Abstract
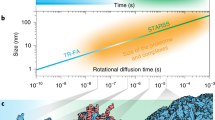
Conformational changes within typical protein molecules are rapid and small, making their quantitative resolution challenging. These changes generally involve rotational motions and may thus be resolved by determining changes in the orientation of a fluorescent label that assumes a unique orientation in each conformation. Here, by analyzing fluorescence intensities collected using a polarization microscope at a rate of 50 frames per second, we follow the changes of 10−16° in the orientation of a single bifunctional rhodamine molecule attached to a regulator of conductance to K+ (RCK) domain of the MthK channel, and thus, the transitions between its three conformational states, with effective standard deviation (σ) of 2−5°. Based on available crystal structures, the position of the fluorophore’s center differs by 3.4−8.1 Å among the states. Thus, the present approach allows the resolution of protein conformational changes involving ångström-scale displacements.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
Data and materials described here will be made available upon reasonable request.
Code availability
Code used in the work will be made available upon reasonable request.
References
Brocchieri, L. & Karlin, S. Protein length in eukaryotic and prokaryotic proteomes. Nucleic Acids Res. 33, 3390–3400 (2005).
Erickson, H. P. Size and shape of protein molecules at the nanometer level determined by sedimentation, gel filtration, and electron miscroscopy In: Li, S (Ed.) Biological Procedures Online 32−51 (Springer,: 2009.).
Sase, I., Miyata, H., Ishiwata, S. & Kinosita, K. Jr. Axial rotation of sliding actin filaments revealed by single-fluorophore imaging. Proc. Natl Acad. Sci. USA 94, 5646–5650 (1997).
Warshaw, D. M. et al. Myosin conformational states determined by single fluorophore polarization. Proc. Natl Acad. Sci. USA 95, 8034–8039 (1998).
Ha, T., Glass, J., Enderle, T., Chemla, D. S. & Weiss, S. Hindered rotational diffusion and rotational jumps of single molecules. Phys. Rev. Lett. 80, 2093–2096 (1998).
Adachi, K. et al. Stepping rotation of F1-ATPase visualized through angle-resolved single-fluorophore imaging. Proc. Natl Acad. Sci. USA 97, 7243–7247 (2000).
Sosa, H., Peterman, E. J., Moerner, W. E. & Goldstein, L. S. ADP-induced rocking of the kinesin motor domain revealed by single-molecule fluorescence polarization microscopy. Nat. Struct. Biol. 8, 540–544 (2001).
Forkey, J. N., Quinlan, M. E., Shaw, M. A., Corrie, J. E. & Goldman, Y. E. Three-dimensional structural dynamics of myosin V by single-molecule fluorescence polarization. Nature 422, 399–404 (2003).
Beausang, J. F., Schroeder, H. W. 3rd, Nelson, P. C. & Goldman, Y. E. Twirling of actin by myosins II and V observed via polarized TIRF in a modified gliding assay. Biophys. J. 95, 5820–5831 (2008).
Rosenberg, S. A., Quinlan, M. E., Forkey, J. N. & Goldman, Y. E. Rotational motions of macro-molecules by single-molecule fluorescence microscopy. Acc. Chem. Res. 38, 583–593 (2005).
Forkey, J. N., Quinlan, M. E. & Goldman, Y. E. Measurement of single macromolecule orientation by total internal reflection fluorescence polarization microscopy. Biophys. J. 89, 1261–1271 (2005).
Backlund, M. P. et al. Simultaneous, accurate measurement of the 3D position and orientation of single molecules. Proc. Natl Acad. Sci. USA 109, 19087–19092 (2012).
Zhang, O., Liu, J., Ding, T. & Lew, M. Imaging the three-dimensional orientation and rotational mobility of fluorescent emitters using the Tri-spot point spread function. Appl. Phys. Lett. 113, 031103 (2018).
