Abstract
Low-frequency sensorineural hearing loss (SNHL) is a rare hearing impairment affecting frequencies below 1000 Hz, previously associated with DIAPH1, WSF1, MYO7A, TNC, SLC26A4 or CCDC50 genes. By exome sequencing, we report a novel nonsense variant in CENPP gene, segregating low-frequency SNHL in five affected members in a Swiss family with autosomal dominant inheritance pattern. Audiological evaluation showed up-sloping audiometric configuration with mild-to-moderate losses below 1000 Hz, that progresses to high-frequencies over time. Protein modeling shows that the variant truncates five amino acids at the end, losing electrostatic interactions that alter protein stability. CENPP gene is expressed in the supporting cells of the organ of Corti and takes part as a subunit of the Constitutive Centromere Associated Network in the kinetochore, that fixes the centromere to the spindle microtubules. We report CENPP as a new candidate gene for low-frequency SNHL. Further functional characterization might enable us to elucidate its molecular role in SNHL.
Similar content being viewed by others
Introduction
Genetic non-syndromic sensorineural hearing loss (SNHL) comprises a group of different disorders, involving high frequencies (>2000 Hz) in 70% of cases [1]. On the other hand, low-frequency SNHL is considered a rare condition, with an estimated prevalence of 18 cases per 100,000 population, showing mostly an autosomal dominant pattern of inheritance [2].
The clinical profile of low-frequency SNHL includes a post-lingual early onset, mild to moderate hearing loss at frequencies below 1000 Hz, usually symmetrical and bilateral, that can be associated with tinnitus, without vestibular symptoms [3]. It is progressive, where thresholds worsen over time extending to all frequencies [4]. When low-tones SNHL is associated with episodes of vertigo, aural fullness or tinnitus, the condition is considered as early stage of Meniere Disease (MD) [5].
Six genes have been associated with low-frequency SNHL, including diaphanous homolog 1 gene (DIAPH1), wolframin gene (WSF1), tenascin-C gene (TNC), CCDC50 gene, pendrin gene (SLC26A4), myosin VIIA (MYO7A), and loci DFNA54, with no gene resolved (Table S1).
Here, we present a Swiss family with low-frequency SNHL segregating a nonsense variant in the CENPP gene and modeling the effect of the variant on the protein structure.
Material and methods
Patients
The study was approved by the local Ethical Committee (KEK-ZH-Nr.2019-01006, Kantonale Ethikkommission, Zurich, Switzerland) in accordance with the Helsinki declaration and its amendments. Written informed consent was obtained from participants. A German-speaking Swiss (CH/DE) family was identified at the University Hospital of Zurich showing low-frequency SNHL in five individuals. An audiovestibular evaluation was performed in affected patients including a 3T gadolinium-enhanced MRI to exclude intra/retrolabyrinthine pathology [6].
Exome sequencing
Patient saliva samples were collected with Oragene®DNA kit (DNA-Genotek, OG-510, Ottawa, Canada) and gDNA was extracted as previously reported [7]. Coding regions capture and subsequent libraries were prepared with SureSelect Human All Exon 500 Mb (Agilent Technologies, Santa Clara, CA, USA). Exome sequencing was performed in HiSeq 6000 platform (Illumina) and pair-end reads were generated with a mean of 100× coverage.
Bioinformatics data processing
Sarek nextflow v2.7.1 (NF-ACore) was used for sequencing data processing [8]. Raw data sequence was aligned with reference genome GRCh38/Hg38 using Burrows-Wheeler Aligner algorithm. BAM files were subsequently marked for duplicates and quality recalibrated (Base Quality Score Recalibration). After clean-up operations, variants were called using HaplotypeCaller from Genome Analysis Toolkit v4.1, generating a Variant Call Format file, with Single Nucleotide Variants and Insertions/Deletions from each individual.
