Abstract
NARROW LEAF 1 (NAL1) is a breeding-valuable pleiotropic gene that affects multiple agronomic traits in rice, although the molecular mechanism is largely unclear. Here, we report that NAL1 is a serine protease and displays a novel hexameric structure consisting of two ATP-mediated doughnut-shaped trimeric complexes. Moreover, we identified TOPLESS-related corepressor OsTPR2 involved in multiple growth and development processes as the substrate of NAL1. We found that NAL1 degraded OsTPR2, thus modulating the expression of downstream genes related to hormone signalling pathways, eventually achieving its pleiotropic physiological function. An elite allele, NAL1A, which may have originated from wild rice, could increase grain yield. Furthermore, the NAL1 homologues in different crops have a similar pleiotropic function to NAL1. Our study uncovers a NAL1–OsTPR2 regulatory module and provides gene resources for the design of high-yield crops.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
ChIP-seq data generated in this study have been deposited in the GEO database under accession no. GSE207132. Coordinates and structure factors of NAL1 in this study have been deposited in the Protein Data Bank under accession code 7Y77. The structure data are obtained from public data in the Protein Data Bank under accession code 1P01, 1SGP, 2R3Y, 3K6Y and 5IL9. The orthologues of wheat, maize, soybean, oil crop were downloaded from Ensembl plants (https://plants.ensembl.org/Oryza_sativa/Gene/Compara_Ortholog?db=core;g=Os04g0615000;r=4:31205267-31214632;t=Os04t0615000-01). Materials used in this study are available upon request. Source data are provided with this paper.
References
Ray, D. K., Mueller, N. D., West, P. C. & Foley, J. A. Yield trends are insufficient to double global crop production by 2050. PLoS ONE 8, e66428 (2013).
Liu, M. et al. Inducible overexpression of Ideal Plant Architecture1 improves both yield and disease resistance in rice. Nat. Plants 5, 389–400 (2019).
Xue, W. et al. Natural variation in Ghd7 is an important regulator of heading date and yield potential in rice. Nat. Genet. 40, 761–767 (2008).
Weng, X. Y. et al. Grain number, plant height, and heading date7 is a central regulator of growth, development, and stress response. Plant Physiol. 164, 735–747 (2014).
Wang, Q. et al. The Ghd7 transcription factor represses ARE1 expression to enhance nitrogen utilization and grain yield in rice. Mol. Plant 14, 1012–1023 (2021).
Qian, Q., Guo, L. B., Smith, S. M. & Li, J. Y. Breeding high-yield superior quality hybrid super rice by rational design. Natl Sci. Rev. 3, 283–294 (2016).
Li, X. et al. Genetic control of the root system in rice under normal and drought stress conditions by genome-wide association study. PLoS Genet. 13, e1006889 (2017).
Taguchi-Shiobara, F. et al. Natural variation in the flag leaf morphology of rice due to a mutation of the NARROW LEAF 1 gene in Oryza sativa L. Genetics 201, 795–808 (2015).
Takai, T. et al. A natural variant of NAL1, selected in high-yield rice breeding programs, pleiotropically increases photosynthesis rate. Sci. Rep. 3, 2149 (2013).
Wang, Q. et al. Genetic architecture of natural variation in rice chlorophyll content revealed by a genome-wide association study. Mol. Plant 8, 946–957 (2015).
Yano, K. et al. Genome-wide association study using whole-genome sequencing rapidly identifies new genes influencing agronomic traits in rice. Nat. Genet. 48, 927–934 (2016).
Zhang, G. H. et al. LSCHL4 from japonica cultivar, which is allelic to NAL1, increases yield of indica super rice 93-11. Mol. Plant 7, 1350–1364 (2014).
Xu, J. L. et al. SS1 (NAL1)- and SS2-mediated genetic networks underlying source-sink and yield traits in rice (Oryza sativa L.). PLoS ONE 10, e0132060 (2015).
Singh, V. K. et al. Effect of qGN4.1 QTL for grain number per panicle in genetic backgrounds of twelve different mega varieties of rice. Rice (N Y) 11, 8 (2018).
Fujita, D. et al. NAL1 allele from a rice landrace greatly increases yield in modern indica cultivars. Proc. Natl Acad. Sci. USA 110, 20431–20436 (2013).
Chen, M. et al. Fine mapping of a major QTL for flag leaf width in rice, qFLW4, which might be caused by alternative splicing of NAL1. Plant Cell Rep. 31, 863–872 (2012).
