Abstract
Dynamic protein molecules are defined by their spatiotemporal characteristics and should thus be represented by models incoporating both characteritics. Structural biology enables determination of atomic structures of individual conformational states of a given protein. Obtaining the complementary temporal information of a given time resolution, which can be directly linked to the corresponding atomic structures, requires identifying at each time point the specific conformational state adopted by the protein. Here, we examine individual regulator of conductance to K+ (RCK) domains in the regulatory module of the MthK channel by monitoring in real time the orientation of an α-helix that is conformational-state-specific. The acquired dynamic information that specifies an RCK domain’s multi-state conformational changes, combined with already available corresponding atomic structures, enables us to establish an experiment-based spatiotemporal representation of an RCK domain, and to deduce a quantitative mechanistic model of the channel.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
Data and materials described here will be made available upon reasonable request.
References
Coontz, R., Fahrenkamp-Uppenbrink, J., Lavine, M. & Vinson, V. Going from strength to strength. Science 343, 1091 (2014).
Murata, K. & Wolf, M. Cryo-electron miscroscopy for strcutural analysis of dynamic biological macromolecules. Biochim. Biophys Acta 1862, 324–334 (2018).
Stryer, L. Fluorescence energy transfer as a spectroscopic ruler. Ann. Rev. Biochem. 47, 819–846 (1978).
Lewis, J. H. & Lu, Z. Resolution of ångström-scale protein-conformational changes by analyzing fluorescence anisotropy. Nat. Struct. Mol. Biol. https://doi.org/10.1038/s41594-019-0274-2 (2019).
Lewis, J. H. & Lu, Z. Energetics of ångström-scale conformational changes in an RCK domain of the MthK K+ channel. Nat. Struct. Mol. Biol. https://doi.org/10.1038/s41594-019-0275-1 (2019).
Jiang, Y. et al. Crystal structure and mechanism of a calcium-gated potassium channel. Nature 417, 515–522 (2002).
Ye, S., Li, Y., Chen, L. & Jiang, Y. Crystal structures of a ligand-free MthK gating ring: insights into the ligand gating mechanism of K+ channels. Cell 126, 1161–1173 (2006).
Zadek, B. & Nimigean, C. M. Calcium-dependent gating of MthK, a prokaryotic potassium channel. J. Gen. Physiol. 127, 673–685 (2006).
Pau, V. P., Barca-Heidemann, K. & Rothberg, B. S. Allosteric mechanism of Ca2+ activation and H+-inhibited gating of the MthK K+ channel. J. Gen. Physiol. 135, 509–526 (2010).
Sakmann, B. & Neher, E. Single-Channel Recording (Plenum Press, 1995).
Miller, C. Ion Channel Reconstitution (Plenum, 1986).
Lewis, J. H., Jamiolkoski, R. M., Woody, E., Ostap, E. M. & Goldman, Y. E. Deconvolution of camera instrument response functions. Biophys J. 112, 1214–1220 (2017).
Pau, V. P. et al. Structure and function of multiple Ca2+-binding sites in a K+ channel regulator of K+ conductance (RCK) domain. Proc. Natl Acad. Sci. USA 108, 17684–17689 (2011).
Zhou, Y., Morais-Cabral, J. H., Kaufman, A. & MacKinnon, R. Chemistry of ion coordination and hydration revealed by a K+ channel-Fab complex at 2.0 A resolution. Nature 414, 43–48 (2001).
Acknowledgements
We thank Y. Zhou for technical support, Y. Jiang and R. MacKinnon for providing the complementary DNA of MthK, V. Pau and B. Rothberg for sharing their published data for comparison and P. De Weer, T. Hoshi and B. Salzberg for critiques of our manuscript at different stages of its development. This study was supported by the grant no. GM055560 from the National Institute of General Medical Sciences of the National Institutes of Health to Z.L.
Author information
Authors and Affiliations
Contributions
J.H.L. and Z.L. designed the study. J.H.L. performed experiments, developed analytical tools, and analyzed the data, with the input from Z.L. J.H.L. and Z.L. interpreted the results and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information: Ines Chen was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1 Model for the whole MthK channel.
The gate (G) and the regulatory module (RM) both are either in an open state (o) or in a closed (c) state; RM can exist in aRM (left box) or bRM (right box). All-S2 species with or without Ca2+ bound in aRM should be excluded from the closed state of the regulatory module, but it was not explicitly done for graphic simplicity.
Supplementary Figure 2 Illustration of a single cycle of RCK’s conformational changes.
