Abstract
Biallelic hypomorphic variants in PRORP have been recently described as causing the autosomal recessive disorder combined oxidative phosphorylation deficiency type 54 (COXPD54). COXPD54 encompasses a phenotypic spectrum of sensorineural hearing loss and ovarian insufficiency (Perrault syndrome) to leukodystrophy. Here, we report three additional families with homozygous missense PRORP variants with pleiotropic phenotypes. Each missense variant altered a highly conserved residue within the metallonuclease domain. In vitro mitochondrial tRNA processing assays with recombinant TRMT10C, SDR5C1 and PRORP indicated two COXPD54-associated PRORP variants, c.1159A>G (p.Thr387Ala) and c.1241C>T (p.Ala414Val), decreased pre-tRNAIle cleavage, consistent with both variants impacting tRNA processing. No significant decrease in tRNA processing was observed with PRORP c.1093T>C (p.Tyr365His), which was identified in an individual with leukodystrophy. These data provide independent evidence that PRORP variants are associated with COXPD54 and that the assessment of 5′ leader mitochondrial tRNA processing is a valuable assay for the functional analysis and clinical interpretation of novel PRORP variants.
Similar content being viewed by others
Introduction
Biallelic variants in Protein Only RNase P Catalytic Subunit (PRORP) have recently been associated with a newly defined syndrome, combined oxidative phosphorylation deficiency 54 (COXPD54) (MIM #619737), encompassing a phenotypic spectrum of Perrault syndrome and leukodystrophy [1]. Perrault syndrome (MIM #233400) is a rare, autosomal recessive, clinically and genetically heterogeneous disorder characterised by bilateral sensorineural hearing loss (SNHL) in both sexes and primary ovarian insufficiency in 46, XX karyotype females [2, 3]. The individuals with biallelic PRORP variants, reported to date, presented with pleiotropic phenotypes ranging in clinical severity, with PRORP variant protein resulting in reduced mitochondrial tRNA processing in vitro [1]. With the small number of reported affected individuals with COXPD54, genotype–phenotype correlations have yet to be established and independently replicated. PRORP (MRPP3) is one of the three subunits constituting the human mitochondrial RNase P (mtRNase P) complex with MRPP1 (TRMT10C) and MRPP2/SDR5C1 (HSD17B10). This multimeric complex aids mitochondrial tRNA maturation by catalysing endonucleolytic cleavage of the 5′ leader sequence from polycistronic mitochondrial RNA transcripts [4].
Here, we present three further unrelated families with homozygous, missense PRORP variants. Consistent with the previously reported cases, the variants are within the functional metallonuclease domain and alter highly conserved residues [1]. The affected individuals have phenotypes overlapping with the previous reports and provide evidence that the clinical severity correlates with the degree of mitochondrial tRNA processing deficit. These findings provide independent supportive evidence for the association of biallelic PRORP variants with defective mitochondrial tRNA processing, resulting in a phenotypic spectrum encompassing Perrault syndrome and COXPD54.
Materials and methods
Variant identification methodology
Biallelic variants in PRORP were identified by exome sequence analysis in all three families (see Supplementary information). Segregation analysis was undertaken using Sanger sequencing for all unaffected and affected relatives where available. Details of respiratory chain activity assays, immunoblotting and mitochondrial tRNA processing assays are also outlined in detail in the Supplementary information.
Results
The three probands presented with diverse clinical phenotypes outlined below. Additional clinical, in vitro and protein modelling data is available in the Supplementary information.
Family F1 is a consanguineous family with a 28-year-old female proband from Iran who presented with mild intellectual impairment, gait abnormality, behavioural problems and hypothyroidism. There is no evidence of SNHL or ovarian insufficiency; however, brain MRI demonstrates cerebellar atrophy and multifocal leukoencephalopathy (Supplementary Fig. S1A). Menstruation is irregular (once every 4–6 months), and she has bilateral polycystic ovaries. Biochemical tests of ovarian function revealed hormone levels were within follicular phase reference values (Supplementary Fig. S1B). Exome sequencing revealed a homozygous missense PRORP (NM_014672.4:c.1093T>C (p.Tyr365His)) variant in the proband. The parents of the proband are heterozygous and her two clinically unaffected siblings, a brother and sister, are wildtype and heterozygous, respectively (Fig. 1A).