Ding, X., Li, X. & Xiong, L. Evaluation of near-isogenic lines for drought resistance QTL and fine mapping of a locus affecting flag leaf width, spikelet number, and root volume in rice. Theor. Appl. Genet. 123, 815–826 (2011).
Wang, Y. et al. Natural sequence variations and combinations of GNP1 and NAL1 determine the grain number per panicle in rice. Rice (N Y) 13, 14 (2020).
Qi, J. et al. Mutation of the rice Narrow leaf1 gene, which encodes a novel protein, affects vein patterning and polar auxin transport. Plant Physiol. 147, 1947–1959 (2008).
Lin, L., Zhao, Y., Liu, F., Chen, Q. & Qi, J. Narrow leaf 1 (NAL1) regulates leaf shape by affecting cell expansion in rice (Oryza sativa L.). Biochem. Biophys. Res. Commun. 516, 957–962 (2019).
Cho, S.-H., Yoo, S.-C., Zhang, H., Lim, J.-H. & Paek, N.-C. Rice NARROW LEAF1 regulates leaf and adventitious root development. Plant Mol. Biol. Rep. 32, 270–281 (2013).
Jiang, D. et al. Characterization of a null allelic mutant of the rice NAL1 gene reveals its role in regulating cell division. PLoS ONE 10, e0118169 (2015).
Huang, Y. et al. Variation in the regulatory region of FZP causes increases in secondary inflorescence branching and grain yield in rice domestication. Plant J. 96, 716–733 (2018).
Ye, L. et al. A trypsin family protein gene controls tillering and leaf shape in barley. Plant Physiol. 181, 701–713 (2019).
Plant, A. R., Larrieu, A. & Causier, B. Repressor for hire! The vital roles of TOPLESS‐mediated transcriptional repression in plants. New Phytol. 231, 963–973 (2021).
Ryu, H., Cho, H., Bae, W. & Hwang, I. Control of early seedling development by BES1/TPL/HDA19-mediated epigenetic regulation of ABI3. Nat. Commun. 5, 4138 (2014).
Wang, L., Kim, J. & Somers, D. E. Transcriptional corepressor TOPLESS complexes with pseudoresponse regulator proteins and histone deacetylases to regulate circadian transcription. Proc. Natl Acad. Sci. USA 110, 761–766 (2013).
Long, J. A., Ohno, C., Smith, Z. R. & Meyerowitz, E. M. TOPLESS regulates apical embryonic fate in Arabidopsis. Science 312, 1520–1523 (2006).
Tang, N. et al. MODD mediates deactivation and degradation of OsbZIP46 to negatively regulate ABA signaling and drought resistance in rice. Plant Cell 28, 2161–2177 (2016).
Deribe, Y. L., Pawson, T. & Dikic, I. Post-translational modifications in signal integration. Nat. Struct. Mol. Biol. 17, 666–672 (2010).
Niu, D. et al. SIZ1-mediated SUMOylation of TPR1 suppresses plant immunity in Arabidopsis. Mol. Plant 12, 215–228 (2019).
Causier, B., Ashworth, M., Guo, W. & Davies, B. The TOPLESS interactome: a framework for gene repression in Arabidopsis. Plant Physiol. 158, 423–438 (2012).
Gao, X. et al. OsLIS-L1 encoding a lissencephaly type-1-like protein with WD40 repeats is required for plant height and male gametophyte formation in rice. Planta 235, 713–727 (2012).
Jiang, L. et al. DWARF 53 acts as a repressor of strigolactone signalling in rice. Nature 504, 401–405 (2013).
Fang, J. et al. The URL1-ROC5-TPL2 transcriptional repressor complex represses the ACL1 gene to modulate leaf rolling in rice. Plant Physiol. 185, 1722–1744 (2021).
Zhuang, H. et al. NONSTOP GLUMES1 encodes a C2H2 zinc finger protein that regulates spikelet development in rice. Plant Cell 32, 392–413 (2020).
Hao, Y. et al. Genome-wide identification, phylogenetic analysis, expression profiling, and protein–protein interaction properties of TOPLESS gene family members in tomato. J. Exp. Bot. 65, 1013–1023 (2014).
Liu, X., Galli, M., Camehl, I. & Gallavotti, A. RAMOSA1 ENHANCER LOCUS2-mediated transcriptional repression regulates vegetative and reproductive architecture. Plant Physiol. 179, 348–363 (2019).