Jiang, Y. et al. Crystal structure and mechanism of a calcium-gated potassium channel. Nature 417, 515–522 (2002).
Ye, S., Li, Y., Chen, L. & Jiang, Y. Crystal structures of a ligand-free MthK gating ring: insights into the ligand gating mechanism of K+ channels. Cell 126, 1161–1173 (2006).
Brunger, A. T. et al. Crystallography & NMR system: A new software suite for macromolecular structure determination. Acta Crystallogr. D. Biol. Crystallogr. 54, 905–921 (1998).
Fourkas, J. T. Rapid determination of the three-dimensional orientation of single molecules. Opt. Lett. 26, 211–213 (2001).
Ohmachi, M. et al. Fluorescence microscopy for simultaneous observation of 3D orientation and movement and its application to quantum rod-tagged myosin V. Proc. Natl Acad. Sci. USA 109, 5294–5298 (2012).
Lippert, L. G. et al. Angular measurements of the dynein ring reveal a stepping mechanism dependent on a flexible stalk. Proc. Natl Acad. Sci. USA 114, E4564–E4573 (2017).
Axelrod, D. Carbocyanine dye orientation in red cell membrane studied by microscopic fluorescence polarization. Biophys. J. 26, 557–574 (1979).
Chen, J. & Gupta, A. K. On change point detection and estimation. Commun. Stat.− Simul. C. 30, 665–697 (2001).
Beausang, J. F., Goldman, Y. E. & Nelson, P. C. Changepoint analysis for single-molecule polarized total internal reflection fluorescence microscopy experiments. Methods Enzymol. 487, 431–463 (2011).
Press, W. H, Teukolsky, S. A, Vetterling, W. T. & Flannery, B. P. Numerical Recipes: The Art of Scientific Computing (Cambridge University Press, 2007).
Horn, R. Statistical methods for model discrimination. Applications to gating kinetics and permeation of the acetylcholine receptor channel. Biophys. J. 51, 255–263 (1987).
Lewis, J. H. & Lu, Z. Energetics of ångström-scale conformational changes in an RCK domain of the MthK K+ channel. Nat. Struct. Mol. Biol. https://doi.org/10.1038/s41594-019-0275-1 (2019).
Lewis, J. H. & Lu, Z. Integrating spatiotemporal features of a ligand-regulated, multistate allosteric protein. Nat. Struct. Mol. Biol. https://doi.org/10.1038/s41594-019-0276-0 (2019).
Axelrod, D. Total internal reflection fluorescence. Annu. Rev. Biophys. Bioeng. 13, 247–268 (1984).
Acknowledgements
We thank Y. Zhou for technical assistance; Y. Goldman for inspiration and the use of microscopes in his laboratory to test our initial fluorescence-labeled protein samples; Y. Jiang and R. MacKinnon for providing the cDNA encoding MthK; L. Lippert and T. Dadosh for discussions; P. De Weer,T. Hoshi and B. Salzberg for critique of our manuscript at different stages of its development; and E. McCleskey for support. This study was supported by the grant no. GM055560 from the National Institute of General Medical Sciences of the National Institutes of Health to Z.L. During the initial period of the project, Z.L. was supported as an investigator by the Howard Hughes Medical Institute.
Author information
Authors and Affiliations
Contributions
J.H.L and Z.L. designed the study; J.H.L assembled the polarization microscope, performed experiments, developed analytical tools, and analyzed the data with input from Z.L.; J.H.L. and Z.L. interpreted the results and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information: Ines Chen was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1 State transitions detection by changepoint analysis.
a,b. 4 simulated intensity traces (b) and their summation (a), each of which has a transition point at time t = 2.65 sec. c. A cumulative distribution plot of photons collected over time. The slopes of all plots change at t = 2.65 sec, where the rate of photon release transitions from the initial state (red line) to the subsequent state (blue line). d. Plot of the log likelihood ratio over time calculated with Eq. 14 in Methods.
