Abstract
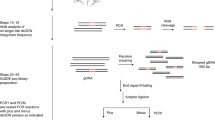
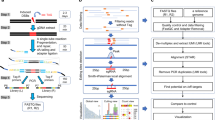
Programmable cytosine base editors show promising approaches for correcting pathogenic mutations; yet, their off-target effects have been of great concern. Detect-seq (dU-detection enabled by C-to-T transition during sequencing) is an unbiased, sensitive method for the off-target evaluation of programmable cytosine base editors. It profiles the editome by tracing the editing intermediate dU, which is introduced inside living cells and edited by programmable cytosine base editors. The genomic DNA is extracted, preprocessed and labeled by successive chemical and enzymatic reactions, followed by biotin pull-down to enrich the dU-containing loci for sequencing. Here, we describe a detailed protocol for performing the Detect-seq experiment, and a customized, open-source, bioinformatic pipeline for analyzing the characteristic Detect-seq data is also provided. Unlike those previous whole-genome sequencing-based methods, Detect-seq uses an enrichment strategy and hence is endowed with great sensitivity, a higher signal-to-noise ratio and no requirement for high sequencing depth. Furthermore, Detect-seq is widely applicable for both mitotic and postmitotic biological systems. The entire protocol typically takes 5 d from the genomic DNA extraction to sequencing and ~1 week for data analysis.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Code availability
All custom codes for Detect-seq data analysis are deposited in GitHub (https://github.com/menghaowei/Detect-seq). To make it easier to repeat the data analysis process of Detect-seq, we also provide Snakemake workflow scripts in the GitHub repository.
References
Komor, A. C., Kim, Y. B., Packer, M. S., Zuris, J. A. & Liu, D. R. Programmable editing of a target base in genomic DNA without double-stranded DNA cleavage. Nature 533, 420–424 (2016).
Nishida, K. et al. Targeted nucleotide editing using hybrid prokaryotic and vertebrate adaptive immune systems. Science 353, aaf8729 (2016).
Gaudelli, N. M. et al. Programmable base editing of A•T to G•C in genomic DNA without DNA cleavage. Nature 551, 464–471 (2017).
Rees, H. A. & Liu, D. R. Base editing: precision chemistry on the genome and transcriptome of living cells. Nat. Rev. Genet. 19, 770–788 (2018).
Zhang, F. Development of CRISPR-Cas systems for genome editing and beyond. Q Rev. Biophys. 52, e6 (2019).
Dunbar, C. E. et al. Gene therapy comes of age. Science 359, eaan4672 (2018).
Doudna, J. A. The promise and challenge of therapeutic genome editing. Nature 578, 229–236 (2020).
Rees, H. A. et al. Improving the DNA specificity and applicability of base editing through protein engineering and protein delivery. Nat. Commun. 8, 15790 (2017).
Kim, Y. B. et al. Increasing the genome-targeting scope and precision of base editing with engineered Cas9-cytidine deaminase fusions. Nat. Biotechnol. 35, 371–376 (2017).
Zuo, E. et al. A rationally engineered cytosine base editor retains high on-target activity while reducing both DNA and RNA off-target effects. Nat. Methods 17, 600–604 (2020).
Doman, J. L., Raguram, A., Newby, G. A. & Liu, D. R. Evaluation and minimization of Cas9-independent off-target DNA editing by cytosine base editors. Nat. Biotechnol. 38, 620–628 (2020).
Wang, L. et al. Eliminating base-editor-induced genome-wide and transcriptome-wide off-target mutations. Nat. Cell Biol. 23, 552–563 (2021).
Lei, Z., Meng, H., Zhuang, Y., Zhu, Q. & Yi, C. Chemical and biological approaches to interrogate off-target effects of genome editing tools. ACS Chem. Biol. 18, 205–217 (2023).
Lei, Z. et al. Detect-seq reveals out-of-protospacer editing and target-strand editing by cytosine base editors. Nat. Methods 18, 643–651 (2021).
Lei, Z. et al. Mitochondrial base editor induces substantial nuclear off-target mutations. Nature 606, 804–811 (2022).
Shu, X. et al. Genome-wide mapping reveals that deoxyuridine is enriched in the human centromeric DNA. Nat. Chem. Biol. 14, 680–687 (2018).
Zhu, C. et al. Single-cell 5-formylcytosine landscapes of mammalian early embryos and ESCs at single-base resolution. Cell Stem Cell 20, 720–731.e5 (2017).
Zeng, H. et al. Bisulfite-free, nanoscale analysis of 5-hydroxymethylcytosine at single base resolution. J. Am. Chem. Soc. 140, 13190–13194 (2018).
Xia, B. et al. Bisulfite-free, base-resolution analysis of 5-formylcytosine at the genome scale. Nat. Methods 12, 1047–1050 (2015).
Song, C. X. et al. Genome-wide profiling of 5-formylcytosine reveals its roles in epigenetic priming. Cell 153, 678–691 (2013).
Chen, L. et al. Re-engineering the adenine deaminase TadA-8e for efficient and specific CRISPR-based cytosine base editing. Nat. Biotechnol. In press https://doi.org/10.1038/s41587-022-01532-7 (2022).