A family variant dataset was created by merging variant files from all individuals (BCFtools, https://samtools.github.io). Final selection of high-quality variants was performed by executing the Variant Quality Score Recalibration separately from both types of variants, removing those with values under quality threshold or average depth of coverage <10. Variants were annotated with the Variant Ensembl Variant Predictor tool (Ensembl).
Filtering and prioritization strategy
Variant filtering and prioritization strategy are summarized in Fig. S1. We selected those variants shared between the five relatives segregating low-frequency SNHL (I-4, II-4, II-5, III-1, and III-2), but not found in II-3. Variants classified as “HIGH” or “MODERATE”, according to Variant Ensembl Variant Predictor protein consequences were retained. A conservative Minor Allele Frequency ≤0.001 in the non-Finnish European population from gnomADv3.1, and a pathogenic phred-like Combined Annotation Dependent Depletion score ≥20 thresholds were used as cut-off. Candidate variants were submitted for publication in ClinVar database (Submission ID: SUB11466822).
We prioritized “Pathogenic” or “Likely Pathogenic” variants classified by the American College of Medical Genetics. Inner ear expression of genes from candidate variants was confirmed in the gEAR portal (umgear.org). Previous associations with hearing loss in humans (deafnessvariationdatabase.org; hereditaryhearingloss.org) and mice (www.informatics.jax.org/vocab/mp_ontology) were also examined (Fig. S1).
Computational modeling
The human CENPP amino acid sequence was retrieved from Uniprot (Q6IPU0). Structural models were generated using fold-recognition and ab initio methods (Table S3), obtaining best structural validation results with Robetta-ab [9]. This model was used to predict the protein stability change of mutation using SCOOP [10].
The mutated CENPP protein (CENPP:p.Cys283*) was modeled by comparative homology modeling using MODELLER 10.1 [11] and the wild-type CENPP protein model as a template.
Results
Characterization of study family
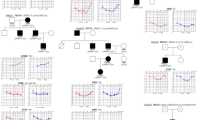
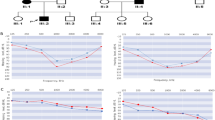
Low-frequency SNHL was segregated in five individuals (I-4, II-4, II-5, III-1, and III-2) across three consecutive generations, suggesting an autosomal dominant inheritance pattern (Fig. 1A). Hearing loss was bilateral and symmetrical and showed anticipation, starting in early 50s, 30s and 20s in first, second and third generation, respectively (Table S2). Air conduction thresholds were used to estimate the progression in the family over time (Fig. 1B). Hearing impairment debuted affecting primarily frequencies below 500 Hz in a mild way (20–40 dB). Deterioration was observed over 50–60 years in a moderate level (30–50 dB) at low tones, also starting a threshold decrease in high frequencies. In the elderly patient, the decline in low-frequencies stabilized, but high-tones worsened to moderate-to-severe levels (60–80 dB), leading to a down-sloping configuration. Interestingly, III-1 was diagnosed with MD in the second decade of life, showing attacks of spinning vertigo of several hours of duration, aural pressure, and low-frequency unilateral hearing loss in left ear. No other comorbidity was reported by any of the patients. Endolymphatic space MRIs revealed endolymphatic hydrops in the ear affected by MD in III-1.
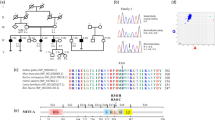
A Familial pedigree segregating autosomal low-frequency SNHL in a dominant inheritance pattern. Full black filled symbols: Meniere Disease; half lower black filled symbols: low-frequency SNHL; strikethrough symbols: deceased individual. Gray numbers at the left of the symbol indicate the age of onset; B Air conduction audiograms in affected individuals. The decade of life of patients during audiometry is shown below the chart. Red series: left ear; blue series: right ear; yo: years old; dB: decibels; kHz: kilohertzs. C CENPP modeled by Robetta-AB method. Shaded area includes the truncated region because of the variant, deleting from cysteine 283 onward. Amplified panels show the loss of polar interactions of glutamate 285 and alanine 284 with arginine 213 between wildtype and mutated protein.
