Abstract
Tandem DNA repeats vary in the size and sequence of each unit (motif). When expanded, these tandem DNA repeats have been associated with more than 40 monogenic disorders1. Their involvement in disorders with complex genetics is largely unknown, as is the extent of their heterogeneity. Here we investigated the genome-wide characteristics of tandem repeats that had motifs with a length of 2–20 base pairs in 17,231 genomes of families containing individuals with autism spectrum disorder (ASD)2,3 and population control individuals4. We found extensive polymorphism in the size and sequence of motifs. Many of the tandem repeat loci that we detected correlated with cytogenetic fragile sites. At 2,588 loci, gene-associated expansions of tandem repeats that were rare among population control individuals were significantly more prevalent among individuals with ASD than their siblings without ASD, particularly in exons and near splice junctions, and in genes related to the development of the nervous system and cardiovascular system or muscle. Rare tandem repeat expansions had a prevalence of 23.3% in children with ASD compared with 20.7% in children without ASD, which suggests that tandem repeat expansions make a collective contribution to the risk of ASD of 2.6%. These rare tandem repeat expansions included previously undescribed ASD-linked expansions in DMPK and FXN, which are associated with neuromuscular conditions, and in previously unknown loci such as FGF14 and CACNB1. Rare tandem repeat expansions were associated with lower IQ and adaptive ability. Our results show that tandem DNA repeat expansions contribute strongly to the genetic aetiology and phenotypic complexity of ASD.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Data availability
Access to the MSSNG and SSC genome-sequencing data can be obtained by completing data access agreements (https://research.mss.ng and https://www.sfari.org/resource/sfari-base, respectively). The 1000G genome-sequencing data are publicly available via Amazon Web Services (s3://1000genomes/1000G_2504_high_coverage/data). Source data are provided with this paper.
Code availability
Code used in this manuscript is available at GitHub (https://github.com/bjtrost/tandem-repeat-expansions-in-ASD).
References
López Castel, A., Cleary, J. D. & Pearson, C. E. Repeat instability as the basis for human diseases and as a potential target for therapy. Nat. Rev. Mol. Cell Biol. 11, 165–170 (2010).
Yuen, R. K. C. et al. Whole genome sequencing resource identifies 18 new candidate genes for autism spectrum disorder. Nat. Neurosci. 20, 602–611 (2017).
Fischbach, G. D. & Lord, C. The Simons Simplex Collection: a resource for identification of autism genetic risk factors. Neuron 68, 192–195 (2010).
The 1000 Genomes Project Consortium. A global reference for human genetic variation. Nature 526, 68–74 (2015).
Bamshad, M. J., Nickerson, D. A. & Chong, J. X. Mendelian gene discovery: fast and furious with no end in sight. Am. J. Hum. Genet. 105, 448–455 (2019).
Manolio, T. A. et al. Finding the missing heritability of complex diseases. Nature 461, 747–753 (2009).
Vorstman, J. A. S. et al. Autism genetics: opportunities and challenges for clinical translation. Nat. Rev. Genet. 18, 362–376 (2017).
Ozonoff, S. et al. Recurrence risk for autism spectrum disorders: a Baby Siblings Research Consortium study. Pediatrics 128, e488–e495 (2011).
Risch, N. et al. Familial recurrence of autism spectrum disorder: evaluating genetic and environmental contributions. Am. J. Psychiatry 171, 1206–1213 (2014).
Fernandez, B. A. & Scherer, S. W. Syndromic autism spectrum disorders: moving from a clinically defined to a molecularly defined approach. Dialogues Clin. Neurosci. 19, 353–371 (2017).
De Rubeis, S. et al. Synaptic, transcriptional and chromatin genes disrupted in autism. Nature 515, 209–215 (2014).
Iossifov, I. et al. The contribution of de novo coding mutations to autism spectrum disorder. Nature 515, 216–221 (2014).
Sanders, S. J. et al. Insights into autism spectrum disorder genomic architecture and biology from 71 risk loci. Neuron 87, 1215–1233 (2015).
Yuen, R. K. C. et al. Genome-wide characteristics of de novo mutations in autism. NPJ Genom. Med. 1, 16027 (2016).