Szemenyei, H., Hannon, M. & Long, J. A. TOPLESS mediates auxin-dependent transcriptional repression during Arabidopsis embryogenesis. Science 319, 1384–1386 (2008).
Martin-Arevalillo, R. et al. Structure of the Arabidopsis TOPLESS corepressor provides insight into the evolution of transcriptional repression. Proc. Natl Acad. Sci. USA 114, 8107–8112 (2017).
Ke, J. et al. Structural basis for recognition of diverse transcriptional repressors by the TOPLESS family of corepressors. Sci. Adv. 1, e1500107 (2015).
Zhang, Z. et al. Gnp4/LAX2, a RAWUL protein, interferes with the OsIAA3–OsARF25 interaction to regulate grain length via the auxin signaling pathway in rice. J. Exp. Bot. 69, 4723–4737 (2018).
Gao, S. et al. CYTOKININ OXIDASE/DEHYDROGENASE4 integrates cytokinin and auxin signaling to control rice crown root formation. Plant Physiol. 165, 1035–1046 (2014).
Song, X. et al. IPA1 functions as a downstream transcription factor repressed by D53 in strigolactone signaling in rice. Cell Res. 27, 1128–1141 (2017).
Huang, X. et al. A map of rice genome variation reveals the origin of cultivated rice. Nature 490, 497–501 (2012).
Wang, W. et al. Genomic variation in 3,010 diverse accessions of Asian cultivated rice. Nature 557, 43–49 (2018).
Ouyang, M. et al. The crystal structure of Deg9 reveals a novel octameric-type HtrA protease. Nat. Plants 3, 973–982 (2017).
Tsiatsiani, L., Gevaert, K. & Van Breusegem, F. Natural substrates of plant proteases: how can protease degradomics extend our knowledge? Physiol. Plant. 145, 28–40 (2012).
Ma, H. et al. A D53 repression motif induces oligomerization of TOPLESS corepressors and promotes assembly of a corepressor–nucleosome complex. Sci. Adv. 3, e1601217 (2017).
Santner, A. & Estelle, M. Recent advances and emerging trends in plant hormone signalling. Nature 459, 1071–1078 (2009).
Pauwels, L. et al. NINJA connects the co-repressor TOPLESS to jasmonate signalling. Nature 464, 788–791 (2010).
Li, C. et al. Arabidopsis ECAP is a new adaptor protein that connects JAZ repressors with the TPR2 co-repressor to suppress jasmonate-responsive anthocyanin accumulation. Mol. Plant 13, 246–265 (2020).
Wang, L. et al. Transcriptional regulation of strigolactone signalling in Arabidopsis. Nature 583, 277–281 (2020).
Xu, M., Zhu, L., Shou, H. & Wu, P. A PIN1 family gene, OsPIN1, involved in auxin-dependent adventitious root emergence and tillering in rice. Plant Cell Physiol. 46, 1674–1681 (2005).
Duan, J. et al. Strigolactone promotes cytokinin degradation through transcriptional activation of CYTOKININ OXIDASE/DEHYDROGENASE 9 in rice. Proc. Natl Acad. Sci. USA 116, 14319–14324 (2019).
Laplaze, L. et al. Cytokinins act directly on lateral root founder cells to inhibit root initiation. Plant Cell 19, 3889–3900 (2007).
Zhao, Y. et al. The interaction between rice ERF3 and WOX11 promotes crown root development by regulating gene expression involved in cytokinin signaling. Plant Cell 27, 2469–2483 (2015).
Xu, X. et al. Pyramiding of the dep1-1 and NAL1NJ6 alleles achieves sustainable improvements in nitrogen-use efficiency and grain yield in japonica rice breeding. J. Genet. Genomics 46, 325–328 (2019).
Ouyang, X. et al. Partially functional NARROW LEAF1 balances leaf photosynthesis and plant architecture for greater rice yield. Plant Physiol. 189, 772–789 (2022).
Zhang, Q. et al. Genomic insights into the recent chromosome reduction of autopolyploid sugarcane Saccharum spontaneum. Nat. Genet. 54, 885–896 (2022).
He, Y. et al. Programmed self-elimination of the CRISPR/Cas9 construct greatly accelerates the isolation of edited and transgene-free rice plants. Mol. Plant 11, 1210–1213 (2018).
Otwinowski, Z. & Minor, W. Processing of X-ray diffraction data collected in oscillation mode. Methods Enzymol. 276, 307–326 (1997).
