allele-level regulation. Nature reviews. Genetics 16, 653–664, https://doi.org/10.1038/nrg3888 (2015).
Perez, J. D., Rubinstein, N. D. & Dulac, C. New Perspectives on Genomic Imprinting, an Essential and Multifaceted Mode of Epigenetic Control in the Developing and Adult Brain. Annu Rev Neurosci, https://doi.org/10.1146/annurev-neuro-061010-113708 (2016).
Gendrel, A. V. et al. Developmental dynamics and disease potential of random monoallelic gene expression. Dev Cell 28, 366–380, https://doi.org/10.1016/j.devcel.2014.01.016 (2014).
Adegbola, A. A. et al. Monoallelic expression of the human FOXP2 speech gene. Proceedings of the National Academy of Sciences of the United States of America 112, 6848–6854, https://doi.org/10.1073/pnas.1411270111 (2015).
Jeffries, A. R. et al. Random or stochastic monoallelic expressed genes are enriched for neurodevelopmental disorder candidate genes. PloS one 8, e85093, https://doi.org/10.1371/journal.pone.0085093 (2013).
Ben-David, E., Shohat, S. & Shifman, S. Allelic expression analysis in the brain suggests a role for heterogeneous insults affecting epigenetic processes in autism spectrum disorders. Human molecular genetics 23, 4111–4124, https://doi.org/10.1093/hmg/ddu128 (2014).
Schaaf, C. P. et al. Truncating mutations of MAGEL2 cause Prader-Willi phenotypes and autism. Nature genetics 45, 1405–1408, https://doi.org/10.1038/ng.2776 (2013).
Cook, E. H. et al. Autism or atypical autism in maternally but not paternally derived proximal 15q duplication. American journal of human genetics 60, 928–934 (1997).
Roach, J. C. et al. Analysis of genetic inheritance in a family quartet by whole-genome sequencing. Science 328, 636–639, https://doi.org/10.1126/science.1186802 (2010).
Browning, S. R. & Browning, B. L. Haplotype phasing: existing methods and new developments. Nat Rev Genet 12, 703–714, https://doi.org/10.1038/nrg3054 (2011).
Park, H. J. et al. Elevated Id2 expression results in precocious neural stem cell depletion and abnormal brain development. Stem Cells 31, 1010–1021, https://doi.org/10.1002/stem.1351 (2013).
Zhao, X. et al. A missense mutation in AGTPBP1 was identified in sheep with a lower motor neuron disease. Heredity (Edinb) 109, 156–162, https://doi.org/10.1038/hdy.2012.23 (2012).
Andrews, W. D., Barber, M., Nemitz, M., Memi, F. & Parnavelas, J. G. Semaphorin3A-neuropilin1 signalling is involved in the generation of cortical interneurons. Brain Struct Funct 222, 2217–2233, https://doi.org/10.1007/s00429-016-1337-3 (2017).
Hicks, S. D. & Middleton, F. A. A Comparative Review of microRNA Expression Patterns in Autism Spectrum Disorder. Front Psychiatry 7, 176, https://doi.org/10.3389/fpsyt.2016.00176 (2016).
Hsu, C. L. et al. Analysis of experience-regulated transcriptome and imprintome during critical periods of mouse visual system development reveals spatiotemporal dynamics. Human molecular genetics, https://doi.org/10.1093/hmg/ddy023 (2018).
Lin, C. Y. et al. Analysis of Genome-Wide Monoallelic Expression Patterns in Three Major Cell Types of Mouse Visual Cortex Using Laser Capture Microdissection. PloS one 11, e0163663, https://doi.org/10.1371/journal.pone.0163663 (2016).
Girardot, M., Cavaille, J. & Feil, R. Small regulatory RNAs controlled by genomic imprinting and their contribution to human disease. Epigenetics: official journal of the DNA Methylation Society 7, 1341–1348, https://doi.org/10.4161/epi.22884 (2012).
Bonthuis, P. J. et al. Noncanonical Genomic Imprinting Effects in Offspring. Cell Rep 12, 979–991, https://doi.org/10.1016/j.celrep.2015.07.017 (2015).
Shen, Y. et al. Activation of the Jnk signaling pathway by a dual-specificity phosphatase, JSP-1. Proceedings of the National Academy of Sciences of the United States of America 98, 13613–13618, https://doi.org/10.1073/pnas.231499098 (2001).
Nishina, H., Wada, T. & Katada, T. Physiological roles of SAPK/JNK signaling pathway. J Biochem 136, 123–126, https://doi.org/10.1093/jb/mvh117 (2004).
Leblond, C. S. et al. Genetic and functional analyses of SHANK2 mutations suggest a multiple hit model of autism spectrum disorders. PLoS genetics 8, e1002521, https://doi.org/10.1371/journal.pgen.1002521 (2012).
Kukurba, K. R. et al. Allelic expression of deleterious protein-coding variants across human tissues. PLoS genetics 10, e1004304, https://doi.org/10.1371/journal.pgen.1004304 (2014).
Li, J. et al. Integrated systems analysis reveals a molecular network underlying autism spectrum disorders. Mol Syst Biol 10, 774, https://doi.org/10.15252/msb.20145487 (2014).
Ren, C. M. et al. Balanced translocation t(3;18)(p13; q22.3) and points mutation in the ZNF407 gene detected in patients with both moderate non-syndromic intellectual disability and autism. Biochim Biophys Acta 1832, 431–438, https://doi.org/10.1016/j.bbadis.2012.11.009 (2013).