Siriwardena, S. U., Chen, K. & Bhagwat, A. S. Functions and malfunctions of mammalian DNA-cytosine deaminases. Chem. Rev. 116, 12688–12710 (2016).
Matsumoto, Y. et al. Up-regulation of activation-induced cytidine deaminase causes genetic aberrations at the CDKN2b-CDKN2a in gastric cancer. Gastroenterology 139, 1984–1994 (2010).
Saraconi, G., Severi, F., Sala, C., Mattiuz, G. & Conticello, S. G. The RNA editing enzyme APOBEC1 induces somatic mutations and a compatible mutational signature is present in esophageal adenocarcinomas. Genome Biol. 15, 417 (2014).
Burns, M. B., Temiz, N. A. & Harris, R. S. Evidence for APOBEC3B mutagenesis in multiple human cancers. Nat. Genet. 45, 977–983 (2013).
Bae, S., Park, J. & Kim, J. S. Cas-OFFinder: a fast and versatile algorithm that searches for potential off-target sites of Cas9 RNA-guided endonucleases. Bioinformatics 30, 1473–1475 (2014).
Concordet, J. P. & Haeussler, M. CRISPOR: intuitive guide selection for CRISPR/Cas9 genome editing experiments and screens. Nucleic Acids Res. 46, W242–W245 (2018).
Montague, T. G., Cruz, J. M., Gagnon, J. A., Church, G. M. & Valen, E. CHOPCHOP: a CRISPR/Cas9 and TALEN web tool for genome editing. Nucleic Acids Res. 42, W401–W407 (2014).
Kim, D. et al. Genome-wide target specificities of CRISPR RNA-guided programmable deaminases. Nat. Biotechnol. 35, 475–480 (2017).
McGrath, E. et al. Targeting specificity of APOBEC-based cytosine base editor in human iPSCs determined by whole genome sequencing. Nat. Commun. 10, 5353 (2019).
Jin, S. et al. Cytosine, but not adenine, base editors induce genome-wide off-target mutations in rice. Science 364, 292–295 (2019).
Zuo, E. et al. Cytosine base editor generates substantial off-target single-nucleotide variants in mouse embryos. Science 364, 289–292 (2019).
Haeussler, M. et al. Evaluation of off-target and on-target scoring algorithms and integration into the guide RNA selection tool CRISPOR. Genome Biol. 17, 148 (2016).
Kim, D., Kang, B. C. & Kim, J. S. Identifying genome-wide off-target sites of CRISPR RNA-guided nucleases and deaminases with Digenome-seq. Nat. Protoc. 16, 1170–1192 (2021).
Wei, Y. et al. Mitochondrial base editor DdCBE causes substantial DNA off-target editing in nuclear genome of embryos. Cell Discov. 8, 27 (2022).
Tao, J., Bauer, D. E. & Chiarle, R. Assessing and advancing the safety of CRISPR-Cas tools: from DNA to RNA editing. Nat. Commun. 14, 212 (2023).
Liu, Y. et al. Bisulfite-free direct detection of 5-methylcytosine and 5-hydroxymethylcytosine at base resolution. Nat. Biotechnol. 37, 424–429 (2019).
Zhang, Y. et al. Model-based analysis of ChIP-Seq (MACS). Genome Biol. 9, R137 (2008).
Livak, K. J. & Schmittgen, T. D. Analysis of relative gene expression data using real-time quantitative PCR and the 2−ΔΔCT Method. Methods 25, 402–408 (2001).
Acknowledgements
Bioinformatics analysis was performed on the High-Performance Computing Platform of the School of Life Sciences and High-Performance Computing Platform of the Center for Life Science. This work was supported by the National Key R&D Program (2019YFA0110900 and 2019YFA0802200), National Natural Science Foundation of China (nos. 21825701, 91953201, 92153303 and 22107006) and China Postdoctoral Science Foundation (2021M700238).
Author information
Authors and Affiliations
Contributions
Z.L., H.M. and C.Y. conceived and led the research. Z.L, H.M. and C.Y. wrote the manuscript with the help of X.R. and H.Z.
Corresponding author
Ethics declarations
Competing interests
Peking University has filed patent applications on Detect-seq described in this study, listing Z.L., H.M. and C.Y. as inventors.
Peer review
Peer review information
Nature Protocols thanks Sangsu Bae, Hui Yang and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Related links
Key references using this protocol
Lei, Z. et al. Nat. Methods 18, 643–651 (2021): https://doi.org/10.1038/s41592-021-01172-w
Lei, Z. et al. Nature 606, 804–811 (2022): https://doi.org/10.1038/s41586-022-04836-5
Supplementary information
Supplementary Table 1
Spike-in model sequence and qPCR primer sequence
Supplementary Table 2
An output example for the enrichment significance test results; this file is related to Fig. 4 and Step 51
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Lei, Z., Meng, H., Rao, X. et al. Detect-seq, a chemical labeling and biotin pull-down approach for the unbiased and genome-wide off-target evaluation of programmable cytosine base editors. Nat Protoc 18, 2221–2255 (2023). https://doi.org/10.1038/s41596-023-00837-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41596-023-00837-4
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.