Prioritization of candidate variants
The variant filtering and prioritization strategy is summarized in Fig. S1. The family variant dataset included 51,324 variants. A total of 17,148 variants were shared between patients I-4, II-4, II-5, III-1, and III-2, segregating the phenotype. After filtering Single Nucleotide variants and Insertions/Deletions observed in the unaffected relative (II-3), 2654 remained and 626 were classified as variants with “HIGH” or “MODERATE” effect according to VEP. Among them, ten heterozygous variants showed a Minor Allele Frequency ≤0.001 and five had a phred-like Combined Annotation Dependent Depletion pathogenic score ≥20. Final candidate variant list consisted of 1 stop-gain and 4 missense variants (Table 1).
A novel nonsense variant in the CENPP gene was top-ranked, being validated by sanger sequencing in patients I-4, II-4, II-5 and III-2 (Fig. S2-A). The variant (CENPP, NC_000009.12:c.92613131T>A p(C283*)) affects only to canonical transcript (ENST00000375587.8) in exon 8 (Fig. S2-B). It generates a premature stop codon which truncates the last 5 of 288 total length amino acids of the protein. The conservation of affected C283 residue is conserved between higher organisms (chimpanzee, macaque, gorilla), but not in animal models such as mice or rats (Fig. S2-C).
Protein modeling
The CENPP protein was modeled using Robetta-ab method (Fig. 1C), being reliable compared to structures solved by experimental techniques at geometric level (Table S3). The variant (CENPP, NC_000009.12:c.92613131T>A p(C283*)) causes the deletion of 5 residues at the end of the protein, from cystein283 and onwards, with a slight reduction in protein stability (SCOOP: ΔΔG = 0.4 (kcal/mol), destabilizing). The deletion could lead to the loss of polar interactions between glutamate 285 and alanine 284 with arginine 213 (Fig. 1C, amplified region).
Discussion
We report CENPP as a new candidate gene for autosomal dominant low-frequency SNHL in a Swiss family. By exome sequencing, we have found a novel heterozygous nonsense variant (CENPP, NC_000009.12:c.92613131T>A p(C283*)), that segregated the phenotype in five family members across three generations with an autosomal dominant inheritance pattern. The stop-gain variant was top-ranked according to its high impact consequence in the protein, its highest phred-like Combined Annotation Dependent Depletion pathogenic score and its “Pathogenic” classification by the American College of Medical Genetics.
The genetic underpinnings of low-frequency SNHL are not well-known. Currently, six genes have been associated with the condition: DIAPH1, WSF1, TNC, SLC26A4, CCDC50, and MYO7A. Martín-Sierra et al. also described a variant in PRKCB gene segregating low-frequency SNHL in a Spanish family with MD. In all cases, the observed inheritance pattern was autosomal dominant [12].
The audiometric profile in this family showed a symmetrical and bilateral decrease in the hearing threshold for frequencies <1 kHz, although for II-5 and III-2 was limited below 500 Hz. In elderly patient, impairment extends to all frequencies (I–4). On the other hand, individual III-1 was diagnosed with MD showing low-frequency SNHL in the left ear. Although the right ear had normal hearing levels in his late 10 s, he could develop a bilateral affection. Of note, the low-frequency SNHL was accompanied by endolymphatic hydrops only in the patient with MD. This finding supports earlier studies, which describe that isolated low-frequency hearing loss is only inconsistently associated with endolymphatic hydrops [13].
The onset of autosomal dominant low-frequency SNHL begins in the first or second decade of life for most of the genes previously reported, as we have observed in this family. In some reports, the onset was delayed up to 30–40 s in carriers of mutations in WFS1 and TNC genes [14, 15]. However, isolated low-frequency SNHL can be unnoticed by many individuals, causing a diagnostic delay.