Charrier, A. et al. Zinc finger protein 407 overexpression upregulates PPAR target gene expression and improves glucose homeostasis in mice. Am J Physiol Endocrinol Metab 311, E869–E880, https://doi.org/10.1152/ajpendo.00234.2016 (2016).
Hiramuki, Y., Sato, T., Furuta, Y., Surani, M. A. & Sehara-Fujisawa, A. Mest but Not MiR-335 Affects Skeletal Muscle Growth and Regeneration. PloS one 10, e0130436, https://doi.org/10.1371/journal.pone.0130436 (2015).
Gregg, C. et al. High-resolution analysis of parent-of-origin allelic expression in the mouse brain. Science 329, 643–648, https://doi.org/10.1126/science.1190830 (2010).
Luedi, P. P. et al. Computational and experimental identification of novel human imprinted genes. Genome research 17, 1723–1730, https://doi.org/10.1101/gr.6584707 (2007).
Bjornsson, H. T. et al. SNP-specific array-based allele-specific expression analysis. Genome Res 18, 771–779, https://doi.org/10.1101/gr.073254.107 (2008).
Gagneur, J. et al. Genome-wide allele- and strand-specific expression profiling. Mol Syst Biol 5, 274, https://doi.org/10.1038/msb.2009.31 (2009).
Babak, T. et al. Global survey of genomic imprinting by transcriptome sequencing. Curr Biol 18, 1735–1741, https://doi.org/10.1016/j.cub.2008.09.044 (2008).
Nariai, N., Kojima, K., Mimori, T., Kawai, Y. & Nagasaki, M. A Bayesian approach for estimating allele-specific expression from RNA-Seq data with diploid genomes. BMC Genomics 17(Suppl 1), 2, https://doi.org/10.1186/s12864-015-2295-5 (2016).
Mosconi, M. W., Wang, Z., Schmitt, L. M., Tsai, P. & Sweeney, J. A. The role of cerebellar circuitry alterations in the pathophysiology of autism spectrum disorders. Front Neurosci 9, 296, https://doi.org/10.3389/fnins.2015.00296 (2015).
Cross-Disorder Group of the Psychiatric Genomics, C. Identification of risk loci with shared effects on five major psychiatric disorders: a genome-wide analysis. Lancet 381, 1371–1379, https://doi.org/10.1016/S0140-6736(12)62129-1 (2013).
Goes, F. S. et al. Exome Sequencing of Familial Bipolar Disorder. JAMA Psychiatry 73, 590–597, https://doi.org/10.1001/jamapsychiatry.2016.0251 (2016).
Levy, S. E. et al. Autism spectrum disorder and co-occurring developmental, psychiatric, and medical conditions among children in multiple populations of the United States. J Dev Behav Pediatr 31, 267–275, https://doi.org/10.1097/DBP.0b013e3181d5d03b (2010).
Acknowledgements
This work was supported by the Ministry of Science and Technology, Taipei, Taiwan (MOST 105-2628-B-002-033-MY3 to H-S.H), the National Health Research Institutes, Miaoli, Taiwan (Career Development Grant, NHRI-EX103-10316NC to H-S.H.), the National Taiwan University, Taipei, Taiwan (AIM for Top University Excellent Research Project, 102C101-42, 103C101-C1 and 104C101-B1 to S.S.G. and H-S.H), the National Taiwan University Hospital (NTUH-UN104-018, NTUH-UN105-014 and NTUH-UN106-011 to S.S.G. and H-S.H.), the Foundation for the Advancement of Outstanding Scholarship, Taipei, Taiwan (Young Scholars’ Creativity Award, to H-S.H), the intramural funding of Genomics Research Center, Academia Sinica, Taiwan and the Ministry of Science and Technology, Taiwan (MOST 103-2628-B-001-001-MY4 to T-J.C). We are grateful for support from the participating family as well as to Jane Pickett at Autism BrainNet for coordinating the pooling of blood- and brain-derived genetic material. We thank Trees-Juen Chuang for valuable reading of the manuscript. We thank Boting Wang for their experimental help. We would like to acknowledge Autism BrainNet, sponsored by the Simons Foundation and Autism Speaks, for providing biological material. Brain specimens were acquired through the Autism Tissue Program, a predecessor to Autism BrainNet, from the Harvard Brain Tissue Resources Center (McLean Hospital, Belmont, MA, USA) for the Autism Tissue Program, a predecessor to Autism BrainNet. Samples and family diagnostic data were originally collected by the University of Utah Autism Research Program (Hilary Coon) supported by RO1MH094400. Samples and data were made available through collaboration with this project and with additional direct permission from the family.
Author information
Authors and Affiliations
Contributions
H.-S.H. designed experiments. H.-S.H. and C.-Y.L. wrote the manuscript. H.-S.H., and S.S.G. supervised the studies. All authors reviewed, edited and approved the manuscript. C.-Y.L., K.-W.C., C.-Y.L., and J.-Y.W. carried out all experiments. P.-H.H., H.-N.H., H.C., and S.A. provided human samples.
Corresponding author
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Lin, CY., Chang, KW., Lin, CY. et al. Allele-specific expression in a family quartet with autism reveals mono-to-biallelic switch and novel transcriptional processes of autism susceptibility genes. Sci Rep 8, 4277 (2018). https://doi.org/10.1038/s41598-018-22753-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-018-22753-4
This article is cited by
-
Mir125b-2 imprinted in human but not mouse brain regulates hippocampal function and circuit in mice
Communications Biology (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.