Winn, M. D. et al. Overview of the CCP4 suite and current developments. Acta Crystallogr. D 67, 235–242 (2011).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D 60, 2126–2132 (2004).
Adams, P. D. et al. PHENIX: building new software for automated crystallographic structure determination. Acta Crystallogr. D 58, 1948–1954 (2002).
Chen, S., Zhou, Y., Chen, Y. & Gu, J. fastp: an ultra-fast all-in-one FASTQ preprocessor. Bioinformatics 34, i884–i890 (2018).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Zhang, Y. et al. Model-based analysis of ChIP-Seq (MACS). Genome Biol. 9, R137 (2008).
Ross-Innes, C. S. et al. Differential oestrogen receptor binding is associated with clinical outcome in breast cancer. Nature 481, 389–393 (2012).
Yu, G., Wang, L. G. & He, Q. Y. ChIPseeker: an R/Bioconductor package for ChIP peak annotation, comparison and visualization. Bioinformatics 31, 2382–2383 (2015).
Ramirez, F., Dundar, F., Diehl, S., Gruning, B. A. & Manke, T. deepTools: a flexible platform for exploring deep-sequencing data. Nucleic Acids Res. 42, W187–W191 (2014).
Robinson, J. T. et al. Integrative Genomics Viewer. Nat. Biotechnol. 29, 24–26 (2011).
Danecek, P. et al. The variant call format and VCFtools. Bioinformatics 27, 2156–2158 (2011).
Acknowledgements
We thank the staff of the BL17U1/BL19U1 beamline of the National Center for Protein Sciences Shanghai (NCPSS) at the Shanghai Synchrotron Radiation Facility for assistance during data collection, and research associate D. Zhang at the Center for Protein Research, Huazhong Agricultural University, for technical support. This work was supported by the National Natural Science Foundation of China (grants 31930080 to L.X., 31821005 to L.X., 32270255 to J.Y.), the Foundation of Hubei Hongshan Laboratory (grants 2021hszd011, 2021hskf003 to J.Y.), and the China Postdoctoral Science Foundation (grant 2019M652669 to W.L.). We thank the BaiChuan fellowship of College of Life Science and Technology, Huazhong Agricultural University, for funding support. The computations in this paper were run on the bioinformatics computing platform of the National Key Laboratory of Crop Genetic Improvement, Huazhong Agricultural University.
Author information
Authors and Affiliations
Contributions
W.L., J.Y., P.Y. and L.X. designed the study. W.L., J.Y., Y.Z., F.Z., Y.Y., X. Li and H.W. performed all experiments. W.L., J.Y., Z.G., Y.C. and H.T. analysed the data. W.L., J.Y., H.X., X. Lai and L.X. wrote and revised the article.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Plants thanks Makoto Matsuoka and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 NAL1 forms a hexamer.
a, The domain organization of NAL1. b, NAL1 trimer in asymmetric unit. c, Analytical ultracentrifugation characterization of NAL1 oligomerization. d, Structural alignment of determined NAL159–463 protomers with the AlphaFold2 predicted NAL11–582. e, Structure superposition of NAL1 with other serine proteases. f, Secondary structural elements of NAL1 protomer. g, Density map of ATP molecules. h, Size-exclusion chromatography analysis of NAL1 mutants. Fractions with the same elution volume from each injection were subjected to SDS-PAGE. The experiment in h was repeated independently three times with similar results. i, Structural alignment of catalytic triad in each protomer. The arrow indicates the orientations of the imidazole ring of H233 are relatively rotated. j, Oligomeric architecture comparison with other Deg proteases.
Extended Data Fig. 2 Plant architecture and yield-related traits of wild type (ZH11), complementary lines (NAL1A-COM and NAL1G-COM), and NAL1-knockout line (nal1-cri).
a, Flag leaf morphology, bar = 5 cm. b, Root system, bar = 5 cm. c, Panicle structure, bar = 5 cm.
