Abstract
Zika virus (ZIKV) is a flavivirus that is closely related to other human pathogens, such as dengue virus (DENV)1. Primary transmission usually involves Aedes aegypti, which has expanded its distribution range considerably2, although rarer infection routes, including mother-to-fetus transmission, sexual contact and blood transfusion, have also been observed3,4,5,6,7. Primary ZIKV infection is usually asymptomatic or mild in adults, with quickly resolved blood viraemia, but ZIKV might persist for months in saliva, urine, semen, breast milk and the central nervous system8,9,10,11,12. During a recent ZIKV outbreak in South America, substantial numbers of neurological complications, such as Guillain–Barré syndrome, were reported13,14 together with cases of microcephaly and associated developmental problems in infants born to women infected with ZIKV during pregnancy15,16,17,18,19,20, highlighting the clinical importance of this infection. Analyses of the human immune response to ZIKV are lacking21,22,23,24,25,26,27,28, but the recent outbreak has provided an opportunity to assess ZIKV immunity using current immunological methods. Here, we comprehensively assess the acute innate and adaptive immune response to ZIKV infection in ten women who were recruited during early infection and followed through reconvalescence. We define a cascade of events that lead to immunological control of ZIKV, with previous exposure to DENV impacting some, but not all, mediators of antiviral immunity.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
The data that support the findings of this study are available from the corresponding authors on reasonable request.
References
Ye, Q. et al. Genomic characterization and phylogenetic analysis of Zika virus circulating in the Americas. Infect. Genet. Evol. 43, 43–49 (2016).
Kraemer, M. U. G. et al. The global distribution of the arbovirus vectors Aedes aegypti and Ae. albopictus. eLife 4, e08347 (2015).
Simeone, R. M. et al. Possible Zika virus infection among pregnant women—United States and territories, May 2016. MMWR Morb. Mortal. Wkly. Rep. 65, 514–519 (2016).
Meaney-Delman, D. et al. Zika virus infection among U.S. pregnant travelers—August 2015-February 2016. MMWR Morb. Mortal. Wkly. Rep. 65, 211–214 (2016).
Foy, B. D. et al. Probable non-vector-borne transmission of Zika virus, Colorado, USA. Emerg. Infect. Dis. 17, 880–882 (2011).
Musso, D. et al. Potential sexual transmission of Zika virus. Emerg. Infect. Dis. 21, 359–361 (2015).
Motta, I. J. F. et al. Evidence for transmission of Zika virus by platelet transfusion. N. Engl. J. Med. 375, 1101–1103 (2016).
Petersen, L. R., Jamieson, D. J., Powers, A. M. & Honein, M. A. Zika Virus. N. Engl. J. Med. 374, 1552–1563 (2016).
Osuna, C. E. et al. Zika viral dynamics and shedding in rhesus and cynomolgus macaques. Nat. Med. 22, 1448–1455 (2016).
Bonaldo, M. C. et al. Isolation of infective Zika virus from urine and saliva of patients in Brazil. PLoS Negl. Trop. Dis. 10, e0004816 (2016).
Sotelo, J. R. et al. Persistence of Zika virus in breast milk after infection in late stage of pregnancy. Emerg. Infect. Dis. 23, 856–857 (2017).
Aid, M. et al. Zika virus persistence in the central nervous system and lymph nodes of rhesus monkeys. Cell 169, 610–620 (2017).
Brasil, P. et al. Guillain-Barré syndrome associated with Zika virus infection. Lancet 387, 1482 (2016).
Lannuzel, A. et al. Long-term outcome in neuroZika: when biological diagnosis matters. Neurology 92, e2406–e2420 (2019).
Brasil, P. et al. Zika virus infection in pregnant women in Rio de Janeiro. N. Engl. J. Med. 375, 2321–2334 (2016).
Rasmussen, S. A., Jamieson, D. J., Honein, M. A. & Petersen, L. R. Zika virus and birth defects—reviewing the evidence for causality. N. Engl. J. Med. 374, 1981–1987 (2016).