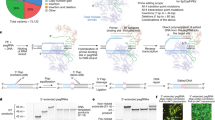
A Pedigrees for families F1–F3, each with homozygous PRORP variants in the proband. B Position of variant residues highlighted in orange. Symbols below the sequence alignments represent level of conservation across species, with * (asterisk) indicating full conservation, : (colon) indicating strongly similar properties, . (period) indicating weakly similar properties and no symbol indicating no conservation. Visualised using Clustal Omega alignment software. C Schematic illustrating the location of all established PRORP variants to date. Novel variants are coloured in orange, whilst previously published variants are in blue. Created using DOG (Domain Graph) software [9]. MTS mitochondrial targeting sequence, PPR pentatricopeptide repeat, CD central domain.
The proband from family F2 is a 7-year-old American male of Mexican background. He presented with severe to profound bilateral SNHL (Supplementary Fig. S1C), undergoing assessment aged 3 years, exhibiting global developmental delay, spastic diplegia and truncal hypotonia. He is also both nonverbal and non-ambulatory. Brain MRI displayed no white matter lesions. Trio exome sequencing of the proband revealed a homozygous variant in PRORP (NM_014672.4:c.1159A>G (p.Thr387Ala)), with no other candidate variants identified. Both parents are heterozygous for this variant and unaffected, whilst his healthy younger sibling is homozygous for the reference allele.
The proband from family F3 is the daughter of a consanguineous couple of Arab Muslim background from Israel. She was affected by isovaleric acidemia detected by newborn screening and homozygous for the IVD: c.941C>T;p.(Ala314Val) hypomorphic variant [5, 6]. The proband did not pass the neonatal hearing screening test on the right ear; however, a formal hearing test was not performed. She presented at 3.5 months with episodes of lactic academia (up to 11.5 mmol/l, normal range 2–4 mmol/l), preceded by a febrile infection, and was additionally diagnosed with atrial septal defect and severe pulmonary hypertension. She had repeatedly high lactate levels (2.6–6.4 mmol/l). She was affected by hypotonia, developmental delay and severe failure to thrive. Aged 1 year, she suffered a lactic acidosis crisis (8.4 mmol/l) with status epilepticus, after which she experienced epilepsy, extrapyramidal movement disorder and loss of developmental skills. She died at age 19 months. Brain MRI scans revealed cerebrospinal space dilation and diffuse restrictive changes in the perirolandic region and basal ganglia (Supplementary Fig. S1D). Exome sequencing identified the homozygous PRORP (NM_014672.4: c.1241C>T (p.Ala414Val)) variant. Both parents are heterozygous for the variant.
All three novel variants alter highly conserved residues (Fig. 1B) and are predicted deleterious by in silico analyses (Supplementary Table S2). The three variants are also absent from gnomAD, providing supportive evidence for pathogenicity [7].
To assess whether the novel PRORP variants altered mitochondrial tRNA processing, we purified recombinant PRORP with the disease-associated variants and tested the endonucleolytic activity of the mtRNase P complex in the presence of pre-tRNAIle in vitro (Supplementary Fig. S3). After incubation with pre-tRNAIle, mtRNase P complex with the p.Thr387Ala (F2) or p.Ala414Val (F3) PRORP variants significantly diminished the levels of 5′ cleavage product (p = 0.0121 and <0.0001 respectively) in comparison to wildtype PRORP (Fig. 2). The percentage decreases in relative intensity were 29% and 59%, respectively. The p.Tyr365His PRORP variant had little impact on 5′ leader processing, with a modest 4% decrease (p = 0.9330).
A Cleavage of the pre-tRNAIle 5′ leader sequence by mtRNase P containing wildtype (WT) or variant PRORP. The relative intensity of the pre-tRNAIle cleavage product was quantified with error bars representing the standard error of the mean. N = 4, *p < 0.05, ****p < 0.0001, one-way ANOVA with Dunnett’s multiple comparisons test, comparing wildtype to variants. B Comparison of mtRNase P activity to previously published PRORP variants. N = 5, *p < 0.05, ****p < 0.0001, one-way ANOVA with Dunnett’s multiple comparisons test, comparing wildtype to variants.