Marshall, C. R. et al. Structural variation of chromosomes in autism spectrum disorder. Am. J. Hum. Genet. 82, 477–488 (2008).
Brandler, W. M. et al. Paternally inherited cis-regulatory structural variants are associated with autism. Science 360, 327–331 (2018).
Bourgeron, T. From the genetic architecture to synaptic plasticity in autism spectrum disorder. Nat. Rev. Neurosci. 16, 551–563 (2015).
Tammimies, K. et al. Molecular diagnostic yield of chromosomal microarray analysis and whole-exome sequencing in children with autism spectrum disorder. J. Am. Med. Assoc. 314, 895–903 (2015).
An, J.-Y. et al. Genome-wide de novo risk score implicates promoter variation in autism spectrum disorder. Science 362, eaat6576 (2018).
Jiang, Y. H. et al. Detection of clinically relevant genetic variants in autism spectrum disorder by whole-genome sequencing. Am. J. Hum. Genet. 93, 249–263 (2013).
Werling, D. M. et al. An analytical framework for whole-genome sequence association studies and its implications for autism spectrum disorder. Nat. Genet. 50, 727–736 (2018).
Gaugler, T. et al. Most genetic risk for autism resides with common variation. Nat. Genet. 46, 881–885 (2014).
Grove, J. et al. Identification of common genetic risk variants for autism spectrum disorder. Nat. Genet. 51, 431–444 (2019).
Hannan, A. J. Tandem repeat polymorphisms: modulators of disease susceptibility and candidates for ‘missing heritability’. Trends Genet. 26, 59–65 (2010).
Bahlo, M. et al. Recent advances in the detection of repeat expansions with short-read next-generation sequencing. F1000Res. 7, 736 (2018).
Cortese, A. et al. Biallelic expansion of an intronic repeat in RFC1 is a common cause of late-onset ataxia. Nat. Genet. 51, 649–658 (2019).
Sato, N. et al. Spinocerebellar ataxia type 31 is associated with “inserted” penta-nucleotide repeats containing (TGGAA)n. Am. J. Hum. Genet. 85, 544–557 (2009).
Rafehi, H. et al. Bioinformatics-based identification of expanded repeats: a non-reference intronic pentamer expansion in RFC1 causes CANVAS. Am. J. Hum. Genet. 105, 151–165 (2019).
Hagerman, R. J. et al. Fragile X-associated neuropsychiatric disorders (FXAND). Front. Psychiatry 9, 564 (2018).
Dolzhenko, E. et al. ExpansionHunter Denovo: a computational method for locating known and novel repeat expansions in short-read sequencing data. Genome Biol. 21, 102 (2020).
Levy, S. et al. The diploid genome sequence of an individual human. PLoS Biol. 5, e254 (2007).
GTEx Consortium. Genetic effects on gene expression across human tissues. Nature 550, 204–213 (2017).
Olson, J. E. et al. Characteristics and utilisation of the Mayo Clinic Biobank, a clinic-based prospective collection in the USA: cohort profile. BMJ Open 9, e032707 (2019).
Subramanian, S., Mishra, R. K. & Singh, L. Genome-wide analysis of microsatellite repeats in humans: their abundance and density in specific genomic regions. Genome Biol. 4, R13 (2003).
Benson, G. Tandem repeats finder: a program to analyze DNA sequences. Nucleic Acids Res. 27, 573–580 (1999).
Willems, T., Gymrek, M., Highnam, G., Mittelman, D. & Erlich, Y. The landscape of human STR variation. Genome Res. 24, 1894–1904 (2014).
Bignell, G. R. et al. Signatures of mutation and selection in the cancer genome. Nature 463, 893–898 (2010).
Hannan, A. J. Tandem repeats mediating genetic plasticity in health and disease. Nat. Rev. Genet. 19, 286–298 (2018).
Lek, M. et al. Analysis of protein-coding genetic variation in 60,706 humans. Nature 536, 285–291 (2016).
Yuen, R. K. C. et al. Whole-genome sequencing of quartet families with autism spectrum disorder. Nat. Med. 21, 185–191 (2015).
Banerjee-Basu, S. & Packer, A. SFARI Gene: an evolving database for the autism research community. Dis. Model. Mech. 3, 133–135 (2010).