Extended Data Fig. 3 C-terminal EAR-1 motif of NAL1 mediates the interaction with OsTPR2.
a, NAL1 interacts with OsTPR family proteins. NAL1 interacts with OsTPR family proteins (OsTPR1, OsTPR2 and OsTPR3) in LCI assays in tobacco leaves. The 10-AA deletion mutant nal1-1 was used as a negative control. b, A schematic domain organization of NAL1 and TPD. c, Size-exclusion chromatography analysis of the interactions between TPD and NAL1 truncations and mutants. Deletions of the C-terminal EAR motifs abolished their interactions. The last EAR motif had no effect on their interaction. The experiment in c was repeated independently three times with similar results. d, Structural model of OsTPD and NAL1EAR-1 complex. The model was obtained based on the structure of OsTPD and NINJA (PDB ID: 6C6V). e, Interaction details between OsTPD and NAL1EAR-1. Key residues were labeled.
Extended Data Fig. 4 OsTPR2 was degraded by NAL1 and partially rescue nal1-cri’s phenotype.
a, Degradation of OsTPR2 in ZF802 and nal1-1 protoplasts after CHX treatment. Transfected rice protoplasts were incubated for 16 h, and then treated with (50 g ml−1) CHX to block protein synthesis. Equal volume protoplasts were collected at different time points for immunoblotting detection of OsTPR2. b, Degradation of OsTPR2 in total proteins extracted from ZF802 and nal1-1 seedlings using cell-free protein degradation assays. His-OsTPR2 was respectively added to extracts from ZH11 and nal1-cri plants. Equal volume protein mix was collected at different time points for immuno-blotting detection of OsTPR2. c, Degradation of OsTPR2 in NIL-IR (NAL1A) and NIL-ZS (NAL1G) protoplasts after CHX treatment. Transfected rice protoplasts were incubated for 16 h, and then treated with (50 g ml-1) CHX to block protein synthesis. Equal volume of protoplasts were collected at different time points for immunoblotting detection of OsTPR2. d, Degradation of OsTPR2 in total protein extract from NIL-IR and NIL-ZS seedlings by cell-free protein degradation assays. His-OsTPR2 was respectively added to total protein extracts from NIL-IR and NIL-ZS plants. Equal volume of protein mix was collected at different time points for immunoblotting detection of OsTPR2. e, Degradation of OsTPR2 in vitro. NAL1 (encoded by NAL1A) or the NAL1H233R (encoded by NAL1G) were separately incubated with OsTPR2 for 0, 30, 60 or 120 min at 37 °C. After terminating the reaction, the reaction mixtures were subjected to SDS/PAGE. The location of OsTPR2 was indicated in the gel. f, OsTPR2 protein levels in ZH11 and nal1-cri in vivo. Anti-OsTPR2 antibody was used to detect OsTPR2 protein level in equal total protein extracts from ZH11 and nal1-cri seedlings. Actin was used as a control. g, OsTPR2 expression levels in ZH11, nal1-cri and the OsTPR2-knockdown lines in the nal1-cri background (named TN lines) seedlings (n = 4). RT-qPCR was repeated at least three times. Data are means ± SD. Different lowercase letters above bars indicate significant difference at P < 0.05 level by one-way ANOVA. The exact P values are listed in Supplementary Table 6. h-j, Plant architecture and yield-related traits of wild type (ZH11), nal1-cri, and OsTPR2-knockdown lines in the nal1-cri background (named TN lines). (h) Flag leaf morphology, bar = 5 cm. (i) Root system, bar = 5 cm. (j) Panicle structure, bar = 5 cm. k, The oligomerization of NAL1 decreased its proteolytic activity. Wild type or the mutated NAL1 (NAL1R120A, NAL1K139A and NAL1W144A) proteins were separately incubated with OsTPR2 for 0, 30, 60 or 120 min at 37 °C. After terminating the reaction, the reaction mixtures were subjected to SDS/PAGE. The location of OsTPR2 was indicated in the gel. The experiments in a-f, k were repeated independently three times with similar results.
Extended Data Fig. 5 NAL1 affects H3K18Ace levels of auxin and SL signaling pathway genes.