This cycle is defined by the beginning time point of one S2 event to the beginning of the next S2 event. The lifetime of S2 itself is denoted by τ2 (equation 17 in its reciprocal form). The interval between the end of the preceding S2 event and the beginning of the trailing S2 event is denoted by τ2→2 (Eq. 18), during which RCK oscillates between S1 and S3.
Supplementary information
Supplementary Information
Supplementary Figs. 1–3, Supplementary Tables 1–4 and Supplementary Notes 1–6.
Supplementary Video 1
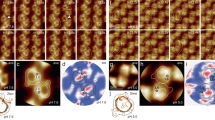
Real-time video of polarized-fluorescence intensities, fluorophore dipole orientations and conformational changes of a tracked RCK domain. Intensities I0, I45, I90 and I135 (leftmost panel) recorded from the particle described in Fig. 1, their individual integrations (left-middle), angles θ, φ and Ω calculated from these intensities (right-middle) and the crystal structure of the corresponding conformational state (rightmost) are displayed point-by-point in real time and at a frame rate of 50 per second. The structures of the tracked RCK are exhibited in the form of electron density against the backdrop of the shadow of the regulatory module comprising eight RCK domains (PDB 1LNQ and 2FY8). States S1, S2 and S3 are colored yellow, blue and orange, respectively.
Supplementary Video 2
Visual illustration of the spatiotemporal model of an isolated regulatory module in the form of video played in real time and at a frame rate of 50 per second. Eight individual RCK domains independently transition among three conformations for the conditions of 0, 0.5 (near EC50) or 50 mM Ca2+ (near saturation), in accordance with temporal templates as exemplified for the Ca2+-free condition in Fig. 4a. The structures of individual conformations are exhibited in the form of a ribbon model overlaid on the space-filling model, and in a side view (left panel) and a view looking down the central axis of the regulatory module (right) (PDB 1LNQ and 2FY8). States S1, S2 and S3 are colored yellow, blue and orange, respectively. The single Ca2+ ion per RCK domain is represented by a cyan sphere.
Supplementary Video 3
Visual illustration of the spatiotemporal model of a whole channel near the EC50 of Ca2+ (1 mM) in the form of video played in real time and at a frame rate of 50 per second. The channel gate of the pore and RCK domains in the regulatory module transition among different conformations in accordance with the temporal templates shown on the left (Fig. 5d), where those rightmost points in the templates appear in real time. In the trace of the gate of the pore (top panel), the open sand closed states are colored chartreuse and maroon, respectively. The structures are simultaneously exhibited in a side view (middle panel) and a view looking down the central axis (right), in which the structure of the pore in the open state is MthK’s but that in the closed state is KcsA’s (PDB 1K4C, 2FY8 and 3RBZ). RCK’s states S1, S2 and S3 are colored yellow, blue and orange, respectively, regardless of whether the regulatory module is in configuration a or b. When all RCKs adopt S2, the pore transitions from the closed state (colored light blue) to the open state (dark blue). Ca2+ ions are presented by cyan spheres.
Supplementary Video 4
Visual illustration of the spatiotemporal model of a whole channel for three Ca2+ conditions in the form of video played in real time and at a frame rate of 50 per second. The Ca2+ concentrations are set to zero, near EC50 (1 mM) or near saturation (50 mM) in terms of the channel’s po. The structures are simultaneously displayed in a side view (left panel) and a view looking down the central axis (right), in which the structure of the pore in the open state is MthK’s but that in the closed state is KcsA’s (PDB 1K4C, 2FY8 and 3RBZ). The gate and RCK domains transition among different conformations in accordance with temporal templates, created as described for the 1 mM Ca2+ condition (Fig. 5d). RCK’s states S1, S2 and S3 are colored yellow, blue and orange, respectively, regardless of whether the regulatory module is in configuration a or b. When all RCK domains adopt S2, the pore proceeds to transition from the closed state (colored light blue) to the open state (dark blue). Ca2+ ions are presented by cyan spheres.
Rights and permissions
About this article
Cite this article
Lewis, J.H., Lu, Z. Integrating spatiotemporal features of a ligand-regulated, multi-state allosteric protein. Nat Struct Mol Biol 26, 816–822 (2019). https://doi.org/10.1038/s41594-019-0276-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41594-019-0276-0
This article is cited by
-
Energetics of ångström-scale conformational changes in an RCK domain of the MthK K+ channel
Nature Structural & Molecular Biology (2019)
-
Resolution of ångström-scale protein conformational changes by analyzing fluorescence anisotropy
Nature Structural & Molecular Biology (2019)