We also independently compared the novel p.Thr387Ala and p.Ala414Val variants to a disease-associated PRORP variant to assess the reproducibility of the tRNA processing assay [1]. The p.Arg421Cys and p.Asn412Ser PRORP variants were evaluated in the tRNA processing assay because they altered the cleavage of pre-tRNAIle to variable degrees [1]. The novel p.Thr387Ala and p.Ala414Val PRORP variants significantly reduced cleavage of the mtRNase P complex by 25% and 52% (p = 0.0153 and <0.0001), whilst the p.Arg421Cys and p.Asn412Ser variants diminished cleavage of the mtRNase P complex by 13% and 75% (p = 0.2873 and <0.0001) respectively, comparable to values in the previous report [1]. The variants p.Thr387Ala and p.Ala414Val were subsequently classified according to ACMG guidelines as likely pathogenic, whereas p.Tyr365His was classified as a variant of uncertain significance (VUS), as submitted to ClinVar.
Discussion
Biallelic PRORP variants have been associated with diverse, overlapping pleiotropic phenotypes, with variable defects in mitochondrial tRNA processing [1]. The aim of this work was to independently confirm that novel PRORP variants identified in additional families are pathogenic and associated with the COXPD54 phenotypic spectrum.
Because all novel variants were located within the functional metallonuclease domain (Fig. 1C), we investigated the ability of mtRNase P complexes with PRORP variants to conduct 5′ leader processing of pre-tRNAIle. mtRNase P complexes with the novel PRORP (p.Thr387Ala and p.Ala414Val) variants significantly decreased the intensity of the cleavage product compared to wildtype, indicating the function of the mtRNase P complex was impaired. The defect in 5′ leader processing was more pronounced with the variant from the F3 proband with a more severe phenotype. Fibroblasts from the affected individual in F3 demonstrated deficiencies in complex I activity (Supplementary Fig. S2B), highlighting a respiratory chain defect.
In contrast, mtRNase P complexes with PRORP p.Tyr365His did not significantly reduce pre-tRNAIle 5′ end cleavage. There are potential explanations for this finding. For example, protein modelling demonstrates that amino acid 365 is not located in the vicinity of the active site or any interaction regions (Supplementary Fig. S4). Furthermore, residues surrounding the amino acid are less conserved than the residues proximate to other disease-associated variants (Fig. 1). These observations indicate this region of the protein may be less essential for regular endonucleolytic function. The p.Tyr365His variant may also marginally reduce mtRNase P processing efficacy to levels which are difficult to detect experimentally but could still diminish processing in specific tissues. It is also possible that a different pathogenic mechanism is responsible for this phenotype, or that this variant does not cause COXPD54.
With the discovery of additional affected families, comparisons can begin to be made between PRORP variants and associated phenotypes (Supplementary Table S3). The proband with the p.Tyr365His PRORP VUS has a similar presentation to the individuals in the initial discovery study with the p.Arg421Cys PRORP variant [1]. Affected individuals in both families presented with leukodystrophy, mild learning disabilities and behavioural abnormalities, with no hearing impairment or ovarian insufficiency. Both variants also had little impact on mitochondrial tRNA processing. The probands from F3 in this study (PRORP p.Ala414Val) and P3 in the discovery study (PRORP p.Arg445Gln;p.Ser400Ilefs*6) were also comparable, presenting with severe childhood-onset features, including lactic acidosis, hypotonia, and seizures with white matter changes. Both missense variants reduced the levels of cleaved pre-tRNAIle in vitro, suggesting a relationship of increased phenotypic severity correlated with diminished mtRNase P function.
The p.Ala414Val PRORP variant reported here is in close proximity to the variant p.Asn412Ser reported previously [1], with both variants resulting in a significant decrease in cleavage product. Interestingly, the phenotype in individual P2 in the previous report [1] of isolated SNHL was less severe than that observed in F3 in this study. This difference in phenotypic severity could be because the F3 proband is homozygous for the p.Ala414Val variant, whereas individual P2 was compound heterozygous for p.Asn412Ser and the less deleterious c.1301C>A (p.Ala434Asp) PRORP variant.
Utilising the ClinGen scoring criteria for gene-disease validity [8], the initial PRORP discovery paper scored 9.75, which is concordant with moderate evidence for disease association. With the addition of our families, the score was increased to 12, upgrading the gene-disease association to strong. A third independent report is required for definitive confirmation that PRORP variants cause disease.
In summary, these data provide the first independent confirmation that biallelic PRORP missense variants can reduce mitochondrial tRNA processing in vitro and are associated with variable, overlapping pleiotropic phenotypes consistent with COXPD54. A possible association between phenotypic severity and mitochondrial tRNA processing deficit is starting to emerge, and further cases need to be defined to determine if robust genotype–phenotype correlations exist. Investigating alternate mechanisms of pathology may elucidate how variants with limited effect on mt-tRNA processing (p.Tyr365His) may cause disease.