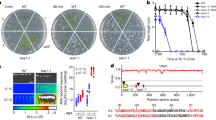
a, Relative histone acetylation and histone levels of nal1-cri mutant in Fig. 4d were normalized based on that of ZH11. The Integrated density of each interested protein band was carried out with FIJI/ImageJ. Data are means ± SD of three biological replicates. * and ** indicate significant differences between ZH11 and nal1-cri at P < 0.05 and 0.01 levels, and ns indicates no significant difference by two-sides Student’s t-test. b, Venn diagram of genes with significantly decreased expression and H3K18Ace levels. RNA-seq and ChIP-seq were used to analyse the expression and H3K18Ace levels, respectively, in ZH11 and nal1-cri seedlings. c, Genome browser traces of H3K18Ace ChIP-seq data in ZH11 and nal1-cri from representative genes. The fragments examined by ChIP-qPCR were indicated. d, H3K18Ace levels of Actin in ZH11 and nal1-cri seedlings. Data are fold-changes relative to levels in ZH11 seedlings. ChIP-qPCR was repeated at least three times. Data are means ± SD of three biological replicates. ns indicates no difference by two-sides Student’s t-test. e, Relative expression levels of auxin and SL signaling pathway genes in the seedlings of nal1-cri and TN lines. RT-qPCR was repeated for four times (n = 4). * and ** indicate significant differences between each complementary line and nal1-cri at P < 0.05 and 0.01 levels, and ns indicates no significant difference by two-sides Student’s t-test. The exact P values are listed in Supplementary Table 6.
Extended Data Fig. 6 Genetic analysis of NAL1 in the indica, japonica and O. rufipogon populations.
a, Representative genotypes of NAL1. b, Nucleotide diversity of NAL1. c, Single-nucleotide diversity.
Extended Data Fig. 7 Conservativeness of NAL1 homolog functions in different crops.
a, Phylogenic tree of NAL1 homologues from different crops. A neighbor-joining tree was built by MEGA-X using a Poisson correction model with gaps to complete deletion. Topological robustness was assessed by bootstrap analysis with 1000 replicates. The bar is an indicator of genetic distance based on branch length. Genes, which were indicated by red box, were selected to construct complementary lines. b-k, Phenotypes of yield-related traits of wild type ZH11, nal1-cri and the complementary (COM) lines constructed by NAL1-homologous genes from wheat (Ta), maize (Zm), rapeseed (Bn), and soybean (Gm). (b) Plant morphology, bar = 20 cm. (c) Flag leaf morphology, bar = 5 cm. (d) Root system, bar = 5 cm. (e) Panicle structure, bar = 5 cm. (f) Plant height (n = 10), (g) Flag leaf length (n = 8), (h) Flag leaf width (n = 8), (i) Root dry weight (n = 8), (j) Spikelet number per panicle (n = 10), (k) Grain yield per plant (n = 10). Data are means ± SD. Different lowercase letters above bars indicate significant difference at P < 0.05 level by one-way ANOVA. The exact P values are listed in Supplementary Table 6.
Extended Data Fig. 8 The expression of ACL in ZH11, nal1-cri and TN lines seedlings (n = 4).
RT-qPCR was repeated at least three times. Data are means ± SD. ** indicates significant differences between ZH11 and nal1-cri at P < 0.01, and * indicates significant differences between each TN lines and nal1-cri at P < 0.05 by two-sides Student’s t-test. The exact P values are listed in Supplementary Table 6.
Extended Data Fig. 9 Sequence alignment of plant NAL1 homologs.
Putative catalytic triad, ear motif, and interface residues are labeled with red star, blue circle, and green rectangle, respectively.
Supplementary information
Supplementary Tables 1–6
Table 1: Data collection, refinement and structural determination. Table 2: Homologous structure search for NAL1 by Dali. Table 3: NAL1 interaction proteins identified by IP–MS. Table 4: List of genes with significantly reduced expression and H3K18Ace levels in NAL1 mutants compared with wild-types. Table 5: Primers used in this study. Table 6: Individual P values of each figure.
Source data
Source Data Figs. 2 and 4 and Extended Data Figs. 1, 3 and 4
Unprocessed western blots and gels.
Source Data Table
Statistical source data for Figs. 1, 3, 4 and 5 and Extended Data Figs. 4, 5, 6, 7 and 8.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Li, W., Yan, J., Zhang, Y. et al. Serine protease NAL1 exerts pleiotropic functions through degradation of TOPLESS-related corepressor in rice. Nat. Plants 9, 1130–1142 (2023). https://doi.org/10.1038/s41477-023-01449-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41477-023-01449-2
This article is cited by
-
The catalytic triad of rice NARROW LEAF1 involves H234
Nature Plants (2024)
-
Genome-wide association study of leaf photosynthesis using a high-throughput gas exchange system in rice
Photosynthesis Research (2024)
-
Genetic and functional mechanisms of yield-related genes in rice
Acta Physiologiae Plantarum (2024)
-
Identification of Genomic Regions for Deep-Water Resistance in Rice for Efficient Weed Control with Reduced Herbicide Use
Rice (2023)