Abstract
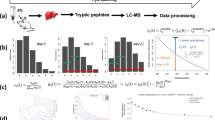
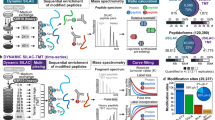
Proteins are continually produced and degraded, to avoid the accumulation of old or damaged molecules and to maintain the efficiency of physiological processes. Despite its importance, protein turnover has been difficult to measure in vivo. Previous approaches to evaluating turnover in vivo have required custom labeling approaches, involved complex mass spectrometry (MS) analyses, or used comparative strategies that do not allow direct quantitative measurements. Here, we describe a robust protocol for quantitative proteome turnover analysis in mice that is based on a commercially available diet for stable isotope labeling of amino acids in mammals (SILAM). We start by discussing fundamental concepts of protein turnover, including different methodological approaches. We then cover in detail the practical aspects of metabolic labeling and explain both the experimental and computational steps that must be taken to obtain accurate in vivo results. Finally, we present a simple experimental workflow that enables measurement of precise turnover rates in a time frame of ~4–5 weeks, including the labeling time. We also provide all the scripts needed for the interpretation of the MS results and for comparing turnover across different conditions. Overall, the workflow presented here comprises several improvements in the determination of protein lifetimes with respect to other available methods, including a minimally invasive labeling strategy and a robust interpretation of MS results, thus enhancing reproducibility across laboratories.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout













Similar content being viewed by others
Data and code availability
The dataset presented in this protocol was originally generated in ref. 4. All data are available from the corresponding authors on reasonable request. All code from Supplementary Data 1 is also publicly available at GitHub: https://github.com/malevra/protein-turnover.
References
Labbadia, J. & Morimoto, R. I. The biology of proteostasis in aging and disease. Annu. Rev. Biochem. 84, 435–464 (2015).
Ong, S.-E. & Mann, M. A practical recipe for stable isotope labeling by amino acids in cell culture (SILAC). Nat. Protoc. 1, 2650–2660 (2006).
Schoenheimer, R. The Dynamic State of Body Constituents (Harvard University Press, 1946).
Fornasiero, E. F. et al. Precisely measured protein lifetimes in the mouse brain reveal differences across tissues and subcellular fractions. Nat. Commun. 9, 4230 (2018).
Chan, X. C. Y., Black, C. M., Lin, A. J., Ping, P. & Lau, E. Mitochondrial protein turnover: Methods to measure turnover rates on a large scale. J. Mol. Cell. Cardiol. 78, 54–61 (2015).
Busch, R. et al. Measurement of protein turnover rates by heavy water labeling of nonessential amino acids. Biochim. Biophys. Acta 1760, 730–744 (2006).
Price, J. C., Guan, S., Burlingame, A., Prusiner, S. B. & Ghaemmaghami, S. Analysis of proteome dynamics in the mouse brain. Proc. Natl. Acad. Sci. USA 107, 14508–14513 (2010).
McClatchy, D. B., Dong, M., Wu, C. C., Venable, J. D. & Yates, J. R. 15N metabolic labeling of mammalian tissue with slow protein turnover. J. Proteome Res. 6, 2005–2010 (2007).
Savas, J. N., Toyama, B. H., Xu, T., Yates, J. R. & Hetzer, M. W. Extremely long-lived nuclear pore proteins in the rat brain. Science 335, 942 (2012).
Toyama, B. H. et al. Identification of long-lived proteins reveals exceptional stability of essential cellular structures. Cell 154, 971–982 (2013).
Heo, S. et al. Identification of long-lived synaptic proteins by proteomic analysis of synaptosome protein turnover. Proc. Natl Acad. Sci. USA 115, E3827–E3836 (2018).
Claydon, A. J. & Beynon, R. J. Proteome dynamics: revisiting turnover with a global perspective. Mol. Cell. Proteom. 11, 1551–1565 (2012).
Basisty, N., Meyer, J. G. & Schilling, B. Protein turnover in aging and longevity. Proteomics 18, e1700108 (2018).
Claydon, A. J. et al. Heterogenous turnover of sperm and seminal vesicle proteins in the mouse revealed by dynamic metabolic labeling. Mol. Cell. Proteom. 11, M111.014993 (2012).
Claydon, A. J., Thom, M. D., Hurst, J. L. & Beynon, R. J. Protein turnover: measurement of proteome dynamics by whole animal metabolic labelling with stable isotope labelled amino acids. Proteomics 12, 1194–1206 (2012).
Guan, S., Price, J. C., Ghaemmaghami, S., Prusiner, S. B. & Burlingame, A. L. Compartment modeling for mammalian protein turnover studies by stable isotope metabolic labeling. Anal. Chem. 84, 4014–4021 (2012).
Zhang, Y. et al. Proteome scale turnover analysis in live animals using stable isotope metabolic labeling. Anal. Chem. 83, 1665–1672 (2011).