Trost, B. et al. A comprehensive workflow for read depth-based identification of copy-number variation from whole-genome sequence data. Am. J. Hum. Genet. 102, 142–155 (2018).
Takiyama, Y. et al. Single sperm analysis of the CAG repeats in the gene for Machado–Joseph disease (MJD1): evidence for non-Mendelian transmission of the MJD1 gene and for the effect of the intragenic CGG/GGG polymorphism on the intergenerational instability. Hum. Mol. Genet. 6, 1063–1068 (1997).
Dean, N. L. et al. Transmission ratio distortion in the myotonic dystrophy locus in human preimplantation embryos. Eur. J. Hum. Genet. 14, 299–306 (2006).
Shoubridge, C. et al. Is there a Mendelian transmission ratio distortion of the c.429_452dup(24bp) polyalanine tract ARX mutation? Eur. J. Hum. Genet. 20, 1311–1314 (2012).
Ekström, A.-B., Hakenäs-Plate, L., Samuelsson, L., Tulinius, M. & Wentz, E. Autism spectrum conditions in myotonic dystrophy type 1: a study on 57 individuals with congenital and childhood forms. Am. J. Med. Genet. B. Neuropsychiatr. Genet. 147B, 918–926 (2008).
Lagrue, E. et al. A large multicenter study of pediatric myotonic dystrophy type 1 for evidence-based management. Neurology 92, e852–e865 (2019).
Dolzhenko, E. et al. Detection of long repeat expansions from PCR-free whole-genome sequence data. Genome Res. 27, 1895–1903 (2017).
Dolzhenko, E. et al. ExpansionHunter: a sequence-graph-based tool to analyze variation in short tandem repeat regions. Bioinformatics 35, 4754–4756 (2019).
Tsai, L. Y. & Beisler, J. M. The development of sex differences in infantile autism. Br. J. Psychiatry 142, 373–378 (1983).
Satterstrom, F. K. et al. Large-scale exome sequencing study implicates both developmental and functional changes in the neurobiology of autism. Cell 180, 568–584 (2020).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows–Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
Purcell, S. et al. PLINK: a tool set for whole-genome association and population-based linkage analyses. Am. J. Hum. Genet. 81, 559–575 (2007).
Alexander, D. H., Novembre, J. & Lange, K. Fast model-based estimation of ancestry in unrelated individuals. Genome Res. 19, 1655–1664 (2009).
Koren, S. et al. Canu: scalable and accurate long-read assembly via adaptive k-mer weighting and repeat separation. Genome Res. 27, 722–736 (2017).
Ester, M., Kriegel, H., Sander, J. & Xu, X. A density-based algorithm for discovering clusters in large spatial databases with noise. In Proc. 2nd International Conference on Knowledge Discovery and Data Mining (AAAI, 1996).
Wang, K., Li, M. & Hakonarson, H. ANNOVAR: functional annotation of genetic variants from high-throughput sequencing data. Nucleic Acids Res. 38, e164 (2010).
de Leeuw, C. A., Mooij, J. M., Heskes, T. & Posthuma, D. MAGMA: generalized gene-set analysis of GWAS data. PLOS Comput. Biol. 11, e1004219 (2015).
Schizophrenia Working Group of the Psychiatric Genomics Consortium. Biological insights from 108 schizophrenia-associated genetic loci. Nature 511, 421–427 (2014).
Demontis, D. et al. Discovery of the first genome-wide significant risk loci for attention deficit/hyperactivity disorder. Nat. Genet. 51, 63–75 (2019).
Lee, J. J. et al. Gene discovery and polygenic prediction from a genome-wide association study of educational attainment in 1.1 million individuals. Nat. Genet. 50, 1112–1121 (2018).
Wood, A. R. et al. Defining the role of common variation in the genomic and biological architecture of adult human height. Nat. Genet. 46, 1173–1186 (2014).
Zhu, M. et al. Using ERDS to infer copy-number variants in high-coverage genomes. Am. J. Hum. Genet. 91, 408–421 (2012).
Abyzov, A., Urban, A. E., Snyder, M. & Gerstein, M. CNVnator: an approach to discover, genotype, and characterize typical and atypical CNVs from family and population genome sequencing. Genome Res. 21, 974–984 (2011).
