Abstract
Chlorination of drinking water protects humans from water-born pathogens, but it also produces low concentrations of dibromoacetonitrile (DBAN), a common disinfectant by-product found in many water supply systems. DBAN is not mutagenic but causes DNA breaks and elevates sister chromatid exchange in mammalian cells. The WHO issued guidelines for DBAN after it was linked with cancer of the liver and stomach in rodents. How this haloacetonitrile promotes malignant cell transformation is unknown. Using fission yeast as a model, we report here that DBAN delays G1-S transition. DBAN does not hinder ongoing DNA replication, but specifically blocks the serine 345 phosphorylation of the DNA damage checkpoint kinase Chk1 by Rad3 (ATR) at broken replication forks. DBAN is particularly damaging for cells with defects in the lagging-strand DNA polymerase delta. This sensitivity can be explained by the dependency of pol delta mutants on Chk1 activation for survival. We conclude that DBAN targets a process or protein that acts at the start of S phase and is required for Chk1 phosphorylation. Taken together, DBAN may precipitate cancer by perturbing S phase and by blocking the Chk1-dependent response to replication fork damage.
Similar content being viewed by others
Introduction
Dibromoacetonitrile (DBAN) is generated at low nanomolar concentrations when bromide reacts with nitrogenous organic matter during the chlorination of drinking water1. A survey of 20 water supply systems in England and Wales revealed DBAN as the most abundant haloacetonitrile (HAN) (≤40.2 nM, ≤8 μg/L)2. Epidemiological studies in the USA linked the consumption of chlorinated water with an increased risk of bladder, brain and rectal cancer3,4,5. Very high concentrations of DBAN (>250 μM) induce cancer of the liver and stomach in rodents6,7. The WHO issued guidelines for DBAN (0.35 μM), dichloroacetonitrile (DCAN; 0.18 μM) and trichloroacetonitrile (TCAN; <0.007 μM) in the response to these findings8.
While DBAN is probably not mutagenic, it effectively alkylates DNA in vitro which may explain how it breaks DNA and elevates sister chromatid exchange in yeast and mammalian cells9,10,11. DBAN increases the levels of reactive oxygen species (ROS) in rat cells as indicated by a rise in 8-hydroxy-2-deoxyguanosine (8OHdG)12,13 and blocks aldehyde dehydrogenase14, dimethylnitrosamine-demethylase15, glutathione-S-transferase16, superoxide dismutase and catalase17 in the liver. Monobromoacetontrile (BAN) was shown to induce endoreplication in Chinese hamster ovary cells by blocking mitosis18. While these findings indicate a potential health risk, it is still unclear how DBAN precipitates cancer.
Using the model organism fission yeast (Schizosaccharomyces pombe), we report here that DBAN delays G1-S transition and specifically blocks the activation of the DNA damage checkpoint kinase Chk1 at broken DNA replication forks.
The replication machinery assembles around the DNA helicase MCM2-7 at dedicated chromosomal sites (origins) in a stepwise process (reviewed in19). The pre-replication complex forms late in G1 when DDK, a heterodimer of Cdc7/Hsk1 kinase and its activating subunit ASK/Dfp1,Dbf4, recruits Sld3 (Treslin) and Cdc45. The cell cycle regulator CDK (cyclin-dependent kinase) activates this complex subsequently at the start of S phase by loading the BRCT-domain protein Rad4 (Dbp11, TopBP1), Sld2, DNA polymerase epsilon and the GINS proteins20. Processive DNA replication begins when DNA polymerase alpha-primase and MCM10 associate with this structure. The lagging strand is displaced in front of the moving replication fork where Cdc45 channels it into DNA polymerase delta. The leading strand runs straight through the fork to be copied by DNA polymerase epsilon21,22. A drop in the nucleotide pool, which can be triggered by the RNR (ribonucleotide reductase) inhibitor hydroxyurea (HU), stalls forks in early S phase by activating the intra-S checkpoint kinase Cds123. Cds1 binds to Mrc1 (Claspin) at the stalled fork when both proteins are phosphorylated by Rad3 (ATR)24. Mrc1 associates also with early origins during the G1-S transition, independently of the checkpoint, where it is modified by DDK25. The collision of replication forks with immobilised topoisomerase 1, which can be trapped on the DNA by camptothecin (CPT), triggers the phosphorylation of the DNA damage checkpoint kinase Chk1 at serine 345 by Rad326. Activation of Cds1 and Chk1 both block the cell cycle activator Cdc2 (CDK1) thereby initiating a transient G2-M arrest27.
We report here that DBAN perturbs G1-S transition and blocks Chk1 phosphorylation at a broken replication fork without affecting the Rad3-dependent modification of other checkpoint proteins. DBAN may therefore act on a process or a protein that is required at the start of S phase and later at damaged DNA replication forks. We conclude that DBAN elicits DNA replication stress, a known driver of cancer development28.