The CENPP gene encodes one of the proteins of the Constitutive Centromere Associated Network in the kinetochore [16]. During last decades, up to 16 proteins have been discovered as part of this complex, associated with five sub-complexes: CENP-C, CENP-LN, CENP-HIKM, CENP-OPQRU, and CENP-TWSX [16, 17]. CENP-P protein is part of the CENP-OPQRU complex, for which there is increasing evidence of its role in chromosome alignment in the spindle equator through direct interaction with microtubules, and motor protein recruitment [17, 18].
In the mouse cochlea, CENPP gene is highly expressed in embryonic and postnatal stages [19], particularly in supporting Pillar and Deiters cells in the organ of Corti [20]. However, no other variants in this gene have been associated with SNHL and further families segregating the phenotype and an animal model will be needed to confirm CENPP as a causal gene for low-frequency SNHL.
Data availability
All data generated or analyzed during this study are included in this published article and its supplementary information file. Anonymized familial variant dataset is available from the corresponding author on reasonable request. The CENPP structural model was submitted to the ModelArchive database (https://www.modelarchive.org/doi/10.5452/ma-ja069; Public access after publication. Temporary access code: FUxtQ7paHI).
References
Sheffield AM, Smith RJH. The epidemiology of deafness. Cold Spring Harb Perspect Med. 2019;9:a033258.
Parving A, Sakihara Y, Christensen B. Inherited sensorineural low-frequency hearing impairment: some aspects of phenotype and epidemiology. Int J Audio. 2000;39:50–60.
Bramhall NF, Kallman JC, Verrall AM, Street VA. A novel WFS1 mutation in a family with dominant low frequency sensorineural hearing loss with normal VEMP and EcochG findings. BMC Med Genet. 2008;9:48.
Oishi N, Inoue Y, Saito H, Kanzaki S, Kanzaki J, Ogawa K. Long-term prognosis of low-frequency hearing loss and predictive factors for the 10-year outcome. Otolaryngol Neck Surg. 2010;142:565–9.
Perez-Carpena P, Lopez-Escamez JA. Current understanding and clinical management of Meniere’s disease: a systematic review. Semin Neurol. 2020;40:138–50.
Baráth K, Schuknecht B, Naldi AM, Schrepfer T, Bockisch CJ, Hegemann SCA. Detection and grading of endolymphatic hydrops in Menière disease using mr imaging. Am J Neuroradiol. 2014;35:1387–92.
Szczepek AJ, Frejo L, Vona B, Trpchevska N, Cederroth CR, Caria H, et al. Recommendations on collecting and storing samples for genetic studies in hearing and tinnitus research. Ear Hear. 2019;40:219–26.
Garcia M, Juhos S, Larsson M, Olason PI, Martin M, Eisfeldt J, et al. Sarek: a portable workflow for whole-genome sequencing analysis of germline and somatic variants. F1000Research. 2020;9:63.
Kim DE, Chivian D, Baker D. Protein structure prediction and analysis using the Robetta server. Nucleic Acids Res. 2004;32(Web Server):W526–31.
Pucci F, Kwasigroch JM, Rooman M. SCooP: an accurate and fast predictor of protein stability curves as a function of temperature. Bioinforma. 2017;33:3415–22. Valencia A, editor.
Fiser A, Šali A. Modeller: generation and refinement of homology-based protein structure models. In: Methods in enzymology [Internet]. Elsevier. 2003. p. 461–91. https://linkinghub.elsevier.com/retrieve/pii/S0076687903740208. Accessed 19 March 2022.
Martín-Sierra C, Requena T, Frejo L, Price SD, Gallego-Martinez A, Batuecas-Caletrio A, et al. A novel missense variant in PRKCB segregates low-frequency hearing loss in an autosomal dominant family with Meniere’s disease. Hum Mol Genet. 2016;25:3407–15.