Acknowledgements
We thank The Centre for Applied Genomics, especially J. Buchanan, B. Kellam, S. Lamoureux, T. Nalpathamkalam, R. Patel, W. Sung and Z. Wang. We also thank G. K. W. Wong, S. Walker and A. Paterson; the participating families in MSSNG (www.mss.ng), and Autism Speaks staff, M. Quirbach and V. Seifer, who manage the MSSNG and AGRE programs; the families at the participating SSC sites, and the principal investigators (A. Beaudet, R. Bernier, J. Constantino, E. Cook, E. Fombonne, D. Geschwind, R. Goin-Kochel, E. Hanson, D. Grice, A. Klin, D. Ledbetter, C. Lord, C. Martin, D. Martin, R. Maxim, J. Miles, O. Ousley, K. Pelphrey, B. Peterson, J. Piggot, C. Saulnier, M. State, W. Stone, J. Sutcliffe, C. Walsh, Z. Warren and E. Wijsman). Project funding is from Autism Speaks, the Canadian Institute for Advanced Research (CIFAR), the KRG Children’s Charitable Foundation, The Petroff Family Fund, The Kazman Family Fund, The Marigold Foundation, Genome Canada, the Canada Foundation for Innovation, the Government of Ontario, Canadian Institutes for Health Research (CIHR), the Natural Sciences and Engineering Research Council (NSERC), Brain Canada, Kids Brain Health Network, Province of Ontario Neurodevelopmental Disorders (POND) Network and the Ontario Brain Institute (OBI). R.K.C.Y. is supported by The Hospital for Sick Children’s Research Institute, SickKids Catalyst Scholar in Genetics, NARSAD Young Investigator award, Dataset Analysis Grant from Autism Speaks, the University of Toronto McLaughlin Centre and the Nancy E.T. Fahrner Award. B. Trost was funded by the Canadian Institutes for Health Research Banting Postdoctoral Fellowship and the Brain Canada Canadian Open Neuroscience Platform Research Scholar Award. C.M.N. and B.A.M. were supported by the Restracomp Award from The Hospital for Sick Children, and M.M. by the Ontario Graduate Scholarship. M.E.S.L. is funded by an Investigator Grant Award Program (IGAP) salary award from BC Children’s Hospital Research Institute and a Genome British Columbia Strategic Initiative Grant, D.G.A. by a National Institute of Mental Health Grant (1R01MH103371), E.A. by the Dr. Stuart D. Sims Chair in Autism, L.Z. by the Stollery Children’s Hospital Foundation Chair in Autism Research, P.S. by the Patsy and Jamie Anderson Chair in Child and Youth Mental Health, C.E.P. by the Canada Research Chair in Disease-Associated Genome Instability and S.W.S. holds the GlaxoSmithKline Chair in Genome Sciences at the University of Toronto and The Hospital for Sick Children.
Author information
Authors and Affiliations
Contributions
R.K.C.Y. conceived and designed the study. B. Trost and C.M.N. developed the tandem repeat detection pipeline, with additional contributions from E.D., M.A.E. and R.K.C.Y. W.E. performed statistical analyses, with additional contributions from B. Trost, B. Thiruvahindrapuram, I.C., G.P. and R.K.C.Y. I.B., M.M., T.P., N. Shum and N. Sato performed laboratory experiments for validation. B. Thiruvahindrapuram, B.A.M., Y.Y., A.D. and G.P. performed miscellaneous data analysis. O.H. and J.W. helped to process cloud-based data and provided general technical assistance. J.L.H. managed the collection of samples and phenotypes. E.W.K., S.B. and A.K.S. assisted with population control datasets. D.G.A., E.A., M.E., B.A.F., N.H., M.E.S.L., X.L., C.S., I.M.S., P.S. and L.Z. chose the phenotypic assessment tools and recruited, diagnosed and examined the participants. D.G., D.H. and S.W.S. supervised, managed and coordinated genomic data from MSSNG. R.K.C.Y. wrote the manuscript, with additional input from B. Trost, W.E., B. Thiruvahindrapuram, N. Sato, C.E.P. and S.W.S. R.K.C.Y. supervised the study, with additional input from C.E.P. and S.W.S. All authors read, reviewed and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
E.D. and M.A.E. are, or were, employees of Illumina, a public company that develops and markets systems for genetic analysis. D.G.A. is on the Scientific Advisory Boards of Stemina Biomarkers Discovery and Axial Therapeutics. E.A. has served as a consultant to Roche and Quadrant, has received grant funding from Roche, has received royalties from APPI and Springer, has received in-kind support from AMO Pharmaceuticals and has received editorial honoraria from Wiley. D.G. is employed by Verily. A.K.S. has a consulting role for Amgen, Bristol-Myers Squibb, Celgene, Ionis, Janssen, Oncopeptides, Ono, Roche, Seattle Genetics and Takeda, and has received research funding from Amgen, Celgene and Janssen. S.W.S. serves on the Scientific Advisory Committees of Population Bio and Deep Genomics; intellectual property originating from his research and held at The Hospital for Sick Children is licensed to Lineagen and separately to Athena Diagnostics. The strategies for genome-wide analysis and interpretation of tandem DNA repeats from genome sequence have been filed under reference H8313086USP (US provisional application number 62/951671) with the US Patent and Trademark Office.