Results
DBAN interferes with S phase
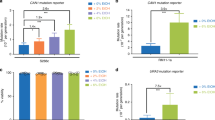
Informed by the ability of bromoacetonitrile (12 μM) to block mitosis in Chinese hamster ovary (CHO) cells18, we first tested whether haloacetonitriles (HANs) impact on cell cycle progression. Wild type cells were synchronised in G2 by lactose gradient centrifugation29 and released into medium with 10 μM bromo-, dibromo-, chloro-, dichloro- or trichloroactonitile (BAN, DBAN, CAN, DCAN, TCAN). Samples were withdrawn every 20 min to score septated G1/S cells. While the monohalogen compounds, BAN and CAN, allowed cells to complete two cell cycle rounds, their dihalogen forms, DCAN and DBAN, delayed entry into the second cycle by 40 min and 60 min, respectively (Fig. 1C,D). Unlike CHO cells18, DBAN-treated yeast cells arrested in G2 before the onset of mitosis (Fig. S1A,B). Since a second cycle G2 arrest is typical for drugs like hydroxyurea (HU) or camptothecin (CPT) that interfere with DNA replication, we concluded that DBAN and DCAN perturb S phase thereby triggering the G2 delay (Fig. 1G). HU stalls DNA replication forks by depleting the dNTP pool, whereas CPT breaks forks by immobilising topoisomerase 1 in front of the advancing replication complex. Both events delay onset of mitosis through the Rad3-dependent activation of the checkpoint kinases Cds1 and Chk1, respectively27,30. To investigate whether the DBAN-induced S phase perturbations activate the checkpoint, we synchronised a checkpoint-defective Δrad3 Δtel1 double mutant (Tel1/ATM is the second checkpoint kinase besides Rad3/ATR) in G2 and released cells into medium with or without 10 μM DBAN. Interestingly, the checkpoint deficient strain arrested like wild type for 80 min (Fig. S1A,C) showing that the G2 delay is independent of the DNA damage response. An unexpected observation was made when we analysed the cell cycle impact of trichloroacetonitrile. Unlike DBAN or DCAN, TCAN (10 μM) blocked entry into the first cycle for around 280 min (Fig. 1E). Since such a first cycle arrest is typical for agents that break DNA30 and because HANs were linked with DNA breaks in mammalian cells10, we tested whether they increase the phosphorylation of histone 2AX at S129 by Rad3 and Tel1, an established marker of chromosomal breaks31. Intriguingly, all HANs (BAN, CAN, DBAN, TCAN), with the exception of DCAN, reduced the phosphorylation of H2AX showing that DNA breaks are not the cause of the arrest (Fig. S1D). Since H2AX is also phosphorylated during unperturbed S phase31, this drop may be caused by depleting the pool of S phase cells due to the G2 arrest. We can however not exclude the possibility that the HANs affect H2AX phosphorylation more directly since BAN and CAN do not show a strong cell cycle arrest but reduce H2AX modification (Fig. 1A,B; Fig. S1D). The decline in H2AX phosphorylation was concentration dependent starting at 8 μM DBAN (Fig. S1E). We next tested whether the first cycle arrest is unique to TCAN and found that a higher concentration of DBAN (20 μM) also delayed cells in the first G2 phase (Fig. 1F). This implies that TCAN is more effective than DBAN in eliciting this response. As in the case of the second cycle arrest, the DNA damage checkpoint was not required (Fig. S1F). A checkpoint-defective Δcds1 Δchk1 double mutant delayed mitosis for 220 min, although this arrest was 40 min shorter compared to wild type cells (Fig. S1F). We did not further investigate this first cycle arrest as we wanted to learn more about how DBAN perturbs S phase given the importance of DNA replication stress in malignant transformation28.
Dibromoacetonitrile (DBAN) arrests cell cycle progression in a concentration dependent manner. Wild type cells (ade6-M210 leu1–32 ura4-D18) were synchronised in G2 by lactose gradient centrifugation29 and released into rich medium (3% glucose, 0.5% yeast extract, 100 mg/L adenine) without (UT = untreated) or with 10 μM of the indicated haloacetonitriles (HANs). All HANs were diluted from a 12 mM stock solution in DMSO to a final concentration of 10 μM. (A) bromoacetonitrile (BAN), (B) chloroacetonitrile (CAN), (C) dichloroacetonitrile (DCAN), (D) dibromoacetonitrile (DBAN), (E) trichloroacetonitrile (TCAN), (F) DBAN at 10 μM or 20 μM. (G) DBAN affects cells in S phase which triggers a second cycle delay.
DBAN delays G1-S progression
To map the execution point of DBAN in S phase, wild type cells were enriched in G1 by nitrogen starvation and released back into the cell cycle by replenishing the medium with a nitrogen source29. The DNA content was measured by flow cytometry over 8 hours to monitor progression from G1 (1 copy of the chromosomes, 1 C) into G2 (two copies, 2 C) (Fig. 2). While untreated cells reached G2 4 h post-release (Fig. 2A), DBAN (10 μM) delayed exit from G1 whereas TCAN (10 μM) blocked cells in G1 (Fig. 2B,C). We also arrested cells in early S phase with 15 mM HU to have an internal marker for unreplicated DNA (Fig. 2B). Eight hours after the release from G1, only HU and TCAN treated cells remained arrested with a 1 C DNA content, while DBAN permitted the completion of DNA replication (Fig. 2B,C). This shows that DBAN only delays G1-S transition, whereas TCAN blocks this step more effectively. None of the other HANs (BAN, CAN, DCAN) had a similar effect on G1-S transition (Fig. 2B,C).
DBAN and TCAN delay G1-S transition. (A) Wild type cells without auxotrophic markers were synchronised in G1 by nitrogen starvation in minimal medium (3% glucose, 0.67% nitrogen base w/o amino acids and ammonium sulphate) at 30 °C29. Cells were washed and released at T = 0 h into pre-warmed minimal medium with ammonium sulphate. Samples were withdrawn at the indicated time points and the DNA content was measured by flow cytometry29. The dotted lines indicate 1 copy of the chromosomes (1 C, G1) and two copies (2 C, G2), respectively. (B,C) DNA content at the start of the experiment (0 h), at 4 h and 8 h post-release. Untreated cells (UT) progress from G1 (1 C) to G2 (2 C) within 4 h. This progression is slowed down by DBAN but blocked by TCAN (indicated by the arrows). The final concentration of all HANs is 10 μM. In a control experiment, cells were arrested in early S (1 C) with 15 mM hydroxyurea (HU).