Costa, F. et al. Emergence of congenital Zika syndrome: viewpoint from the front lines. Ann. Intern. Med. 164, 689–691 (2016).
França, G. V. A. et al. Congenital Zika virus syndrome in Brazil: a case series of the first 1501 livebirths with complete investigation. Lancet 388, 891–897 (2016).
Hazin, A. N. et al. Computed tomographic findings in microcephaly associated with Zika virus. N. Engl. J. Med. 374, 2193–2195 (2016).
Honein, M. A. et al. Birth defects among fetuses and infants of US women with evidence of possible Zika virus infection during pregnancy. JAMA 317, 59–68 (2017).
Stettler, K. et al. Specificity, cross-reactivity, and function of antibodies elicited by Zika virus infection. Science 353, 823–826 (2016).
Wang, Q. et al. Molecular determinants of human neutralizing antibodies isolated from a patient infected with Zika virus. Sci. Transl. Med. 8, 369ra179 (2016).
Robbiani, D. F. et al. Recurrent potent human neutralizing antibodies to Zika virus in brazil and mexico. Cell 169, 597–609 (2017).
Grifoni, A. et al. Prior dengue virus exposure shapes T cell immunity to Zika virus in humans. J. Virol. 91, e01469-17 (2017).
Ricciardi, M. J. et al. Ontogeny of the B- and T-cell response in a primary Zika virus infection of a dengue-naïve individual during the 2016 outbreak in Miami, FL. PLoS Negl. Trop. Dis. 11, e0006000 (2017).
Lai, L. et al. Innate, T-, and B-cell responses in acute human Zika patients. Clin. Infect. Dis. 66, 1–10 (2018).
Delgado, F. G. et al. Improved immune responses against Zika virus after sequential dengue and Zika virus infection in humans. Viruses 10, 480 (2018).
Carlin, A. F. et al. A longitudinal systems immunologic investigation of acute Zika virus infection in an individual infected while traveling to Caracas, Venezuela. PLoS Negl. Trop. Dis. 12, e0007053 (2018).
Michlmayr, D., Andrade, P., Gonzalez, K., Balmaseda, A. & Harris, E. CD14+CD16+ monocytes are the main target of Zika virus infection in peripheral blood mononuclear cells in a paediatric study in Nicaragua. Nat. Microbiol. 2, 1462–1470 (2017).
Foo, S. -S. et al. Asian Zika virus strains target CD14+ blood monocytes and induce M2-skewed immunosuppression during pregnancy. Nat. Microbiol. 2, 1558–1570 (2017).
Sun, X. et al. Transcriptional changes during naturally acquired Zika virus infection render dendritic cells highly conducive to viral replication. Cell Rep. 21, 3471–3482 (2017).
Wang, C. et al. Myeloid-derived suppressor cells inhibit T follicular helper cell immune response in Japanese encephalitis virus infection. J. Immunol. 199, 3094–3105 (2017).
Everett, H. & McFadden, G. Apoptosis: an innate immune response to virus infection. Trends Microbiol. 7, 160–165 (1999).
Shi, C. & Pamer, E. G. Monocyte recruitment during infection and inflammation. Nat. Rev. Immunol. 11, 762–774 (2011).
Ishikawa, T., Yamanaka, A. & Konishi, E. A review of successful flavivirus vaccines and the problems with those flaviviruses for which vaccines are not yet available. Vaccine 32, 1326–1337 (2014).
Barouch, D. H., Thomas, S. J. & Michael, N. L. Prospects for a Zika virus vaccine. Immunity 46, 176–182 (2017).
Abbink, P. et al. Protective efficacy of multiple vaccine platforms against Zika virus challenge in rhesus monkeys. Science 353, 1129–1132 (2016).
Larocca, R. A. et al. Vaccine protection against Zika virus from Brazil. Nature 536, 474–478 (2016).
Rogers, T. F. et al. Zika virus activates de novo and cross-reactive memory B cell responses in dengue-experienced donors. Sci. Immunol. 2, eaan6809 (2017).