Additional information
Peer review information Nature thanks Thomas Bourgeron, Anders Børglum and Anthony Hannan for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
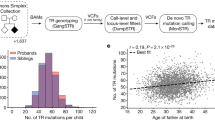
Extended Data Fig. 1 Study design.
a, Schematic workflow of the tandem repeat detection and analyses. 1Tandem repeats here are defined as those with 2–20-bp repeat motifs that span at least 150 bp. 2Rare expansions here are defined as tandem repeat expansions that are outliers according to size and occur in <0.1% of population controls from the 1000G. Note that EHdn only approximates the size and location of a given tandem repeat; thus, we use the term ‘region’ to refer to a genomic segment detected in this way, and reserve ‘location’ or ‘locus’ for sites that have been more precisely mapped. b, Genome-sequencing cohorts used for each analysis performed in this study. Numbers above each cohort represent the number of samples that remained after curation (Supplementary Notes).
Extended Data Fig. 2 Distribution of the number of tandem repeats detected by EHdn.
The number of tandem repeats detected by EHdn in a given sample is stratified by cohort, sequencing platform and DNA library preparation method (a; n = 2,504, 594, 1,220, 6,634 and 9,096 for 1000G/Illumina NovaSeq/PCR-free, MSSNG/Illumina HiSeq 2000 or 2500/PCR-based, MSSNG/Illumina HiSeq X/PCR-based, MSSNG/Illumina HiSeq X/PCR-free and SSC/Illumina HiSeq X/PCR-free, respectively) and predicted ancestry for samples in the ‘MSSNG/Illumina HiSeq X/PCR-free’ category (b; n = 157, 301, 247, 287, 4,841, 687 and 114 for admixed, AFR, AMR, EAS, EUR, OTH and SAS, respectively). Ancestry designations were derived from the 1000G ‘super populations’ (https://www.internationalgenome.org/category/population): AFR, African; AMR, admixed American; EAS, East Asian; EUR, European; OTH, other; SAS, South Asian. The centre of each box plot indicates the median, the lower and upper hinges correspond to the first and third quartiles, and the minima and maxima are 1.5× the interquartile range below or above the median, respectively.
Extended Data Fig. 3 Quality control for the detection of tandem repeats.
a–c, Histogram (left) and normal QQ plots (right) of the number of tandem repeats detected by EHdn for all samples (a), samples for which the number of tandem repeats was within mean ± 2 s.d. (b) and samples for which the number of tandem repeats was within mean ± 3 s.d. (c). Of the three distributions, data in c are the closest to a normal distribution.
Extended Data Fig. 4 The number of unique motifs in each repeat-containing region.
The number of unique motifs (y axis) in each repeat-containing region (x axis) is shown for all autosomal chromosomes and chromosomes X and Y.
Extended Data Fig. 5 Distributions of gnomAD gene constraints.