To exclude the possibility that the G1 arrest protocol impacts on this interesting finding, we synchronised nda3-KM311 cells in mitotic prophase. This cold sensitive beta-tubulin mutant stops with condensed chromosomes without a mitotic spindle at 20 °C and returns to the cell cycle within minutes upon a temperature up-shift to 30 °C32. In line with the first experiment, TCAN prevented the accumulation of G2 cells whereas DBAN only delayed it (Fig. S1G).
We next arrested cells in early S phase by incubating a wild type strain in 15 mM HU for 3.5 h29 to test whether addition of DBAN or TCAN (10 μM) after the G1-S transition would still delay the cell cycle. While the latter was not the case for DBAN (Fig. S2B), TCAN-treated cells delayed progression through S phase by 20 min compared to untreated cells (Fig. S2C). Since DNA replication was complete within 60 min in untreated cells (Fig. S2A), a 20 min difference (1/3 of S phase) may well be significant.
Taken together, these data show that DBAN delays G1-S transition while allowing cells to complete DNA replication. In contrast, TCAN blocks cells effectively before the G1-S transition and slightly delays DNA replication. This conclusion is in line with the higher potency of TCAN as a G2 blocker (Fig. 1E).
DBAN affects DNA polymerase delta
Since DBAN allows DNA replication to proceed after an initial G1-S delay (Figs 2C, S1G), we tested whether this affects the three replicative DNA polymerases, alpha (Pol1, swi7), epsilon (Pol2, cdc20) or delta (Pol3, cdc 6). Since these essential genes cannot be deleted, we used temperature-sensitive mutants at the semi-restrictive temperature of 30 °C. Serial dilutions of the strains were applied to rich medium plates containing no HAN or 10 μM DBAN. We also incubated one plate at 37 °C to confirm the temperature sensitivity. While mutations in the three essential subunits of Pol delta (cdc6.23 [catalytic], cdc 2 7.P11 [non-catalytic], cdc1.P13 [non-catalytic]) impaired cell viability, mutations in the catalytic subunits of Pol epsilon (cdc20.M10) or Pol alpha (swi7.H4) did not (Fig. 3B). Interestingly, deletion of the fourth, non-essential Pol delta subunit, cdm1, had also no effect (Fig. 3C). We next tested mutations in the MCM2-7 helicase that unwinds the DNA template (MCM2 [cdc19.P1], MCM4 [cdc21-M 6 8] or MCM5 [nda4-108]), but failed to detect loss of viability (Fig. 3C). Also no impact on cell growth was found for mutations in DDK (Cdc7/Hsk1) kinase (hsk1-1312) and in the replication factor Rad4 (TopBP1) (rad4.11 6) (Fig. 3C). Whether a mutation in Ctf4 (mcl1-1), which binds Pol alpha to the replication complex, impairs cell viability was difficult to judge since the strain grew very poorly even in the absence of DBAN (Fig. 3B). Interestingly, none of the DNA pol delta mutants showed a growth defect on TCAN plates even at 20 μM (Fig. 3D). This was unexpected given the high impact of TCAN on cell cycle progression. A possible explanation is provided by the replication delay caused by TCAN (Fig. S2C) that may prevent loss of viability of pol delta mutants. The latter mutants were also not sensitive to CAN, BAN or DCAN at 10 μM (Fig. 3E).
DBAN kills cells mutated in DNA polymerase delta. (A) Model of the replication fork (adapted from22). (B–E) Serial dilutions of the indicated strains were applied to rich medium plates at 30 °C. One plate was incubated at 37 °C. Incubation time: 3 days. The final concentration of DBAN was 10 μM, but 20 μM for TCAN. The mutant alleles are: pol alpha (swi7.H4), Ctf4 (mcl1-1), pol delta (cdc6.23, cdc27.P11, cdc1.p13), pol epsilon (cdc20.M10), S phase cell cycle inhibitor Mik1 (mik1::ura4+ ), cell cycle inhibitor Wee1 (wee1::ura4+ ), Δcdm1 (non-essential pol delta subunit, cdm1::ura4+ ), MCM2 (cdc19.P1), MCM4 (cdc21-M68), MCM5 (nda4-108), DDK/Cdc7 (hsk1-1312), Rad4 (TopBP1) (rad4.116), MAP kinase Sty1 (sty1::ura4+ ). Final concentration of BAN, CAN and DCAN is 10 μM.
DBAN overcomes the intra-S arrest of a pol delta mutant
To find out why pol delta mutants are DBAN sensitive, we synchronised wild type, pol alpha (swi7.H4), pol delta (cdc 2 7.P11) and pol epsilon (cdc20.M10) strains in early S phase using the HU arrest protocol29. After HU was washed out, cells were released into medium with or without 10 μM DBAN. Flow cytometry showed that untreated wild type cells completed S phase within 60 min (Fig. 4A). As previously reported33, the untreated mutant strains delayed S phase progression at the semi-permissive temperature of 30 °C (Fig. 4B–D). While DBAN had no impact on S phase within the first 60 min in the case of the pol alpha and pol epsilon mutants, it did significantly advance DNA replication of the pol delta (cdc27.P11) strain (Fig. 4C). This advancement was clearly detectable at the 60 min and 90 min time points. After 2 h, the DBAN-treated cdc27.P11 cells had initiated already the next cell cycle round compared to the untreated sample (Fig. 4C). Cdc27 connects the catalytic (Cdc6) and non-catalytic (Cdc1) subunits, and binds Pol delta to the DNA sliding clamp PCNA34. DBAN also advanced DNA replication in the other two mutant strains but later and to a lesser extent (Fig. 4B,D).