Dejnirattisai, W. et al. Dengue virus sero-cross-reactivity drives antibody-dependent enhancement of infection with Zika virus. Nat. Immunol. 17, 1102–1108 (2016).
Bardina, S. V. et al. Enhancement of Zika virus pathogenesis by preexisting antiflavivirus immunity. Science 356, 175–180 (2017).
Pantoja, P. et al. Zika virus pathogenesis in rhesus macaques is unaffected by pre-existing immunity to dengue virus. Nat. Commun. 8, 15674 (2017).
Terzian, A. C. B. et al. Viral load and cytokine response profile does not support antibody-dependent enhancement in dengue-primed Zika virus-infected patients. Clin. Infect. Dis. 65, 1260–1265 (2017).
George, J. et al. Prior exposure to Zika virus significantly enhances peak dengue-2 viremia in rhesus macaques. Sci. Rep. 7, 10498 (2017).
Wen, J. et al. Identification of Zika virus epitopes reveals immunodominant and protective roles for dengue virus cross-reactive CD8+ T cells. Nat. Microbiol. 2, 17036 (2017).
Elong Ngono, A. et al. Mapping and role of the CD8+ T cell response during primary Zika virus infection in mice. Cell Host Microbe 21, 35–46 (2017).
Duangchinda, T. et al. Immunodominant T-cell responses to dengue virus NS3 are associated with DHF. Proc. Natl Acad. Sci. USA 107, 16922–16927 (2010).
Tian, Y., Sette, A. & Weiskopf, D. Cytotoxic CD4 T Cells: differentiation, function, and application to dengue virus infection. Front. Immunol. 7, 531 (2016).
Dung, N. T. P. et al. Timing of CD8+ T cell responses in relation to commencement of capillary leakage in children with dengue. J. Immunol. 184, 7281–7287 (2010).
Crosby, E. J., Goldschmidt, M. H., Wherry, E. J. & Scott, P. Engagement of NKG2D on bystander memory CD8 T cells promotes increased immunopathology following Leishmania major infection. PLoS Pathog. 10, e1003970 (2014).
Kim, J. et al. Innate-like cytotoxic function of bystander-activated CD8+ T cells is associated with liver injury in acute hepatitis A. Immunity 48, 161–173 (2018).
Maini, M. K. et al. The role of virus-specific CD8+ cells in liver damage and viral control during persistent hepatitis B virus infection. J. Exp. Med. 191, 1269–1280 (2000).
Baylis, S. A. et al. Harmonization of nucleic acid testing for Zika virus: development of the 1st world health organization international standard. Transfusion 57, 748–761 (2017).
Trösemeier, J. -H. et al. Genome sequence of a candidate World Health Organization reference strain of Zika virus for nucleic acid testing. Genome. Announc. 4, e00917-16 (2016).
Acknowledgements
We thank the patients as well as the physicians and medical staff of the Viral Hepatitis Clinic in Brazil who provided care to the patients; the Flavivirus Laboratory staff at FIOCRUZ as well as J. Quick and N. Loman from the University of Birmingham for their assistance in analysing patient samples for ZIKV; B. da Silva Baptista as well as the Massachusetts General Hospital HSCI CRM flow cytometry core facility members for their technical assistance. This work was supported in part by National Institutes of Health grants U01 AI131314 (to G.M.L. and L.L.L.-X.), U19 AI066345 (to G.M.L. and L.L.L.-X.), HHSN27220140045C (to A.S.), 1PO1AI106695-01A1 (to A.S.), U19AI118626-01 (to A.S.), 75N9301900065 (to A.S.), the EU grant 734584 (to A.S.), Conselho Nacional de Desenvolvimento Tecnológico (CNPq 470092/2014-9; 200099/2016-7), Fundação de Amparo à Pesquisa do Estado do Rio de Janeiro (Faperj E-26/202.930/2016) and ‘Sicherheit von Blut(produkten) und Geweben hinsichtlich der Abwesenheit von Zikaviren’ from the German Ministry of Health.