Qin H, He B, Wu H, Li Y, Chen J, Wang W, et al. Visualization of endolymphatic hydrops in patients with unilateral idiopathic sudden sensorineural hearing loss with four types according to Chinese criterion. Front Surg. 2021;8:682245.
Gurtler N, Kim Y, Mhatre A, Schlegel C, Mathis A, Lalwani AK. DFNA54, a third locus for low-frequency hearing loss. J Mol Med. 2004;82:775–80.
Zhao Y, Zhao F, Zong L, Zhang P, Guan L, Zhang J, et al. Exome sequencing and linkage analysis identified Tenascin-C (TNC) as a novel causative gene in nonsyndromic hearing loss. Janecke AR, editor. PLoS One. 2013;8:e69549.
Musacchio A, Desai A. A molecular view of kinetochore assembly and function. Biology. 2017;6:5.
Pesenti ME, Prumbaum D, Auckland P, Smith CM, Faesen AC, Petrovic A, et al. Reconstitution of a 26-subunit human kinetochore reveals cooperative microtubule binding by CENP-OPQUR and NDC80. Mol Cell. 2018;71:923–39.e10.
Bancroft J, Auckland P, Samora CP, McAinsh AD. Chromosome congression is promoted by CENP-Q- and CENP-E-dependent pathways. J Cell Sci. 2014; https://doi.org/10.1242/jcs.163659.
Scheffer DI, Shen J, Corey DP, Chen ZY. Gene expression by mouse inner ear hair cells during development. J Neurosci. 2015;35:6366–80.
Liu H, Chen L, Giffen KP, Stringham ST, Li Y, Judge PD, et al. Cell-specific transcriptome analysis shows that adult pillar and deiters’ cells express genes encoding machinery for specializations of cochlear hair cells. Front Mol Neurosci. 2018;11:356.
Acknowledgements
We would like to thank the patients of this family for their enthusiastic participation.
Funding
This project was partially funded by H2020-SC1-2019-848261: Unification of Treatments and Interventions for Tinnitus Patients—UNITI; and the Swiss Schmieder-Bohrisch Foundation. JAL-E is partially funded by the Asociacion Sindrome de Meniere España (ASMES) and Meniere Society, UK. PR-B is a PhD student in the Biomedicine Program at Universidad de Granada and his salary was supported by H2020-SC1-2019-848261. DB is supported by a national MD-PhD scholarship from the Swiss National Science Foundation. Additional funding:- Papel de la regulación epigenómica en la penetrancia y expresividad en la enfermedad de Meniere (EPIMEN) - CECEU PY20-00303.- Regional Ministry of Economic Transformation, Industry, Knowledge, and Universities of Junta de Andalucía - PREDOC2021/00343.- Career development grant (“Filling the Gap”) from the University of Zurich, Switzerland.
Author information
Authors and Affiliations
Contributions
AE and DB recruited and obtained clinical data from the family. PR-B, PR-N, and AE-B conducted DNA extraction and quality controls. PR-B, AE-B and AG-M performed bioinformatics analyses in WES data. AMP-P performed the protein modeling. PR-B and JAL-E drafted the manuscript. All authors approved the final version of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethical approval
This study involves human participants and was approved by the local Ethical Committee (KEK-ZH-Nr. 2019-01006, Kantonale Ethikkommission, Zurich, Switzerland) in accordance with the Helsinki declaration and its amendments. Written informed general consent was obtained from all participants.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Robles-Bolivar, P., Bächinger, D., Parra-Perez, A.M. et al. A novel nonsense variant in the CENPP gene segregates in a Swiss family with autosomal dominant low-frequency sensorineural hearing loss. Eur J Hum Genet 30, 1301–1305 (2022). https://doi.org/10.1038/s41431-022-01184-w
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41431-022-01184-w
This article is cited by
-
Genome sequencing—do you know what you are getting into?
European Journal of Human Genetics (2022)