DBAN advances DNA replication in a pol delta mutant. (A) Wild type cells (ade6-M210 leu1–32 ura4-D18) were grown in rich medium at 30 °C and synchronised in early S phase for 3.5 h in 15 mM hydroxyurea (HU)29. HU was washed out and cells were released into pre-warmed rich medium. The DNA content was measured at the indicated times. The dotted lines indicate 1 copy of the chromosomes (1 C, G1) and two copies (2 C, G2), respectively. The DNA content of nitrogen starved cells (no N) was measured in a parallel experiment to have an internal standard. (B–D). The indicated mutant strains were HU-synchronised and released into rich medium at 30 °C with or without 10 μM DBAN. The green histogram is the DNA content of untreated cells, the red histogram is the DNA content in the presence of DBAN. The brown colour indicates that both histograms overlap.
These results imply that DBAN abolishes the intra-S phase arrest of the pol delta (cdc27.P11) strain, which may be linked with its loss of viability on DBAN plates (Fig. 3B). Since the viability of pol delta mutants depends on Chk1 kinase33,35 we next tested whether DBAN interferes with the activation of the checkpoint kinases Cds1 and Chk1.
DBAN suppresses the activation of Chk1
To find out whether DBAN perturbs activation of the intra-S checkpoint kinase Cds1 at stalled forks, asynchronous cds1-His 6 HA 2 cells36 were incubated with 10 μM DBAN, 12 mM HU or with both chemicals simultaneously for 4 h. Total protein extracts were loaded onto a 6% phos-tag SDS gel to assay the phosphorylation status of Cds1. Phostag electrophoresis reveals the phosphorylation pattern of proteins as their mobility is inversely related to the extent of their modification37. While DBAN did not promote the modification of Cds1, the kinase was intensively phosphorylated when DNA replication forks stalled in the presence of HU. Although DBAN did not impact on this hyperphosphorylation, it induced a faster migrating band (Fig. 5A). The latter band could be a hypophosphorylated form of Cds1.
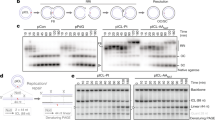
DBAN suppresses Chk1 phosphorylation. (A,B) cds1-His 6 HA 2 (55 kDa) and chk1-HA 3 (60 kDa) cells were grown in rich medium at 30 °C and treated for 4 h with 10 μM DBAN, 12 μM camptothecin (CPT), 12 mM hydroxyurea (HU) or the combination as indicated. Total protein extracts were loaded on a 6% phostag gel. Full phostag gels are shown. H2AX-S129-P was detected on a 20% acrylamide (37.5:1 acrylamide:bisacrylamide) gel. The Chk1 shift was detected on a 10% (100:1 acrylamide:bisacrylamide) gel. H2AX and Chk1 (normal) panels were cropped. The full images are shown in Figs S3 and S4, respectively. The arrows indicate the smaller Cds1 band, Chk1-S345 phosphorylation and the hyper-phosphorylated forms of Chk1. (C) Model of Chk1 activation by Rad3 at broken replication forks. CPT immobilises Topoisomerase 1 and the Rad9-Rad1-Hus1 ring aids Chk1 phosphorylation. Rad9 is also phosphorylated by Rad341. (D) Drop test of the indicated strains. Chk1-D155E-HA3 is a kinase-dead mutant. DBAN: 10 μM; CPT: 12 μM. (E,F) chk1-HA 3 cells were HU synchronised (3.5 h, 15 mM HU, rich medium) and released into medium without (UT) or with CPT (12 μM), CPT (12 μM) + DBAN (10 μM) or CPT (12 μM) + TCAN (10 μM). Total protein extracts were separated on a 8% (Mrc1) and 10% (Chk1) acrylamide gel. P = Phospho-Chk1-S345 shift band). (G) rad9-HA 3 cells were HU-synchronised and released into rich medium without (UT) or with CPT (12 μM), DBAN (10 μM) or CPT (12 μM) + DBAN (10 μM). P = Phospho-Rad9 shift band. Full images: Mrc1: Fig. S5; Chk1: Fig. S6; Rad9: Fig. S7.
We then repeated this experiment with a chk1-HA 3 strain38 but replaced HU with 12 μM camptothecin (CPT) to break DNA replication forks. In contrast to Cds1, DBAN effectively suppressed Chk1 phosphorylation at serine 345 (Fig. 5B). On normal SDS page, S345 phosphorylation was detected as a band shift, as previously reported, which disappeared upon DBAN exposure (Fig. 5B)38. DBAN also induced very slowly migrating, hyperphosphorylated bands of Chk1 independently of CPT. The suppression of Chk1 phosphorylation at damaged replication forks could explain why DBAN impairs the viability of DNA polymerase delta mutants which rely on this kinase for survival (Fig. 3B)35.
To test whether DBAN is an inhibitor of Rad3 kinase, we replaced CPT with MMS (methyl-methanesulfonate) that damages DNA by alkylation39. Since DBAN did not to block the MMS-induced phosphorylation of Chk1 (Fig. S1H), it is unlikely that the HAN impairs Rad3 kinase directly. We then exposed wild type cells (chk1-HA 3) to CPT or to the combination of DBAN and CPT on plates to gauge whether replication forks still break. The latter seems to be the case as DBAN rendered wild type cells (chk1-HA 3) CPT sensitive (Fig. 5D). This increase in sensitivity, which was similar to the sensitivity of a kinase-dead chk1 mutant (chk1-D155E-HA 3), indicates that forks still break while DBAN prevents activation of Chk1. We can however not exclude the possibility that CPT generates a block to DNA replication like a mutation in DNA polymerase delta that renders cells sensitive to DBAN.