Author information
Authors and Affiliations
Contributions
P.T., J.G.M. and A.T.-C. conceived and designed the study. P.T., J.G.M., A.T.-C. and R.C.H. performed immunophenotyping and intracellular cytokine stainings by flow cytometry. J.G.M., P.S.F.d.S., V.d.M.d.M. and L.L.L.-X. collected clinical data and biological samples. C.Y., A.M.B.d.F. and S.A.B. performed ZIKV-RNA detection assays. J.G.M., C.Y., M.A.P, J.M. and S.A.B. performed anti-ZIKV antibody ELISA assays. C.Y. and J.B. performed ZIKV neutralizing antibody assays. D.W., A.G., A.S., D.H.B., S.A.B. and L.L.L.-X. contributed to the study design and data interpretation. G.M.L. conceived and supervised the study, and provided funding. P.T. and G.M.L. analysed the data and wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
J.M. is the owner of Dr. Julio Moran Laboratories, a company owning and, for research purposes only, distributing the ELISA assays DIACHECK for the detection of anti-human ZIKV antibodies. The other authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
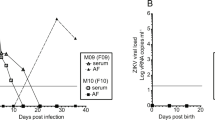
Extended Data Fig. 1 Patient’s symptoms and ZIKV-RNA detection data.
(a) Patient’s symptoms information and associated plasma ZIKV-RNA detection, over time. (b) Quantification of ZIKV RNA in the plasma. Each patient and previous exposure to DENV status, as defined by the positive detection of anti-DENV IgG at symptom onset, is displayed through unique symbols and connecting lines. X-axis, Time (days) represents the time from onset of symptoms. Gray dashed line notes the assay’s detection limit. Data are representative of n=2 independent experiments.
Extended Data Fig. 2 Gating strategies for blood immunophenotyping.
Flow cytometry gating strategies for the identification of the different cell subsets of monocytes (NC= non-classical; I= intermediate; C= classical), dendritic cells (pDCs= plasmacytoid dendritic cells; mDCs= myeloid dendritic cells) and MDSCs (myeloid-derived suppressor cells) (a), as well as plasmablasts and activated CD8+ T cells (b).
Extended Data Fig. 3 Changes in immune cell frequencies following acute ZIKV infection.
Representative flow-cytometry plots showing changes in the frequency of monocytes (a), dendritic cells (b), plasmablasts (c) and activated CD8+ T cells (d), at different time points following ZIKV infection. Numerical values of the measured frequencies in a total n=10 patients, at different time points (acute n=10; recovery n=5; follow-up n=4) are displayed in Fig. 1d. Times (in days) from onset of symptoms are indicated.
Extended Data Fig. 4 Humoral immune response to acute ZIKV infection.
Linear regression analysis to model the relationship between plasma anti-ZIKV IgM and IgA detection signals (n=41) using DIACHECK ELISA assays. (b) Plasma detection of anti-ZIKV IgM antibodies using EUROIMMUNE ELISA assay (left chart) and linear regression analysis to model the relationship between plasma anti-ZIKV IgM detection signals obtained with EUROIMMUNE and DIACHECK ELISA assays (n=41) (right chart). (c) Plasma detection of anti-ZIKV IgG antibodies using EUROIMMUNE ELISA assay (left chart) and linear regression analysis to model the relationship between plasma anti-ZIKV IgG detection signals obtained with EUROIMMUNE and DIACHECK ELISA assays (n=41) (right chart). (a-c) Each patient and previous exposure to DENV status, as defined by the positive detection of anti-DENV IgG at symptom onset, is displayed through unique symbols and connecting lines. Gray dashed lines note assay’s detection limits. X-axis, (Time (days)) represents the time from onset of symptoms. Data are expressed as mean values of OD/CO ratios from two independent experiments. Pearson correlation coefficient R and significance p (two-sided) values are reported from the linear regression analysis performed with GraphPad Prism v.6 software.
Extended Data Fig. 5 Titration of anti-ZIKV neutralizing antibodies.