Data availability
The PRORP variants were submitted to ClinVar (https://www.ncbi.nlm.nih.gov/clinvar/) (GenBank: NM_014672.4; accession numbers SCV002820061–SCV002820063).
References
Hochberg I, Demain LAM, Richer J, Thompson K, Urquhart JE, Rea A, et al. Bi-allelic variants in the mitochondrial RNase P subunit PRORP cause mitochondrial tRNA processing defects and pleiotropic multisystem presentations. Am J Hum Genet. 2021;108:2195–204.
Perrault M, Klotz B, Housset E. Two cases of Turner syndrome with deaf-mutism in two sisters. Bull Mem Soc Med Hop Paris. 1951;67:79–84.
Faridi R, Rea A, Fenollar-Ferrer C, O’Keefe RT, Gu S, Munir Z, et al. New insights into Perrault syndrome, a clinically and genetically heterogeneous disorder. Hum Genet. 2022;141:805–19.
Holzmann J, Frank P, Löffler E, Bennett KL, Gerner C, Rossmanith W. RNase P without RNA: identification and functional reconstitution of the human mitochondrial tRNA processing enzyme. Cell. 2008;135:462–74.
Mohsen AW, Anderson BD, Volchenboum SL, Battaile KP, Tiffany K, Roberts D, et al. Characterization of molecular defects in isovaleryl-CoA dehydrogenase in patients with isovaleric acidemia. Biochemistry. 1998;37:10325–35.
Ensenauer R, Vockley J, Willard JM, Huey JC, Sass JO, Edland SD, et al. A common mutation is associated with a mild, potentially asymptomatic phenotype in patients with isovaleric acidemia diagnosed by newborn screening. Am J Hum Genet. 2004;75:1136–42.
Karczewski KJ, Francioli LC, Tiao G, Cummings BB, Alföldi J, Wang Q, et al. The mutational constraint spectrum quantified from variation in 141,456 humans. Nature. 2020;581:434–43.
Rehm HL, Berg JS, Brooks LD, Bustamante CD, Evans JP, Landrum MJ, et al. ClinGen–the clinical genome resource. N Engl J Med. 2015;372:2235–42.
Ren J, Wen L, Gao X, Jin C, Xue Y, Yao X. DOG 1.0: illustrator of protein domain structures. Cell Res. 2009;19:271–73.
Acknowledgements
We would like to thank the families for their participation.
Funding
This study was supported by the Medical Research Council (MR/W019027/1), the Royal National Institute for the Deaf/Masonic Charitable Foundation (S60_Newman) and the NIHR Manchester Biomedical Research Centre (NIHR 203308). RWT is supported by the Wellcome Centre for Mitochondrial Research (203105/Z/16/Z), the Mitochondrial Disease Patient Cohort (UK) (G0800674), the Medical Research Council International Centre for Genomic Medicine in Neuromuscular Disease (MR/S005021/1), the Lily Foundation, Mito Foundation, the Pathological Society, the UK NIHR Biomedical Research Centre for Ageing and Age-related disease award to the Newcastle upon Tyne Foundation Hospitals NHS Trust and the UK NHS Highly Specialised Service for Rare Mitochondrial Disorders of Adults and Children. MO is funded by Mito Foundation, The Lily Foundation and The Pathological Society.
Author information
Authors and Affiliations
Contributions
TBS, WGN and RTO contributed to the conception and design of the study. RWT, WGN and RTO participated in the project supervision. RM, MZ, LH, SS, HG, NSL, TG, KCH and HH were involved in patients’ medical care and interpretation of genetic analyses. TBS, AR, HBT, KT and MO contributed to functional work. TJM and WWY oversaw modelling predictions. All authors contributed to the acquisition and/or analysis of data. TBS, RWT, WGN and RTO wrote the manuscript. All authors revised it critically and approved the final version.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics approval and consent to participate
All individuals (or their legal guardians) provided written informed consent in accordance with local regulations. Ethical approval for this study was granted by the National Health Service Ethics Committee (16/WA/0017) and University of Manchester.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Smith, T.B., Rea, A., Thomas, H.B. et al. Novel homozygous variants in PRORP expand the genotypic spectrum of combined oxidative phosphorylation deficiency 54. Eur J Hum Genet 31, 1190–1194 (2023). https://doi.org/10.1038/s41431-023-01437-2
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41431-023-01437-2
This article is cited by
-
Expanding what we know about rare genetic diseases
European Journal of Human Genetics (2023)