To find out when DBAN acts on Chk1 in the cell cycle, we synchronised chk1-HA 3 cells in early S phase with 15 mM HU and released them into medium with CPT (12 μM) or with CPT and DBAN or TCAN (10 μM). In line with the idea that Chk1 is activated once bulk DNA synthesis had been completed40, the shift band of Chk1, indicative of serine 345 phosphorylation, appeared 60 min post-release after the levels of the DNA replication marker Mrc1 had declined (Fig. 5E)24. Since Chk1 is weakly phosphorylated during the HU arrest, all samples displayed a weak shift band that increased strongly when forks were damaged by CPT (Fig. 5E, bottom panel). The presence of DBAN or TCAN effectively suppressed activation of Chk1 (Fig. 5F). Since Rad3 modifies also the Rad9 subunit of the 9-1-1 checkpoint ring at broken forks (Fig. 5C)41, we repeated this experiment with a HU-synchronised rad9-HA 3 strain42. As in the case of Chk1, Rad9 phosphorylation is detectable as a band shift. This shift was not affected by DBAN strongly suggesting that Chk1 is specifically targeted by the haloacetonitrile (Fig. 5G). We noticed however that the Rad9 phosphorylation peaked 30 min earlier in the presence of DBAN (Fig. 5G, panels 3 + 4). The phos-tag assay did not reveal any changes in the Rad9 phosphorylation pattern which were DBAN specific (Fig. S1I).
Collectively, these results suggest that DBAN kills pol delta mutants (Fig. 3B) and renders wild type cells sensitive to camptothecin (Fig. 5D) by preventing the phosphorylation of Chk1 at damaged DNA replication forks.
Discussion
The evidence presented here reveals novel activities of DBAN at two stages during the cell cycle, at the G1-S transition and later at damaged replication forks (Fig. 6). DBAN affects both processes in a negative way as it delays entry into S phase (Fig. 2) and suppresses phosphorylation of Chk1 (Fig. 5B,F) without affecting the activation of Rad9 (Fig. 5G) or Cds1 (Fig. 5A). Replication fork damage can originate from the inhibition of topoisomerase 1 by camptothecin or from mutations in the lagging strand DNA polymerase delta26,33. Collectively, these findings imply that DBAN blocks an event or protein that is required for both entry into S phase and activation of Chk1 at damaged forks (Fig. 6). Early in S phase, human and yeast cells phosphorylate the histone H2AX in a cell cycle specific manner independently of DNA breaks31,43. The same chromatin modification is later required for the recruitment of Crb2 (53BP1) to a broken fork where the scaffold protein associates with Chk144,45. DBAN and TCAN may therefore compromise both processes by reducing H2AX phosphorylation (Fig. S1D,E). For example, the HANs may up-regulate a phosphatase, like human PP2A-B56ϵ, that dephosphorylates H2AX at damaged forks upon CPT treatment46. What argues against this model is the survival of the DNA pol delta mutants in the presence of BAN, CAN or TCAN (Fig. 3) which all reduce H2AX phosphorylation (Fig. S1D). An alternative explanation is provided by the dual function of Rad4/Cut5 (TopBP1) during G1-S transition and in the activation of Chk147,48. Rad4 associates with Sld3 and Sld2/Drc1 at start and with Crb2 at broken DNA45,47. What argues however against Rad4 is its requirement for the activation of Cds1 at stalled forks48 which is not affected by DBAN (Fig. 5A). A third candidate is DDK kinase as it initiates the assembly of the replication complex at the end of G1 (reviewed in19) and terminates Chk1 activation by phosphorylating the Rad9 subunit of the 9-1-1 complex49. We have however not found any evidence that the Rad9 phosphorylation pattern changes upon DBAN exposure (Fig. S1I). A fourth possibility is provided by Mrc1 (Claspin) that associates with early replication origins at start25 and recruits human Chk1 to broken forks50. A similar interaction between S. pombe Chk1 and Mrc1 has not yet been reported. Finally, DBAN may act directly on Chk1 as indicated by its hyper-phosphorylation (Fig. 5B). Chk1 is required in S. pombe and human cells at the start of the G1-S transition51,52 and at broken forks26. Further work will however be required to dissect the different possibilities that are of great interest as it is still enigmatic why human Chk1 occupies a dominant role in S phase whereas yeast Chk1 appears to act mainly in G2 (Fig. 5E)40.
DBAN is known to block a large number of enzymes in vitro 7 but it is as yet unclear how it interacts with proteins. Halogen atoms like bromine and chlorine have a positive charge, known as the alpha-hole, that can make electrostatic contacts with the protein backbone or amino acid side chains53. While bromine interacts preferentially with the side chain of arginine, chlorine prefers leucine54. Whether this explains why TCAN is a stronger cell cycle inhibitor than DBAN (Fig. 1) is difficult to tell as it is unclear to which protein they bind. It is even not entirely safe to conclude that they bind to the same protein. DBAN and TCAN behave in a similar way regarding the G1-S delay (Fig. 2) and the inhibition of Chk1 phosphorylation (Fig. 5F). Both HANs differ however in their lethality when DNA polymerase delta is mutated (Fig. 3D). Although this appears to contradict the earlier notion that DBAN kills these mutant strains by blocking Chk1 phosphorylation at damaged forks, it could be explained by the ability of TCAN to delay DNA replication (Fig. 4C). If this delay were to prevent fork damage in pol delta mutant cells, the inhibition of Chk1 would not affect cell viability.
Although DNA replication stress is a good explanation for why DBAN (10 μM) triggers a second cycle delay (Fig. 1D), it would not provide an answer to why 20 μM DBAN block cells in the first G2 (Fig. 1F). The concentration dependency implies that DBAN has either more than one target in cells with different affinities for the HAN or that DBAN upregulates the expression or activity of a protein in a concentration dependent manner. From the different options discussed earlier, the up-regulation of a phosphatase like PPA2 would connect the diverse cell cycle activities of DBAN and TCAN. Interestingly, induction of PPA2 arrests cells in G2 independently of the DNA damage checkpoint when the accessory protein Vpr of human immunodeficiency virus type 1 (HIV-1) is over-expressed in S. pombe 55. Since this arrest shares a similar independence from the Rad3-Tel1 checkpoint as the G2 arrest produced by DBAN and TCAN (Figs 1E,F; S1F), it is quite possible that both HANs delay cell cycle progression in G2 through a mechanism that involves the activation of a phosphatase acting on the cell cycle machinery.