Vogt, J. A. et al. Determination of fractional synthesis rates of mouse hepatic proteins via metabolic 13C-labeling, MALDI-TOF MS and analysis of relative isotopologue abundances using average masses. Anal. Chem. 77, 2034–2042 (2005).
Rahman, M., Previs, S. F., Kasumov, T. & Sadygov, R. G. Gaussian process modeling of protein turnover. J. Proteome Res. 15, 2115–2122 (2016).
Toyama, B. H. et al. Visualization of long-lived proteins reveals age mosaicism within nuclei of postmitotic cells. J. Cell Biol. 218, 1–16 (2018).
Lau, E. et al. A large dataset of protein dynamics in the mammalian heart proteome. Sci. Data 3, 1–15 (2016).
Sadygov, R. G. et al. d2ome, software for in vivo protein turnover analysis using heavy water labeling and LC-MS, reveals alterations of hepatic proteome dynamics in a mouse model of NAFLD. J. Proteome Res. 17, 3740–3748 (2018).
Naylor, B. C. et al. DeuteRater: a tool for quantifying peptide isotope precision and kinetic proteomics. Bioinformatics 33, 1514–1520 (2017).
Dufner, D. & Previs, S. F. Measuring in vivo metabolism using heavy water. Curr. Opin. Clin. Nutr. Metab. Care 6, 511–517 (2003).
Krüger, M. et al. SILAC mouse for quantitative proteomics uncovers kindlin-3 as an essential factor for red blood cell function. Cell 134, 353–364 (2008).
Chen, X., Wei, S., Ji, Y., Guo, X. & Yang, F. Quantitative proteomics using SILAC: principles, applications, and developments. Proteomics 15, 3175–3192 (2015).
Hanke, S., Besir, H., Oesterhelt, D. & Mann, M. Absolute SILAC for accurate quantitation of proteins in complex mixtures down to the attomole level. J. Proteome Res. 7, 1118–1130 (2008).
Zanivan, S., Krueger, M. & Mann, M. In vivo quantitative proteomics: the SILAC mouse. Methods Mol. Biol. 757, 435–450 (2012).
Pena, I. A. et al. Mouse lysine catabolism to aminoadipate occurs primarily through the saccharopine pathway; implications for pyridoxine dependent epilepsy (PDE). Biochim. Biophys. Acta 1863, 121–128 (2017).
Mandad, S. et al. The codon sequences predict protein lifetimes and other parameters of the protein life cycle in the mouse brain. Sci. Rep. 8, 16913 (2018).
Cox, J. et al. Accurate proteome-wide label-free quantification by delayed normalization and maximal peptide ratio extraction, termed MaxLFQ. Mol. Cell. Proteomics 13, 2513–2526 (2014).
Phillip, Y. & Schreiber, G. Formation of protein complexes in crowded environments-from in vitro to in vivo. FEBS Lett. 587, 1046–1052 (2013).
Venable, J. D., Dong, M.-Q., Wohlschlegel, J., Dillin, A. & Yates, J. R. Automated approach for quantitative analysis of complex peptide mixtures from tandem mass spectra. Nat. Methods 1, 39–45 (2004).
Collins, B. C. et al. Multi-laboratory assessment of reproducibility, qualitative and quantitative performance of SWATH-mass spectrometry. Nat. Commun. 8, 1–11 (2017).
Meier, F., Geyer, P. E., Virreira Winter, S., Cox, J. & Mann, M. BoxCar acquisition method enables single-shot proteomics at a depth of 10,000 proteins in 100 minutes. Nat. Methods 15, 440–448 (2018).
McShane, E. et al. Kinetic analysis of protein stability reveals age-dependent degradation. Cell 167, 803–815 (2016).
Masters, P. M., Bada, J. L. & Zigler, J. S. Aspartic acid racemisation in the human lens during ageing and in cataract formation. Nature 268, 71–73 (1977).
John, A. M. & Bell, J. M. Amino acid requirements of the growing mouse. J. Nutr. 106, 1361–1367 (1976).
Overmyer, K. A. et al. Multiplexed proteome analysis with neutron-encoded stable isotope labeling in cells and mice. Nat. Protoc. 13, 293–306 (2018).
Shevchenko, A., Tomas, H., Havlis, J., Olsen, J. V. & Mann, M. In-gel digestion for mass spectrometric characterization of proteins and proteomes. Nat. Protoc. 1, 2856–2860 (2006).
Schirle, M., Heurtier, M.-A. & Kuster, B. Profiling core proteomes of human cell lines by one-dimensional PAGE and liquid chromatography-tandem mass spectrometry. Mol. Cell. Proteom. 2, 1297–1305 (2003).
Batth, T. S., Francavilla, C. & Olsen, J. V. Off-line high-pH reversed-phase fractionation for in-depth phosphoproteomics. J. Proteome Res. 13, 6176–6186 (2014).