The distributions of gnomAD observed/expected (o/e) upper bounds are shown for genes with rare tandem repeat expansions near TSSs (n = 32 genes) and splice junctions (n = 80 genes), compared with other genes (n = 19,567 genes) (one-sided Wilcoxon rank-sum test). The minima and maxima indicate 3× the interquartile range-deviated o/e upper bounds from the median and the centre indicates the median of the o/e upper bounds.
Extended Data Fig. 6 Transmission tests.
a–c, Odds ratios calculated as ratios of the transmission events of genic large tandem repeats and those in intergenic regions. Only individuals with ASD of European ancestry in SSC (a; n = 1,808), MSSNG (b; n = 2,010) and both SSC and MSSNG (c; n = 3,818) were considered. d–f, Odds ratios calculated as ratios of the transmission events of large tandem repeats (99th percentile of length distribution) in a particular functional element to those in intergenic regions. Only individuals with ASD of European ancestry in SSC (d), MSSNG (e) and both SSC and MSSNG (f) were considered. Fisher’s exact tests were used to estimate the odds ratios and 95% confidence intervals are shown as error bars.
Extended Data Fig. 7 Transmission gene-set enrichment.
Odds ratios calculated as ratios of the transmission events of large tandem repeats (99th percentile of length distribution) in particular gene sets to those in intergenic regions. Only individuals with ASD of European ancestry in SSC (a; n = 1,808), MSSNG (b; n = 2,010) and both SSC and MSSNG (c; n = 3,818) were considered. Gene sets that were enriched in the burden analysis of rare tandem repeat expansions between children with ASD and their siblings without ASD in SSC are labelled. Red bars indicate significant enrichment in individuals with ASD (FWER < 25%). Fisher’s exact tests were used to estimate the odds ratios and 95% confidence intervals are shown as error bars.
Extended Data Fig. 8 Methods for sizing of the CTG repeat in DMPK.
a, Although short CTG repeats were correctly sized by ExpansionHunter (the results were perfectly matched with fragment analysis), slight discrepancies were observed in the estimates for premutation alleles between ExpansionHunter (EH) and PCR-based fragment analysis. Note that the length of the premutation CTG repeats (42 CTGs) was close to the read length of the HiSeq X platform (150 bp). N/A, not available. b, Predictions of the presence of longer CTG repeats were validated by repeat-primed PCR, although the estimated size by ExpansionHunter was shown to be an underestimate (the saw-tooth pattern of repeat-primed PCR extended longer than the predicted size). Repeat-primed PCR experiments were consistently reproduced at least three times for the large expansions. Repeat sizing experiments of PCR-amplifiable samples were consistently reproduced at least twice.
Extended Data Fig. 9 Validation of tandem repeats detected by EHdn.
a–d, Tandem repeats detected in CACNB1. e–h, Tandem repeats detected in FXN. a, e, Integrative Genomics Viewer read pile-up showing the reads aligning to the loci in CACNB1 and FXN in two families for which tandem repeat expansions were detected in the child (bottom). In both families, the expansion is transmitted from the mother to the child (samples highlighted in red in b and f). b, f, Image of the gel electrophoresis showing two bands that correspond to the expanded and unexpanded alleles in the mother and child. The father has only the unexpanded allele. Results from PCR and gel electrophoresis were consistently reproduced at least twice for CACNB1 and FXN loci (Supplementary Fig. 9). c, g, Chromatogram of the Sanger sequencing analysis of the expanded non-reference tandem repeat in the mother. d, h, Chromatogram of the Sanger sequencing analysis of the expanded non-reference tandem repeat in the child. Sanger sequencing was performed using the DNA of the expanded alleles, which was extracted from the gels.
Supplementary information
Supplementary Information
This file contains Supplementary Notes (additional results that could not be included in the manuscript due to space constraints) and Supplementary Figures 1-9.
Supplementary Tables
This file contains Supplementary Tables 1-15.
Supplementary Data
This file contains source data for Supplementary Figures 1, 3-5 and 7.
Rights and permissions
About this article
Cite this article
Trost, B., Engchuan, W., Nguyen, C.M. et al. Genome-wide detection of tandem DNA repeats that are expanded in autism. Nature 586, 80–86 (2020). https://doi.org/10.1038/s41586-020-2579-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-020-2579-z
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.