The final point to consider is whether DBAN, which is frequently found in water supply systems2, posses a serious cancer risk. The concentration of 10 μM used in this study is approximately 30 times higher than the WHO guideline of 0.35 μM (70 μg/L). It is therefore unlikely that DBAN levels reach such high concentrations in tap water. The peak concentration found in water supply systems in Western Australia, for example, was 0.13 μM (26.6 μg/L)56, whereas the peak value in the United Kingdom was with 0.04 μM (8 μg/L) much lower2. It is however not yet clear whether haloacetonitriles accumulate over a longer consumption period in the liver, gastrointestinal tract or the kidneys7. The latter may explain why the consumption of chlorinated drinking water was linked with cancer3.
Materials and Methods
Yeast Strains
The genotype of the strains used in this study is ade6-M210 leu1–32 ura4-D18. Wild type cells contained no additional mutation and the different mutant alleles are mentioned in the text. The cds1-HA 2 His 6 [URA4 + ] ura4-D18 strain is described in36 and the chk1-HA 3 strain in38. Before cells were synchronised in G1 by nitrogen starvation, all auxotrophic markers were crossed out.
Cell synchronisation
Cells were synchronised as described in29. HU was used at a final concentration of 15 mM for 3.5 h at 30 °C in rich medium. For the G1 arrest, cells without auxotrophic markers were first grown in minimal medium with nitrogen before being transferred to minimal medium without nitrogen for 16 h at 30 °C. Lactose gradients were centrifuged for 8 min at 800 rpm. The nda 3 -KM311 mitotic arrest was performed in rich medium as reported in57.
Flow cytometry
The DNA content was measured using a CUBE 8 (Sysmex) instrument as described in29. The histograms were produced using the free Flowing Software (http://flowingsoftware.btk.fi/).
Phos-tag SDS page
Phostag gels (6%) were prepared and run as reported in58.
Antibodies
Anti-HA antibody (BioScource, Covance MMS-101P-200), anti-Mrc1 antibody (ABCAM, ab188269), anti-Cdc2 antibody (ab5467) and anti-Histone 2AX-S129-P antibody (ab17576). Secondary mouse-HRP (Dako, P0447), secondary rabbit-HRP (Dako, P0217).
Protein extracts
Total protein extracts were prepared as described in42.
Chemicals
Bromoacetonitrile (Sigma Aldrich, 242489), dibromoacetonitrile (Alfa Aesar, A16994), chloroacetonitrile (Sigma Aldrich, C19651), dichloroacetonitrile (Alfa Aesar, A10612), trichloroacetonitrile (Sigma Aldrich, T53805), phostag-acrylamide (AlphaLabs, AAL-107, 300-93523), hydroxyurea (Formedium, HDU0025).
References
Jia, A., Wu, C. & Duan, Y. Precursors and factors affecting formation of haloacetonitriles and chloropicrin during chlor(am)ination of nitrogenous organic compounds in drinking water. J. Hazard. Mater. 308, 411–418 (2016).
Bond, T., Templeton, M. R., Mokhtar Kamal, N. H., Graham, N. & Kanda, R. Nitrogenous disinfection byproducts in English drinking water supply systems: Occurrence, bromine substitution and correlation analysis. Water Res. 85, 85–94 (2015).
Morris, R. D., Audet, A. M., Angelillo, I. F., Chalmers, T. C. & Mosteller, F. Chlorination, chlorination by-products, and cancer: a meta-analysis. Am. J. Public Health 82, 955–963 (1992).
McGeehin, M. A., Reif, J. S., Becher, J. C. & Mangione, E. J. Case-control study of bladder cancer and water disinfection methods in Colorado. Am. J. Epidemiol. 138, 492–501 (1993).
Cantor, K. P. et al. Drinking water source and chlorination byproducts in Iowa. III. Risk of brain cancer. Am. J. Epidemiol. 150, 552–560 (1999).
Bull, R. J. et al. Evaluation of mutagenic and carcinogenic properties of brominated and chlorinated acetonitriles: by-products of chlorination. Fundam. Appl. Toxicol. Off. J. Soc. Toxicol. 5, 1065–1074 (1985).
National Toxicology Program. Toxicology and carcinogenesis studies of dibromoacetonitrile (CAS No. 3252-43-5) in F344/N rats and B6C3F1 mice (drinking water studies). Natl. Toxicol. Program Tech. Rep. Ser. 1–193 (2010).
Hu, Y. et al. Detection of genotoxic effects of drinking water disinfection by-products using Vicia faba bioassay. Environ. Sci. Pollut. Res. Int. 24, 1509–1517 (2017).
Muller-Pillet, V., Joyeux, M., Ambroise, D. & Hartemann, P. Genotoxic activity of five haloacetonitriles: comparative investigations in the single cell gel electrophoresis (comet) assay and the ames-fluctuation test. Environ. Mol. Mutagen. 36, 52–58 (2000).
Lin, E. L., Daniel, F. B., Herren-Freund, S. L. & Pereira, M. A. Haloacetonitriles: metabolism, genotoxicity, and tumor-initiating activity. Environ. Health Perspect. 69, 67–71 (1986).
Zhang, S.-H., Miao, D.-Y., Tan, L., Liu, A.-L. & Lu, W.-Q. Comparative cytotoxic and genotoxic potential of 13 drinking water disinfection by-products using a microplate-based cytotoxicity assay and a developed SOS/umu assay. Mutagenesis 31, 35–41 (2016).