Lam, M. P. Y. et al. Protein kinetic signatures of the remodeling heart following isoproterenol stimulation. J. Clin. Invest. 124, 1734–1744 (2014).
Ahmed, S., Holt, M., Riedel, D. & Jahn, R. Small-scale isolation of synaptic vesicles from mammalian brain. Nat. Protoc. 8, 998–1009 (2013).
Dunkley, P. R., Jarvie, P. E. & Robinson, P. J. A rapid Percoll gradient procedure for preparation of synaptosomes. Nat. Protoc. 3, 1718–1728 (2008).
Sims, N. R. & Anderson, M. F. Isolation of mitochondria from rat brain using Percoll density gradient centrifugation. Nat. Protoc. 3, 1228–1239 (2008).
Cox, B. & Emili, A. Tissue subcellular fractionation and protein extraction for use in mass-spectrometry-based proteomics. Nat. Protoc. 1, 1872–1878 (2006).
Alvarez-Castelao, B., Schanzenbächer, C. T., Langer, J. D. & Schuman, E. M. Cell-type-specific metabolic labeling, detection and identification of nascent proteomes in vivo. Nat. Protoc. 14, 556–575 (2019).
Tu, R. Comparison between confidence intervals of linear regression models with and without restrication. Commun. Stat. Theory Methods 28, 2879–2898 (1999).
Feil, R., Brocard, J., Mascrez, B. & Lemeur, M. Ligand-activated site-specific recombination in mice. Proc. Natl. Acad. Sci. USA 93, 10887–10890 (1996).
Erdmann, G., Schütz, G. & Berger, S. Inducible gene inactivation in neurons of the adult mouse forebrain. BMC Neurosci. 8, 63 (2007).
Gage, G. J., Kipke, D. R. & Shain, W. Whole animal perfusion fixation for rodents. J. Vis. Exp. 65, 3564 (2012).
Cox, J. & Mann, M. MaxQuant enables high peptide identification rates, individualized p.p.b.-range mass accuracies and proteome-wide protein quantification. Nat. Biotechnol. 26, 1367–1372 (2008).
Cox, J. et al. Andromeda: a peptide search engine integrated into the MaxQuant environment. J. Proteome Res. 10, 1794–1805 (2011).
Tyanova, S., Temu, T. & Cox, J. The MaxQuant computational platform for mass spectrometry-based shotgun proteomics. Nat. Protoc. 11, 2301–2319 (2016).
Acknowledgements
We thank S. Truckenbrodt (IST, Austria) for initial suggestions on manuscript preparation. We thank S. Reshetniak (University Medical Center Göttingen), T. Dankovich (University Medical Center Göttingen) and I. Atanassov (Max Planck Institute for Biology of Ageing, Germany) for carefully reading and commenting on the final version of the manuscript. We thank U. Plessmann (University Medical Center Göttingen) for setting up and maintaining the LC–MS configuration. The work was supported by grants to S. O. R. from the European Research Council (ERC-2013-CoG NeuroMolAnatomy) and the Deutsche Forschungsgemeinschaft (DFG 1967/7-1, SFB1286/A3, ExC 2067/Multiscale Bioimaging) and to HU from the SFB-1286/A10.
Author information
Authors and Affiliations
Contributions
Conceptualization: E.F.F. and S.O.R. Methodology: E.F.F., S.M., T.I. and H.U. Software: M.A. Formal analysis: M.A., E.F.F. and S.O.R. Resources and funding acquisition: S.O.R. Writing—initial draft: E.F.F. and S.O.R. Writing—final draft, review and editing: M.A., S.M., T.I., H.U., S.O.R. and E.F.F. Supervision: E.F.F.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Related link
Key references using this protocol
Fornasiero, E. F. et al. Nat. Commun. 9, 4230 (2018): https://www.nature.com/articles/s41467-018-06519-0
Supplementary information
Supplementary Information
Supplementary Tables 1 and 2
Supplementary Data 1
MATLAB scripts, also available online at GitHub
Supplementary Data 2
Testing dataset for evaluation of the MATLAB scripts
Supplementary Data 3
Simulated dataset for evaluation of the MATLAB scripts (pools)
Rights and permissions
About this article
Cite this article
Alevra, M., Mandad, S., Ischebeck, T. et al. A mass spectrometry workflow for measuring protein turnover rates in vivo. Nat Protoc 14, 3333–3365 (2019). https://doi.org/10.1038/s41596-019-0222-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41596-019-0222-y
This article is cited by
-
NanoSIMS observations of mouse retinal cells reveal strict metabolic controls on nitrogen turnover
BMC Molecular and Cell Biology (2021)
-
An atlas of protein turnover rates in mouse tissues
Nature Communications (2021)
-
Understanding the “SMART” features of hematopoietic stem cells and beyond
Science China Life Sciences (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.