Representative titration assays for the detection of anti-ZIKV neutralizing antibodies, overtime. Titers were measured by endpoint titration. A color mapping of the OD/CO ratio values for the detection anti-ZIKV IgM and IgG using EUROIMUNE ELISA assays are indicated. The data are representative of n=2 (patients CR8587, CR8592, CR8602, CR8663, CR4434, CR8597 and CR8622), n=3 (patients CR4965 and 8603) or n=4 (CR8623) independent experiments.
Extended Data Fig. 6 ZIKV-specific T cell memory differentiation following acute ZIKV infection.
T cell memory differentiation based on CCR7 and CD45RA co-expression (naïve: CCR7+CD45RA+; CM: CCR7+CD45RA-; EM: CCR7-CD45RA-; TEMRA: CCR7-CD45RA+). Frequencies of ZIKV-specific CD4+ (a) and CD8+ (b) T cells across the different memory subsets over time, from 5 different patients, are indicated. The analysis was performed on a total of n=6 patients, at different time points (Data from patient CR4965 are available in Fig. 4a, b).
Extended Data Fig. 7 ZIKV-specific T cell functional profiles overtime.
Detailed representation of the overlapping pie charts presented in Fig. 4d, e. The data represent the different sub-groups of cytokine secreting and cytotoxic CD154+CD4+ (a) and CD69+CD8+ (b) T cells after stimulation with 15-mer overlapping peptide pools covering all ZIKV-proteins, by ex vivo intracellular cytokine stainings (ICSs). Frequencies of IL-2, TNFa, IFNγ and CD107a co-expressing cells are indicated. Baseline signals of IL-2, TNFα, IFNγ and CD107a-expressing cells from unstimulated controls have been subtracted to the stimulated conditions to allow the visualization of ZIKV-specific CD4+ and CD8+ T cell signals. Only time points with detectable CD154+IFNγ+CD4+ (acute n=24, recovery n=26, follow-up n=29) and CD69+IFNγ+CD8+ T cells (acute n=17, recovery n=19, follow-up n=24) from patients CR4965, CR8623, CR8603 and CR8622, as depicted in Fig. 3a, b, have been used for this analysis. Black bars correspond to the median of expression in each condition.
Extended Data Fig. 8 ZIKV-specific T cell functional profiles across the different viral proteins targeted.
Overlapping pie charts describing the polyfunctionality of ZIKV-specific CD4+ (a) and CD8+ (b) T cells according to the ZIKV-overlapping peptide pools used for T cell stimulation and determined by ex vivo intracellular cytokine staining (ICS) as defined in Fig. 3. Baseline signals of TNFα, IL-2, CD107a and IFNγ-producing cells in unstimulated controls have been subtracted from ZIKV-stimulated assays to allow the visualization of ZIKV-specific CD4+ and CD8+ T cell signals. Only time points with detectable CD154+IFNγ+CD4+ (acute n=24, recovery n=26, follow-up n=29) and CD69+IFNγ+CD8+ T cells (acute n=17, recovery n=19, follow-up n=24) from patients CR4965, CR8623, CR8603 and CR8622, as depicted in Fig. 3a, b, were used for this analysis. Distribution of the numbers (n) of T cell responses across the different ZIKV-peptide pools are reported.
Extended Data Fig. 9
General overview of the dynamics of immune responses following acute ZIKV infection in human.
Extended Data Fig. 10
Patient’s demographics and HLA-types.
Supplementary information
Rights and permissions
About this article
Cite this article
Tonnerre, P., Melgaço, J.G., Torres-Cornejo, A. et al. Evolution of the innate and adaptive immune response in women with acute Zika virus infection. Nat Microbiol 5, 76–83 (2020). https://doi.org/10.1038/s41564-019-0618-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41564-019-0618-z
This article is cited by
-
Zika virus infection during pregnancy and vertical transmission: case reports and peptide-specific cell-mediated immune responses
Archives of Virology (2024)
-
Immune phenotypes that are associated with subsequent COVID-19 severity inferred from post-recovery samples
Nature Communications (2022)
-
Aberrant NAD+ metabolism underlies Zika virus–induced microcephaly
Nature Metabolism (2021)