Abstract
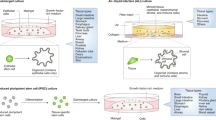
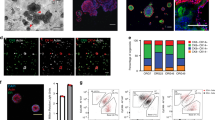
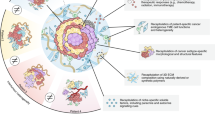
Organoid technology has revolutionized the study of human organ development, disease and therapy response tailored to the individual. Although detailed protocols are available for the generation and long-term propagation of human organoids from various organs, such methods are lacking for breast tissue. Here we provide an optimized, highly versatile protocol for long-term culture of organoids derived from either normal human breast tissues or breast cancer (BC) tissues, as well as culturing conditions for a panel of 45 biobanked samples, including BC organoids covering all major disease subtypes (triple-negative, estrogen receptor-positive/progesterone receptor-positive and human epidermal growth receptor 2-positive). Additionally, we provide methods for genetic manipulation by Lipofectamine 2000, electroporation or lentivirus and subsequent organoid selection and clonal culture. Finally, we introduce an optimized method for orthotopic organoid transplantation in mice, which includes injection of organoids and estrogen pellets without the need for surgery. Organoid derivation from tissue fragments until the first split takes 7–21 d; generation of genetically manipulated clonal organoid cultures takes 14–21 d; and organoid expansion for xenotransplantation takes >4 weeks.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
All data described in this protocol are included in the article figures or were published previously in the supporting primary research papers (https://doi.org/10.1016/j.cell.2017.11.010, https://doi.org/10.1038/s41467-020-15548-7 and https://doi.org/10.1038/s41596-019-0160-8).
References
DeSantis, C. E. et al. Breast cancer statistics, 2019. CA Cancer J. Clin. 69, 438–451 (2019).
Nik-Zainal, S. et al. Landscape of somatic mutations in 560 breast cancer whole-genome sequences. Nature 534, 47–54 (2016).
Curtis, C. et al. The genomic and transcriptomic architecture of 2,000 breast tumours reveals novel subgroups. Nature 486, 346–352 (2012).
Rios, A. C. et al. Intraclonal plasticity in mammary tumors revealed through large-scale single-cell resolution 3D imaging. Cancer Cell 35, 618–632 (2019).
Hoon Tan, P. et al The 2019 World Health Organization classification of tumours of the breast. Histopathology 77, 181–185 (2020).
Sato, T. et al. Single Lgr5 stem cells build crypt-villus structures in vitro without a mesenchymal niche. Nature 459, 262–265 (2009).
Sachs, N. et al. A living biobank of breast cancer organoids captures disease heterogeneity. Cell 172, 373–386 (2018).
Rosenbluth, J. M. et al. Organoid cultures from normal and cancer-prone human breast tissues preserve complex epithelial lineages. Nat. Commun. 11, 1711 (2020).
Dekkers, J. F. et al. Modeling breast cancer using CRISPR–Cas9-mediated engineering of human breast organoids. J. Natl. Cancer Inst 112, 540–544 (2020).
Kedrin, D. et al. Intravital imaging of metastatic behavior through a mammary imaging window. Nat. Methods 5, 1019–1021 (2008).
Holliday, D. L. & Speirs, V. Choosing the right cell line for breast cancer research. Breast Cancer Res. 13, 76–99 (2011).
Kapałczyńska, M. et al. 2D and 3D cell cultures—a comparison of different types of cancer cell cultures. Arch. Med. Sci. 14, 910–919 (2018).
Vargo-Gogola, T. & Rosen, J. M. Modelling breast cancer: one size does not fit all. Nat. Rev. Cancer 7, 659–672 (2007).
Neve, R. M. et al. A collection of breast cancer cell lines for the study of functionally distinct cancer subtypes. Cancer Cell 10, 515–527 (2006).
Miller, D. H., Sokol, E. S. & Gupta, P. B. 3D primary culture model to study human mammary development. Methods Mol. Biol. 1612, 139–147 (2017).
Linnemann, J. R., Meixner, L. K., Miura, H. & Scheel, C. H. An organotypic 3D assay for primary human mammary epithelial cells that recapitulates branching morphogenesis. Methods Mol. Biol. 1612, 125–137 (2017).
Lim, E. et al. Aberrant luminal progenitors as the candidate target population for basal tumor development in BRCA1 mutation carriers. Nat. Med. 15, 907–913 (2009).
Kopper, O. et al. An organoid platform for ovarian cancer captures intra- and interpatient heterogeneity. Nat. Med. 25, 838–849 (2019).
Cattaneo, C. M. et al. Tumor organoid–T-cell coculture systems. Nat. Protoc. 15, 15–39 (2020).
Drost, J. et al. Organoid culture systems for prostate epithelial and cancer tissue. Nat. Protoc. 11, 347–358 (2016).