Jacob, S., Kaphalia, B. S., Jacob, N. & Ahmed, A. E. The water disinfectant byproduct dibromoacetonitrile induces apoptosis in rat intestinal epithelial cells: possible role of redox imbalance. Toxicol. Mech. Methods 16, 227–234 (2006).
Ahmed, A. E., Jacob, S., Nagy, A. A. & Abdel-Naim, A. B. Dibromoacetonitrile-induced protein oxidation and inhibition of proteasomal activity in rat glioma cells. Toxicol. Lett. 179, 29–33 (2008).
Poon, R., Chu, I., LeBel, G., Yagminas, A. & Valli, V. E. Effects of dibromoacetonitrile on rats following 13-week drinking water exposure. Food Chem. Toxicol. Int. J. Publ. Br. Ind. Biol. Res. Assoc. 41, 1051–1061 (2003).
Pereira, M. A., Lin, L. H. & Mattox, J. K. Haloacetonitrile excretion as thiocyanate and inhibition of dimethylnitrosamine demethylase: a proposed metabolic scheme. J. Toxicol. Environ. Health 13, 633–641 (1984).
Ahmed, A. E., Soliman, S. A., Loh, J. P. & Hussein, G. I. Studies on the mechanism of haloacetonitriles toxicity: inhibition of rat hepatic glutathione S-transferases in vitro. Toxicol. Appl. Pharmacol. 100, 271–279 (1989).
Abdel-Wahab, M. H., Arafa, H. M. M., El-Mahdy, M. A. & Abdel-Naim, A. B. Potential protective effect of melatonin against dibromoacetonitrile-induced oxidative stress in mouse stomach. Pharmacol. Res. 46, 287–293 (2002).
Komaki, Y., Mariñas, B. J. & Plewa, M. J. Toxicity of drinking water disinfection byproducts: cell cycle alterations induced by the monohaloacetonitriles. Environ. Sci. Technol. 48, 11662–11669 (2014).
Tognetti, S., Riera, A. & Speck, C. Switch on the engine: how the eukaryotic replicative helicase MCM2-7 becomes activated. Chromosoma 124, 13–26 (2015).
Yabuuchi, H. et al. Ordered assembly of Sld3, GINS and Cdc45 is distinctly regulated by DDK and CDK for activation of replication origins. EMBO J. 25, 4663–4674 (2006).
Masai, H. et al. Phosphorylation of MCM4 by Cdc7 kinase facilitates its interaction with Cdc45 on the chromatin. J. Biol. Chem. 281, 39249–39261 (2006).
Georgescu, R. et al. Structure of eukaryotic CMG helicase at a replication fork and implications to replisome architecture and origin initiation. Proc. Natl. Acad. Sci. USA 114, E697–E706 (2017).
Xu, Y., Davenport, M. & Kelly, T. J. Two-stage mechanism for activation of the DNA replication checkpoint kinase Cds1 in fission yeast. Genes Dev. 20, 990–1003 (2006).
Tanaka, K. & Russell, P. Mrc1 channels the DNA replication arrest signal to checkpoint kinase Cds1. Nat. Cell Biol. 3, 966–972 (2001).
Matsumoto, S. et al. Checkpoint-Independent Regulation of Origin Firing by Mrc1 through Interaction with Hsk1 Kinase. Mol. Cell. Biol. 37 (2017).
Wan, S., Capasso, H. & Walworth, N. C. The topoisomerase I poison camptothecin generates a Chk1-dependent DNA damage checkpoint signal in fission yeast. Yeast Chichester Engl. 15, 821–828 (1999).
Lindsay, H. D. et al. S-phase-specific activation of Cds1 kinase defines a subpathway of the checkpoint response in Schizosaccharomyces pombe. Genes Dev. 12, 382–395 (1998).
Gaillard, H., García-Muse, T. & Aguilera, A. Replication stress and cancer. Nat. Rev. Cancer 15, 276–289 (2015).
Luche, D. D. & Forsburg, S. L. Cell-cycle synchrony for analysis of S. pombe DNA replication. Methods Mol. Biol. Clifton NJ 521, 437–448 (2009).
al-Khodairy, F. et al. Identification and characterization of new elements involved in checkpoint and feedback controls in fission yeast. Mol. Biol. Cell 5, 147–160 (1994).
Rozenzhak, S. et al. Rad3 decorates critical chromosomal domains with gammaH2A to protect genome integrity during S-Phase in fission yeast. PLoS Genet. 6, e1001032 (2010).
Hiraoka, Y., Toda, T. & Yanagida, M. The NDA3 gene of fission yeast encodes beta-tubulin: a cold-sensitive nda3 mutation reversibly blocks spindle formation and chromosome movement in mitosis. Cell 39, 349–358 (1984).
Francesconi, S., Grenon, M., Bouvier, D. & Baldacci, G. p56(chk1) protein kinase is required for the DNA replication checkpoint at 37 degrees C in fission yeast. EMBO J. 16, 1332–1341 (1997).
Bermudez, V. P., MacNeill, S. A., Tappin, I. & Hurwitz, J. The influence of the Cdc27 subunit on the properties of the Schizosaccharomyces pombe DNA polymerase delta. J. Biol. Chem. 277, 36853–36862 (2002).
Yin, L., Locovei, A. M. & D’Urso, G. Activation of the DNA damage checkpoint in mutants defective in DNA replication initiation. Mol. Biol. Cell 19, 4374–4382 (2008).
Aono, N., Sutani, T., Tomonaga, T., Mochida, S. & Yanagida, M. Cnd2 has dual roles in mitotic condensation and interphase. Nature 417, 197–202 (2002).