Supplementary Figure 2 State identification by k-means clustering analysis.
The φ angle is plotted against the θ angle, where d1, d2, or d3 is the distance of a given data point to the mean (μ1, μ2, or μ3) of one of the three distributions.
Supplementary Figure 3 Illustration of translational and rotational offsets between imaging planes, and integration of measured intensities from a point source in n different imaging channels.
a,b. Depiction of positions of three particles, i = 1, 2, 3, imaged in the reference channel at positions xi,ref, yi,ref (left) and separately in a compared channel n at positions xi,n, yi,n (right). In each case, the three positions defined the imaging plane. The offsets between each compared channel n and the reference channel was then defined by the translational offsets Δx, Δy (a) and the rotational angles α (around z-axis; a) and γ (around x-axis; b). c. Illustration of the translational and rotational operations relating the corresponding images of individual particles among the four channels. Simulated images of point sources, as described in d, on the EMCCD camera’s four sections (or dubbed channels in the main text) designated for I0, I45, I90 and I135 (left panel); the corresponding images in the four sections, identified on the basis of the translational and rotational operations, are indicated by numbers (right panel). d. Illustration of intensity integration with a simulated point source (left panel). The center of the point source was located at the pixel with the maximum intensity, which corresponded to the actual center here (middle panel). In the right panel, the area, labeled ‘I’, was designated to the data-capturing zone that was set to harbor ~90% of the total recorded intensity signal. This data-capturing zone was separated from the background zone (Bk) by a buffer zone (Bf). The total background intensity was obtained by integrating the pixels within the entire Bk zone.
Supplementary Figure 4 The relations among the directions of fluorophore dipole, the propagation of a photon and its electric field.
a. Light propagates from the dipole D along vector P, with electric field ED perpendicular to P and in the plane defined by P and D. The surface contour represents the probabilities of photon release in space. ω denotes the inclination angle between propagation vector P and dipole D along the z-axis, whereas γ denotes the rotation angle in the x-y plane, with respect to the x-axis. b. Integrating ED over all possible propagation directions defines a net electric field vector, E.
Supplementary Figure 5 Calibration of the microscope.
Light was linearly polarized perpendicular to the optical z-axis, and in the x-y plane with a φ value ranging from 0 to 180° in 5° increments. The light was separated into four polarized components I0, I45, I90 and I135, and recorded in four separate imaging channels of an EMCCD camera. The curves superimposed on the data points are the fits of Eq. 64. The fitted values of ψ, fψ and gψ are summarized in Supplementary Table 1.
Supplementary Figure 6 Illustration of tethered diffusion model of a fluorophore.
In the model, the fluorophore’s motion is constrained to a volume described by a cone of half-angle δ, which effectively reduces the magnitude of the net electric-field vector E, and thus the ratio of Ip over Itot.
Supplementary information
Supplementary Information
Supplementary Figures 1–6, Supplementary Notes 1–6, Supplementary Table 1
Supplementary Video 1
Movie of intensities recorded at four polarization angles from a single rhodamine molecule (boxed in Fig. 2b; analyzed in Fig. 3), attached to one of the eight RCK domain in an isolated regulatory module of the MthK channel. The four intensity components, I0, I45, I90 and I135 from left to right, were simultaneously captured on an EMCCD at a frame rate of 50 per second. The movie was created to display in real time.
Rights and permissions
About this article
Cite this article
Lewis, J.H., Lu, Z. Resolution of ångström-scale protein conformational changes by analyzing fluorescence anisotropy. Nat Struct Mol Biol 26, 802–807 (2019). https://doi.org/10.1038/s41594-019-0274-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41594-019-0274-2
This article is cited by
-
Integrating spatiotemporal features of a ligand-regulated, multi-state allosteric protein
Nature Structural & Molecular Biology (2019)
-
Energetics of ångström-scale conformational changes in an RCK domain of the MthK K+ channel
Nature Structural & Molecular Biology (2019)