Fujii, M., Matano, M., Nanki, K. & Sato, T. Efficient genetic engineering of human intestinal organoids using electroporation. Nat. Protoc. 10, 1474–1485 (2015).
Broutier, L. et al. Culture and establishment of self-renewing human and mouse adult liver and pancreas 3D organoids and their genetic manipulation. Nat. Protoc. 11, 1724–1743 (2016).
Drost, J. et al. Sequential cancer mutations in cultured human intestinal stem cells. Nature 521, 43–47 (2015).
Dall, G., Vieusseux, J., Unsworth, A., Anderson, R. & Britt, K. Low dose, low cost estradiol pellets can support MCF-7 tumour growth in nude mice without bladder symptoms. J. Cancer 6, 1331–1336 (2015).
Dobrolecki, L. E. et al. Patient-derived xenograft (PDX) models in basic and translational breast cancer research. Cancer Metastasis Rev. 35, 547–573 (2016).
Gakhar, G., Wight-Carter, M., Andrews, G., Olson, S. & Nguyen, T. A. Hydronephrosis and urine retention in estrogen-implanted athymic nude mice. Vet. Pathol. 46, 505–508 (2009).
Whittle, J. R. et al. Dual targeting of CDK4/6 and BCL2 pathways augments tumor response in estrogen receptor positive breast cancer. Clin. Cancer Res. 26, 4120–4134 (2020).
Acknowledgements
We are very grateful for the technical support of the Princess Máxima Center for Pediatric Oncology, the Hubrecht Institute, the Foundation Hubrecht Organoid Technology (HUB), the Dana-Farber Cancer Institute and the Walter and Eliza Hall Institute of Medical Research for generation of the described methods and the HUB for managing the breast organoid biobank. This work was financially supported by the Princess Máxima Center for Pediatric Oncology. J.F.D. was supported by a Marie Curie Global Fellowship and a VENI grant from the Dutch Organization for Scientific Research. J.M.R. and J.S.B. were supported by an R35 grant (CA242428) from the National Cancer Institute and acknowledge the support of the Susan G. Komen Breast Cancer Foundation. J.M.R. was supported by the American Cancer Society. J.E.V. was supported by the Australian National Health and Medical Research Council. A.C.R. was supported by a Starting Grant 2019 (project 804412) from the European Research Council.
Author information
Authors and Affiliations
Contributions
J.F.D., E.J.V., N.S., J.M.R., O.K., H.G.R. and C.P. performed experiments and developed the protocol. J.F.D., E.J.V., N.S., J.M.R., H.G.R., C.S.V. and S.F.B. generated figures. J.F.D. and E.J.V. conceptualized and wrote the manuscript. H.G.R., E.J.W., C.S.V. and S.F.B. co-wrote the manuscript. J.E.V., C.S.V., S.F.B., J.S.B., H.C. and A.C.R. supervised the work.
Corresponding author
Ethics declarations
Competing interests
J.F.D., N.S., S.F.B., H.C. and A.C.R. are inventors on patents related to the organoid technology (WO201309812-A3/WO2016083612-Al/P309013GB).
Additional information
Peer review information Nature Protocols thanks the anonymous reviewers for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Related links
Key references using this protocol:
Sachs, N. et al. Cell 172, E10 (2018): https://doi.org/10.1016/j.cell.2017.11.010
Rosenbluth, J. M. et al. Nat. Commun. 11, 1171 (2020): https://doi.org/10.1038/s41467-020-15548-7
Dekkers, J. F. et al. J. Natl Cancer Inst. 112, djz196 (2019): https://doi.org/10.1038/s41596-019-0160-8
Supplementary Information
Rights and permissions
About this article
Cite this article
Dekkers, J.F., van Vliet, E.J., Sachs, N. et al. Long-term culture, genetic manipulation and xenotransplantation of human normal and breast cancer organoids. Nat Protoc 16, 1936–1965 (2021). https://doi.org/10.1038/s41596-020-00474-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41596-020-00474-1
This article is cited by
-
Predictive, preventive, and personalized medicine in breast cancer: targeting the PI3K pathway
Journal of Translational Medicine (2024)
-
Establishing conditions for the generation and maintenance of estrogen receptor-positive organoid models of breast cancer
Breast Cancer Research (2024)
-
One-Pot Biopreparation of Trimetallic ZnO–MgO–CuO Nanoparticles: Enhanced Cytotoxicity, Antibacterial Activities and Molecular Docking Studies
Chemistry Africa (2024)
-
BEHAV3D: a 3D live imaging platform for comprehensive analysis of engineered T cell behavior and tumor response
Nature Protocols (2024)
-
Exploring the interaction between extracellular matrix components in a 3D organoid disease model to replicate the pathophysiology of breast cancer
Journal of Experimental & Clinical Cancer Research (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.