Kinoshita, E., Kinoshita-Kikuta, E., Takiyama, K. & Koike, T. Phosphate-binding tag, a new tool to visualize phosphorylated proteins. Mol. Cell. Proteomics MCP 5, 749–757 (2006).
Walworth, N. C. & Bernards, R. rad-dependent response of the chk1-encoded protein kinase at the DNA damage checkpoint. Science 271, 353–356 (1996).
Kosoy, A. & O’Connell, M. J. Regulation of Chk1 by its C-terminal domain. Mol. Biol. Cell 19, 4546–4553 (2008).
Redon, C. et al. Yeast histone 2A serine 129 is essential for the efficient repair of checkpoint-blind DNA damage. EMBO Rep. 4, 678–684 (2003).
Furuya, K., Poitelea, M., Guo, L., Caspari, T. & Carr, A. M. Chk1 activation requires Rad9 S/TQ-site phosphorylation to promote association with C-terminal BRCT domains of Rad4TOPBP1. Genes Dev. 18, 1154–1164 (2004).
Caspari, T. et al. Characterization of Schizosaccharomyces pombe Hus1: a PCNA-related protein that associates with Rad1 and Rad9. Mol. Cell. Biol. 20, 1254–1262 (2000).
Kurose, A. et al. Effects of hydroxyurea and aphidicolin on phosphorylation of ataxia telangiectasia mutated on Ser 1981 and histone H2AX on Ser 139 in relation to cell cycle phase and induction of apoptosis. Cytom. Part J. Int. Soc. Anal. Cytol. 69, 212–221 (2006).
Nakamura, T. M., Du, L.-L., Redon, C. & Russell, P. Histone H2A phosphorylation controls Crb2 recruitment at DNA breaks, maintains checkpoint arrest, and influences DNA repair in fission yeast. Mol. Cell. Biol. 24, 6215–6230 (2004).
Saka, Y., Esashi, F., Matsusaka, T., Mochida, S. & Yanagida, M. Damage and replication checkpoint control in fission yeast is ensured by interactions of Crb2, a protein with BRCT motif, with Cut5 and Chk1. Genes Dev. 11, 3387–3400 (1997).
Li, X., Nan, A., Xiao, Y., Chen, Y. & Lai, Y. PP2A-B56ϵ complex is involved in dephosphorylation of γ-H2AX in the repair process of CPT-induced DNA double-strand breaks. Toxicology 331, 57–65 (2015).
Fukuura, M. et al. CDK promotes interactions of Sld3 and Drc1 with Cut5 for initiation of DNA replication in fission yeast. Mol. Biol. Cell 22, 2620–2633 (2011).
Harris, S. et al. Delineating the position of rad4+/cut5+ within the DNA-structure checkpoint pathways in Schizosaccharomyces pombe. J. Cell Sci. 116, 3519–3529 (2003).
Furuya, K. et al. DDK phosphorylates checkpoint clamp component Rad9 and promotes its release from damaged chromatin. Mol. Cell 40, 606–618 (2010).
Chini, C. C. S. & Chen, J. Human claspin is required for replication checkpoint control. J. Biol. Chem. 278, 30057–30062 (2003).
Carr, A. M. et al. The chk1 pathway is required to prevent mitosis following cell-cycle arrest at ‘start’. Curr. Biol. CB 5, 1179–1190 (1995).
Ge, X. Q. & Blow, J. J. Chk1 inhibits replication factory activation but allows dormant origin firing in existing factories. J. Cell Biol. 191, 1285–1297 (2010).
Sirimulla, S., Bailey, J. B., Vegesna, R. & Narayan, M. Halogen Interactions in Protein–Ligand Complexes: Implications of Halogen Bonding for Rational Drug Design. J. Chem. Inf. Model. 53, 2781–2791 (2013).
Kortagere, S., Ekins, S. & Welsh, W. J. Halogenated ligands and their interactions with amino acids: implications for structure-activity and structure-toxicity relationships. J. Mol. Graph. Model. 27, 170–177 (2008).
Elder, R. T. et al. HIV-1 Vpr induces cell cycle G2 arrest in fission yeast (Schizosaccharomyces pombe) through a pathway involving regulatory and catalytic subunits of PP2A and acting on both Wee1 and Cdc25. Virology 287, 359–370 (2001).
Liew, D., Linge, K. L. & Joll, C. A. Formation of nitrogenous disinfection by-products in 10 chlorinated and chloraminated drinking water supply systems. Environ. Monit. Assess. 188, 518 (2016).
Nakazawa, N., Mehrotra, R., Ebe, M. & Yanagida, M. Condensin phosphorylated by the Aurora-B-like kinase Ark1 is continuously required until telophase in a mode distinct from Top2. J. Cell Sci. 124, 1795–1807 (2011).
Caspari, T. & Hilditch, V. Two Distinct Cdc2 Pools Regulate Cell Cycle Progression and the DNA Damage Response in the Fission Yeast S. pombe. PloS One 10, e0130748 (2015).
Acknowledgements
We are grateful to George Meyrick for matched funding in support of the KESS (Knowledge Economy Skills Scholarships) Master project (KESS Mini 111).
Author information
Authors and Affiliations
Contributions
T.C. wrote the manuscript, designed the study, performed most experiments and analysed the data. J.D. performed some experiments. N.F., C.D. and C.F. are co-applicants with T.C. on the KESS grant and initiated the project. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Competing Interests
The authors declare that they have no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Caspari, T., Dyer, J., Fenner, N. et al. The drinking water contaminant dibromoacetonitrile delays G1-S transition and suppresses Chk1 activation at broken replication forks. Sci Rep 7, 12730 (2017). https://doi.org/10.1038/s41598-017-13033-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-017-13033-8
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.